Assays and Tools to Monitor Biology in 3D Cell Cultures
Researchers are turning to 3D culture methods such as spheroids and organoids to create model systems for studying cellular biology. 3D cell culture mimics the tissue or tumor environment more closely than a 2D environment by facilitating more natural interactions with a matrix and even allowing cells to secrete their own matrix. 3D cell culture also offers more natural cell-to-cell contacts and produces a gradient of access to O2 and nutrients.
Often, 3D cell culture can reveal dramatic differences in cellular responses compared to monolayer culture. Monitoring the biology of cells in 3D culture can be approached by looking at changes in cell health, metabolism and expression. Ultimately, these phenotypic changes are related back to the genotype of the cell or the alterations in the genotype that led to a disease state. We offer assays and tools to address each step in your workflow, from cell health and metabolic changes to gene expression and analysis.
New to 3D Cell Culture?
Learn about 3D cell cultures, why and how they are used, and how to choose assays for 3D models.
How to monitor biology in 3D culture?
When working with 3D culture models, choosing the right assay system is crucial. Learn about tools to monitor biology in 3D culture.
Cell Health Changes
Most cell health assays were designed for monitoring cells cultured in monolayers. Using these assays with cells cultured in 3D, be it spheroid cultures, organoids or on a hydrogel matrix, offer challenges. Standard protocols are not likely to work with 3D cultures if you need to isolate a protein or a metabolite from the cytoplasm. The protocol and reagent(s) must be optimized for 3D cultures. The CellTiter- Glo® 3D Assay is optimized with more detergent and a specialized protocol. We have developed protocols optimized for use with 3D culture for several of our other assays typically used in monolayer culture.
FEATURED PRODUCT
CellTiter-Glo® 3D Cell Viability Assay
Lytic Assay
Monitor cell viability through measurement of ATP with a reagent specifically designed for 3D cultures. Utilizes the same ATP measurement luciferase chemistry of the classic CellTiter-Glo® Assay with greater lytic power, which allows deeper penetration yielding greater ATP release for quantitation. Co-measure dead cells with same-well multiplexing with CellTox™ Green Cytotoxicity assay or media sampling with the LDH-Glo™ Cytotoxicity Assay.
Lytic Capacity of CellTiter-Glo® 3D Compared to ATPlite 1Step Reagent

FEATURED APPLICATION
Drug Screening Using Single Organoids
Organoids provide an essential tool for more accurately predicting in vivo responses from drug screening campaigns as they are more representative of the cellular heterogeneity within human tissue. Despite the pressing need for such models in drug screening, adoption of organoid in vitro assays for HTS is still limited due to workflow challenges, including lack of reproducibility and scalability. Highlighted here, Cell Microsystems and Promega provide an automated solution for reproducibly generating hundreds of organoids ready for use in plate-based assays that have been optimized and validated for 3D cell culture models.

Viability of unsorted compared to sorted organoids.
CELL VIABILITY
RealTime-Glo™ MT Cell Viability Assay
Non-Lytic Assay
Monitor cell viability through a bioluminescent assay continuously for up to 72 hours. Extremely sensitive assay monitors cellular reducing equivalents. Isolate nucleic acid from the same well for gene expression or genome analysis. Multiplex with CellTox™ Green to also measure dead cells.
Response of RealTime-Glo™ MT Cell Viability Assay to Increasing HCT116 Spheroid Diameter

Details in Application Note (link below).
CYTOTOXICITY
LDH-Glo™ Cytotoxicity Assay
Non-Lytic Assay
Measure cytotoxicity through lactate dehydrogenase release with this sensitive bioluminescent assay. The assay requires only 2–5μl of culture media per time point allowing multiple samplings from the same well. Use to measure cytotoxicity prior to other cellular assays like CellTiter- Glo® 3D or Caspase-Glo® 3/7 Assay.
Multiplexing the LDH-Glo™ Assay and the CellTiter-Glo® Assay

