The Bioluminescence Advantage
Keith V. Wood
Promega Corporation
Publication Date: 2007
Abstract
Although scientists may draw upon a range of photon-emitting chemistries to serve different experimental purposes, increasingly bioluminescence is becoming a preferred choice for many types of analyses. This article outlines the underlying reasons for the advantage bioluminescence brings to bioanalytical methodologies.
The Case for Bioluminescence
Of the many assay technologies used in life science research, photon emission chemistries are among the most popular due to their inherently high sensitivity and simplicity of use. These chemistries are categorized according to how the energy necessary for photon generation is acquired. Fluorescence, arguably the most widely used chemistry, is also called photoluminescence because it relies on photons as the energy source. Other examples include chemiluminescence, which relies on chemical energy; radioluminescence, which relies on radioactivity (e.g., scintillants); and electroluminescence, which relies on electricity.
Bioluminescence is a form of chemiluminescence where the light-yielding reaction is derived from a naturally catalyzed process, such as the enzymatic reactions found in luminous fireflies, jellyfish, bacteria and others. Bioluminescence chemistries are becoming increasingly popular in a variety of bioanalytical methods because they can deliver 10- to 1,000-fold higher assay sensitivity than fluorescence assays. This greatly increased sensitivity can substantially improve assay performance when applied in complex biological samples. The reason is evident upon examining the mechanism for light production and how it affects implementation into assay methods.(1)
In all luminescent chemistries, energy is absorbed by light-emitting molecules to create excited-state intermediates. Light is produced as this energy is released from the excited states in their transition back to the corresponding ground states (Figure 1). As luminescent chemistries are differentiated by how the excited states are created, this also fundamentally affects how they perform as assay technologies. In fluorescence and bioluminescence, this difference is reflected in their relative signal strengths and signal-to-background ratios.
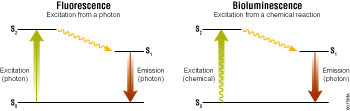
Assays using fluorescence can be relatively bright since the photons generating the excited states may be introduced into a sample at very high rates. However, this high influx of photons can also lead to higher backgrounds largely due to two factors: the burden on the photodetector to discriminate between excitation and emission photons, and interference from other weak fluorophores typically present in biological samples. In fluorogenic assays, the “prefluorophores” added to a sample may be the worst offenders in terms of contributing to high backgrounds.
Although the signal from fluorescent reporters in an assay should be relatively efficient compared to these other processes, it is important to remember that the number of photons being introduced into a sample is typically enormous relative to those produced by the reporters. Thus, the net background in fluorescent assays can often be significant. In contrast, although bioluminescent assays typically yield lower light intensities due to the slower rate by which excited states can be created, assay backgrounds are virtually absent since no photons are introduced into the sample.
This relationship between brightness and background has significant consequences for the suitability of these assay technologies in various applications. Under circumstances where photon detection is limiting, assay backgrounds are largely determined by the capabilities of the detection instrumentation. This is the situation for techniques such as cellular microscopy and flow cytometry, where the optics required for detecting individual cells restrict the light-gathering capacity. In these cases, the brightness of the assay chemistry becomes the primary consideration, and thus fluorescence is usually preferred.
However, with larger samples where optical requirements are simpler and photon detection becomes more efficient, the background inherent in the assay chemistry becomes more important. This is the case for measurements made in individual sample tubes or in multiwell plates (e.g., 96- and 384-well plates), or for image analysis in larger organisms such as mice. In these cases the very low background characteristic of bioluminescent assays can provide substantially greater sensitivity. Moreover, photon detection is typically more efficient for bioluminescence since optical filters or monochromators are not needed for spectral resolution and the photodetectors can usually be positioned closer to the sample.
The result is that bioluminescence can be measured routinely to levels approaching 10–20 moles—less than 10,000 molecules per sample. For a typical biological sample, this is equivalent to a few molecules per cell. In addition, this high sensitivity allows for a very large dynamic range. Most bioluminescent assays are linear for 6 to 8 logs of analyte concentration. Fortunately luminometers, the instruments of choice for quantifying bioluminescence, are capable of operating over these very large ranges.
