qPCR and RT-qPCR
Promega real-time PCR (qPCR) and RT-qPCR assays provide a choice of probe-based or dye-based qPCR methods. We also offer XpressAmp® Reagents that are compatible for use with the GoTaq® qPCR and RT-qPCR Systems and enable direct PCR amplification from viral samples without having to perform RNA or DNA extraction.
Both probe and dye-based qPCR assay systems provide sensitive detection for reproducible and earlier quantification of low- and high-copy-number targets over a broad dynamic range, together with resistance to a wide range of PCR inhibitors. They provide sensitive detection on most real time PCR instruments and are amenable to both fast and standard cycling methods.
All real-time qPCR and RT-qPCR systems also include the convenience of room-temperature reaction set-up and easy-to-use master mix formulations.
Shop qPCR and RT-qPCR Products
Filter By
Showing 12 of 12 Products
Need a product modified?
Contact us to discuss custom packaging, formulation and format options for protein analysis products.

Download the MIQE Guidelines Checklist

Download the PCR and Real-Time PCR Brochure

What is Real-Time qPCR?
Real-time quantitative PCR, or qPCR, is a standard method for detecting and quantifying a specific target sequence, or quantifying gene expression levels in a sample. In qPCR, fluorescently labeled probes or nucleic acids, or fluorescent double-stranded DNA-binding dyes are incorporated into the PCR reaction and product formation is monitored in real time following each PCR cycle.
Probe-based qPCR uses a fluorescent primer or probe to detect the amplification product. An advantage of this method is that it allows sequence-specific detection of the target and reduces the likelihood of detecting nonspecific artifacts. Probe-based detection also can accommodate multiplex amplification by using differentially labeled probes. There are several different approaches to probe-based qPCR detection including hydrolysis probes such as TaqMan®, hybridization probes, hairpin probes and labeled primers.
The use of fluorescent DNA-binding dyes, like BRYT® Green or SYBR® Green, is one of the most straightforward qPCR approaches. A dye is added to the reaction, and fluorescence is measured at each PCR cycle. Because fluorescence of these dyes increases dramatically in the presence of double-stranded DNA, DNA synthesis can be monitored as an increase in fluorescent signal.
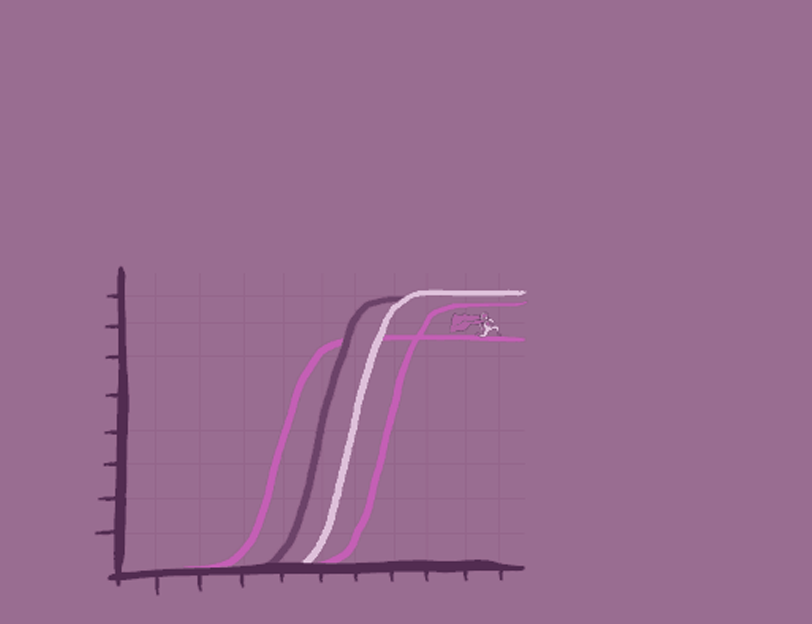
The primary output from a real-time PCR experiment is an amplification curve showing the accumulation of amplified product as fluorescent signal versus the cycle number.
Quantifying the amount of starting material is done using the quantification cycle (Cq, also known as Ct). Cq or Ct (threshold cycle) is the amplification cycle at which the fluorescence signal detected reaches an amplification threshold in a specific well. The amplification threshold is calculated based on the background fluorescence over the baseline region of all the curves in the group and represents the point where the reporter signal is significantly higher than the background levels. Cq is inversely related to the amount of starting template in the reaction and is a relative measure of the concentration of target in the PCR reaction.