Human liver microtissue spheroids were treated with aflatoxin B1 for 48 hours. Media samples were collected and assayed with the LDH-Glo™ Assay. The remaining cells were assayed for viability with the CellTiter-Glo® 3D Assay. Details available in Technical Manual TM548.
CYTOTOXICITY
CellTox™ Green Cytotoxicity Assay
Non-Lytic Assay
Monitor cytotoxicity through a fluorescent assay continuously for 72 hours. Cell impermeable DNA binding dye enters cells with damaged membranes and stains cellular DNA. Multiplex with RealTime- Glo™ MT for continuous monitoring of live and dead cells or use upstream of CellTiter-Glo® 3D end-point analysis.
Same Well Multiplexing to Monitor Live and Dead Cells

Details in Scientific Poster (link below).
APOPTOSIS
Caspase-Glo® 3/7 3D Assay
Lytic Assay
Monitor apoptosis using a sensitive bioluminescent assay that measures caspase-3 and -7 activities. Multiplex with cytotoxicity and cell viability assays for a more complete characterization of the apoptotic response.
Multiplexing the CellTox™ Green Cytotoxicity Assay, CellTiter-Fluor™ Cell Viability Assay and Caspase-Glo® 3/7 3D Assay

Same-well multiplexing on A549 spheroids after a 24-hour exposure to bortezomib. Fluorescence and luminescence were measured using the GloMax® Discover Instrument.
APOPTOSIS
RealTime-Glo™ Annexin V Apoptosis and Necrosis Assay
Non-Lytic Assay
Monitor apoptosis and secondary necrosis through a multiplexed bioluminescent annexin V assay and a fluorescent necrosis assay continuously for up to 48 hours. Isolate nucleic acid from the same well for gene expression or genome analysis.
Measurement of Annexin V Exposure and Secondary Necrosis of HepG2 Spheroids

Measurement of annexin V exposure and secondary necrosis of HepG2 spheroids to 48 hour exposure to paclitaxel using the RealTime-Glo™ Annexin V Apoptosis and Necrosis Assay. Luminescence and fluorescence measured with the GloMax® Discover Instrument.
ADME
P450‐Glo™ CYP3A4 Assay and Screening System
Non-Lytic Assay Option
Monitor activation of cytochrome P450 enzymes (CYP1A2 and CYP2C9 also available) through a bioluminescent assay. Non-lytic protocol option uses same protocol for monolayer and 3D cultures where prosubstrates diffuse into cells get converted to luciferin through CYP activity. Luciferin diffuses out of the cell and assay is performed on culture media. Cells can be used for other cell-based assays or nucleic acid extraction.
Monitoring Human Liver Microtissue CYP3A4 Activity and Viability

Monitoring human liver microtissue Cytochrome P450 3A4 activity and viability in response to rifampicin. CYP3A4 activity was measured using non-lytic P450-Glo™ 3A4 Assay with Luciferin-IPA followed by same-well measurement of viability with the CellTiter- Glo® 3D Assay. Measured with a GloMax® Discover Instrument.
AUTOPHAGY
Autophagy LC3 HiBiT Reporter Assay System
Non-Lytic Assay
Monitor autophagic LC3-II flux with a bioluminescent plate-based assay. Stably transfect cells with the LC3 HiBiT Reporter, grow in 3D culture and monitor using the Lytic HiBiT detection System. Reporter also allows monitoring of LC3-II protein through fluorescent microscopy.
Response of LC3 HiBiT Reporter- Expressing HEK293 Spheroids to Autophagy Stimulators and Inhibitors