The utility of bioluminescence as an assay tool is strengthened further by two additional features. First, the photon-emitting capability is naturally evolved within a biological context and thus is broadly compatible with biological systems. Second, the photon emitting molecule, luciferin, is insulated within the structure of the enzyme. This protects the energy transition process and supports optimal quantum yield.
Why Not Fluorescent Proteins?
The most common application of bioluminescence in life science research is as genetic reporters, typically as a means for monitoring the dynamics of genetic transcription. Firefly luciferase is usually the preferred choice—a 61kDa monomeric protein that acts on firefly luciferin, ATP and oxygen to generate yellow-green light (560nm). As such, firefly luciferase serves as an intracellular probe of cellular physiology, and accordingly, should be configured to minimize any impact on normal cellular processes.
One of the best ways to minimize the effects of an intracellular probe is to use it at low concentrations, which can minimize physiological stresses that may be caused by introducing an exogenous component. Beyond this, intracellular probes generally act by coupling or integrating into a specific cellular process, and thus their presence can affect this process. For example, the concentration of the probe may begin to exceed the normal capacity of these processes, potentially leading to diminished or artifactual behaviors. Thus, it is generally recommended to keep the concentration of intracellular probes at a minimum.
This is where the sensitivity of bioluminescence provides a substantial advantage, especially in comparison to fluorescent proteins such as GFP, which are widely used for imaging molecular processes within living cells. Fluorescent proteins have been enormously valuable for this purpose, where the photon emission is bright enough to provide vivid microscopic images. Yet to achieve clearly discernable signals, the fluorescent proteins typically are expressed from strong promoters such as CMV. Although luciferases are less suitable for cellular imaging, they can be used as probes for transcriptional regulation at much lower concentrations than fluorescent reporters.
Another aspect important to the performance of a genetic reporter is the response dynamics. A highly stable reporter tends to resist changes in its intracellular concentration in association with changes in the transcriptional rate. Consequently the reporter may not accurately represent the underlying transcriptional dynamics in living cells. The dynamic response can be improved by incorporating protein degradation signals into the reporter (e.g., PEST sequences), but these also reduce intracellular accumulation of the reporter and can reduce assay sensitivity. Nonetheless, this strategy works well for luciferase reporters because of the high sensitivity inherent in luminescent assays. In contrast, fluorescent proteins are generally highly stable, and addition of degradation signals can restrict assay sensitivity.
The dynamic capabilities of luciferase reporters can be applied broadly to investigations of cellular physiology. In the example shown in Figure 2, the native β2-adrenergic receptor in HEK293 cells is stimulated by adding an agonist, isoproterenol. In this Gαs-coupled receptor, stimulation at the membrane is transmitted within the cell by increasing intracellular cAMP, which in turn activates gene expression through a cAMP-response element (CRE). Receptor activity can be monitored by coupling a CRE upstream of the luciferase reporter gene.
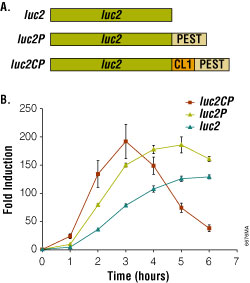
Figure 2. Bioluminescent reporter assay of the beta2-adrenergic receptor. Degradation sequences incorporated into the luciferase reporter increase both the rate and relative magnitude of response. Panel A. luc2, firefly luciferase; luc2P, firefly luciferase with PEST sequence (-Pro-Glu-Ser-Thr); luc2CP, firefly luciferase with CL1 and PEST sequences. Panel B. HEK293 cells stably expressing luciferase reporter genes coupled to a cAMP-response element (CRE). Endogenous β2-adrenergic receptors were induced with 1μM isoproterenol/ 100μM RO-20-1724.
Using the optimized pGL4 Luciferase Reporter Vectors, reporter expression is stimulated 200-fold upon addition of the agonist (Figure 2). Moreover, the rate and magnitude of this response is improved by adding degradation sequences. The luciferase genes in the pGL4 Vectors are codon optimized for use in mammalian cells to ensure maximum expression efficiency. In addition, consensus binding sites for transcription factors have also been removed to minimize the potential for unintended host interactions with the vector.