Details in Scientific Poster (link below).
IMMUNOGENIC CELL DEATH
RealTime-Glo™ Extracellular ATP Assay
Non-Lytic Assay
The RealTime-Glo™ Extracellular ATP Assay is a bioluminescent assay designed for kinetic monitoring of ATP release from dying, stressed or activated cells. Extracellular ATP can function as a damage associated molecular pattern (DAMP) and is a key biomarker for determining whether a treatment induces immunogenic cell death, a specialized form of cell death that results in an immune response.
ATP Release from HCT-116 Spheroids Treated with Staurosporine.
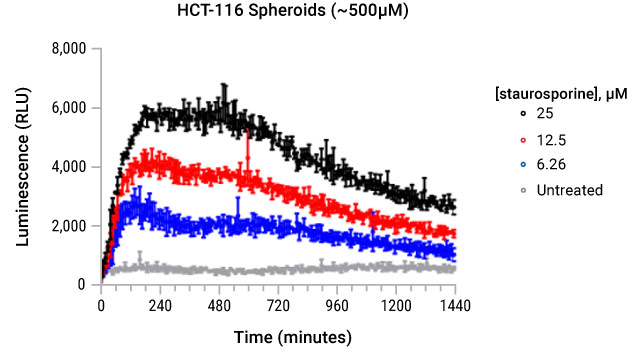
ATP release was monitored with the RealTime-Glo™ Extracellular ATP Assay and measured every 10 minutes over a 24 hour period.
Metabolic Changes
Energy metabolism is critical for cellular health and function and metabolites are linked to cellular energy, creation of cellular building blocks and signaling pathways. We offer several assays to measure metabolic activity with optimized protocols for 3D cell culture applications, including lactate, glutamine, oxidative stress and dinucleotide detection assays. Common sample preparation allows co-measurement with Glucose-Glo™, Lactate-Glo™, Glutamate-Glo™ and Glutamine/Glutamate-Glo™ Assays from the same sample.
FEATURED PRODUCT
Glucose-Glo™ Assay
Sensitive bioluminescent measure of glucose. Requires only 2–5μl of conditioned cell culture media per time point allowing multiple samples per well for time course studies. Tested on microspheroid culture conditioned media.
Insulin-Mediated Inhibition of Gluconeogenesis of iCell® Hepatocyte Spheroids

NUCLEOTIDE & CO-FACTOR DETECTION
NAD/NADH-Glo™ and NADP/ NADPH-Glo™ Assays
Lytic Assay
Monitor NAD+/NADH or NADP/NADPH ratio with this lytic bioluminescent assay. Lysate is divided and through acid/base reaction measure either oxidized or reduced form of NAD+ or NADH. Slight modification to the standard monolayer protocol (details in poster link below).
Triglyceride Levels in Human Liver Microtissues

For full details, see the technical article linked below.
METABOLITE DETECTION
Lactate-Glo™ Assay
Non-Lytic Assay Option
Monitor lactate changes in 3D cultures using the non-lytic, sensitive bioluminescent assay option. Assay requires only 2–5μl of culture media per time point allowing multiple samples from the same well. Cells may be used for other cell-based assays or nucleic acid isolation.
OXIDATIVE STRESS
GSH/GSSG-Glo™ Assay
Lytic Assay
Monitor changes in GSH/GSSG ratios with sensitive bioluminescent assays. Parallel wells are processed for total GSH + GSSG and GSSG only and results yield the GSH/ GSSG ratio. Minor protocol changes from monolayer (5 min shake) to 3D culture (30 min shake). CellTox™ Green can be used upstream to monitor cytotoxicity.
Monitoring Total Glutathione and Viability in Parallel Wells of HCT 116 Spheroids

Spheroids treated for 48 hours with buthionine sulfoximine prior to measurement of total glutathione with the GSH/GSSG-Glo™ Assay or CellTiter-Glo® 3D Assay. Details in Scientific Poster (link below).
METABOLITE DETECTION
Glutamate-Glo™ Assay
Non-Lytic Assay Option
Monitor glutamate changes in 3D cultures using the non-lytic, sensitive bioluminescent assay option. Assay requires on 2–5μl of culture media per time point allowing multiple samples from the same well. Cells may be used for other cell-based assays or nucleic acid isolation.
METABOLITE DETECTION
Glutamine/Glutamate-Glo™ Assay
Non-Lytic Assay Option
Monitor glutamine and glutamate changes in 3D cultures using the non-lytic, sensitive bioluminescent assay option. Glutamine measurement based on conversion of glutamine to glutamate. Assay requires only 2–5μl of culture media per time point allowing multiple samples from the same well. Cells may be used for other cell-based assays or nucleic acid extraction.
OXIDATIVE STRESS
ROS-Glo™ H2O2 Assay
Non-Lytic Assay Option
Monitor reactive oxygen species through a bioluminescent H2O2 assay. Proluciferin substrate reacts directly with H2O2 and assay components convert the activated prosubstrate to luciferin for bioluminescent detection. Same protocol for monolayer and 3D cultures. Cells may be used for other cell-based assays or nucleic acid extraction.
Response of HepG2 Spheroids to ROS-inducing Menadione

Response of HepG2 spheroids of differing diameters to different levels of menadione. Cells incubated 4 days in ultra-low attachment plates (Corning). H2O2 levels measured with the ROS-Glo™ H2O2 assay and read on a GloMax® Discover Instrument. Details in Application Note (link below).
LIPID METABOLISM
Glycerol-Glo™ Assay
Non-Lytic Assay Option
The Glycerol-Glo™ Assay is a bioluminescent assay for rapid and sensitive measurement of glycerol in a variety of biological samples including cells cultured in 3D. Glycerol is often measured as the product of lipolysis, where it is released from triglycerides. Any processes that result in changes in glycerol concentration, both extracellular and intracellular, can be studied with the Glycerol-Glo™ Assay.
LIPID METABOLISM
Triglyceride-Glo™ Assay
Lytic Assay
The Triglyceride-Glo™ Assay is a bioluminescent assay for rapid and sensitive measurement of triglycerides in cultured cell lysates. The assay is ideal for measuring triglyceride accumulation and clearance in normal and pathological conditions.
LIPID METABOLISM
Cholesterol/Cholesterol Ester-Glo™ Assay
Lytic Assay
The Cholesterol/Cholesterol Ester-Glo™ Assay is a bioluminescent assay for rapid and sensitive method for measuring cholesterol and cholesterol esters in cultured cell lysates and other biological samples, such as cell culture medium. Cholesterol is an essential lipid involved in steroidogenesis, bile acid synthesis, cell signaling and maintenance of membrane structure.
Expression Changes
Cells grown in 3D culture can have differences in gene expression when compared to monolayer cultures. The changes in cell-to-cell contacts and cell-to-matrix contacts will be very different and the gradient of oxygen and nutrients within the culture will also influence gene expression. To analyze differences in gene expression, RNA must be isolated and quantified. Then, specific genes are analyzed by RT-qPCR or cellular changes are monitored through techniques like RNA-seq.
FEATURED PRODUCT
ReliaPrep™ RNA Miniprep Systems
Isolate application-ready total RNA or miRNA from 3D cultures to monitor changes in expression. The ReliaPrep™ RNA Miniprep Systems synergize with non-lytic cellular assays like the RealTime-Glo™ MT Cell Viability Assay.
Extraction of RNA from HEK293 Microtissue Spheroids

Find more products for Expression Changes
Genome Analysis
Understanding genotype-phenotype or phenotype-genotype differences is important, especially when studying primary cells or tumors. You may analyze a genotype and use 3D cell culture to understand the phenotype or you may choose to study the phenotype and then look at the genotype to understand why these changes occur. For genomic DNA analysis, you will need to extract and quantify the DNA. Then you will look at specific genes through qPCR or choose to examine the genome with NGS. If you are working with primary cells, you must be able to differentiate which sample came from which donor—STR profiles are often chosen for that task. The STR profile is also important for confirming that the cells you are working with are truly the cells you desire.
FEATURED PRODUCT
Maxwell® RSC Cultured Cells DNA Kit
Isolate application-ready genomic DNA from 3D cultures for genomic analyses with a simple, walk-away automated protocol using a Maxwell® RSC Instrument. Purified DNA is ready for analysis in about 45 minutes. Isolated gDNA can be used for qPCR-based analyses for gene copy number and biomarkers, STR-based analysis for sample tracking or verification, and sequencing-based analyses.
Concentration of DNA Isolated with the Maxwell® RSC Cultured Cells DNA Kit From Various Starting Amounts of HCT116 Cells