Broader Applicability for Bioluminescence
Light intensity generated by bioluminescent reactions is dependent on the concentrations of the component reactants, and through chemical couplings to those components, can be associated with a variety of molecular processes. Typically, the concentrations of all components in an assay are held constant except one, which is allowed to vary in connection with the process of interest. For reporter gene assays, the variable component is the enzyme itself coupled to gene expression. But the substrates, such as ATP or luciferin, may also serve as the variable components (Figure 3).
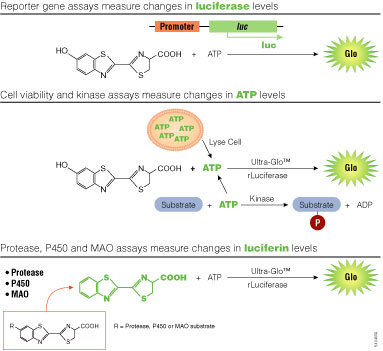
Figure 3. Assay strategies using bioluminescence. luc, firefly luciferase gene; Ultra-Glo™ rLuciferase, a highly stabilized variant of firefly luciferase; P450, cytochrome p450; MAO, monoamine oxidase.
Luciferin may be incorporated into assay designs in a manner similar to fluorogenic assays. By attaching a modifying group, the luciferin becomes unavailable to the luminescent reaction until the modifier is removed through some biochemical process. For example, the luciferin derivative, Asp-Glu-Val-Asp-6′-aminoluciferin, cannot support luminescence until the tetrapeptide sequence is cleaved by the caspase-3 protease. By this means, the luciferin derivative provides a sensitive assay for detecting apoptosis. Analogous fluorescent assays for apoptosis may be made by coupling the tetrapeptide to appropriate fluorophores such as rhodamine. However, when compared under similar conditions, the bioluminescent assay is nearly 100-fold more sensitive (Figure 4). The increased sensitivity is due to greatly reduced background, which also provides an extended linear range.
This approach can be applied to other proteases including caspase-8, caspase-9, DPPIV, calpain and the proteasome with similar results. It may also be used to measure CYP450 activity, monoamine oxidase activity, or a variety of other enzymatic cleavage reactions.
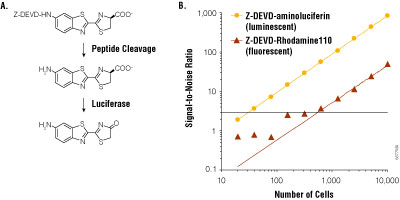
Figure 4. Bioluminescent and fluorescent measurements of caspase-3/7 activity. Panel A. Assay schema of caspase cleavage on Z-DEVD-aminoluciferin. Panel B. Assays were performed on Jurkat cells treated with anti-FAS and serially diluted into 96-well plates. Measurements were made 1 hour after reagent addition, and background was measured from wells not containing cells. Signal-to-noise ratio was calculated as mean measurement minus mean background, divided by the standard deviation of the background. Panel B reprinted with permission from O’Brien, M.A. et al. (2005) J. Biomol. Screen. 10, 137–48.
Detection of ATP is one of the oldest applications for bioluminescence and continues to gain popularity as a rapid assay for cell viability. The assay for mammalian cells is over 100-fold more sensitive than commonly used tetrazolium-based assays and requires only ~5 minutes. Similar bioluminescent assays for bacterial viability are sensitive to ~100 cells, depending on the cell type. Bioluminescent assays for ATP may also be used to measure enzymes that consume ATP, most notably kinases. This approach has the advantage of being nearly universal for kinase activity, regardless of whether the phosphate acceptor is a protein, lipid or polysaccharide.
A recent strategy combined luciferase and protein kinase A (PKA) in an assay for cAMP. cAMP released from cultured cells regulates PKA activity in the assay reagent, which in turn acts on ATP needed for the bioluminescent reaction. The combination of these two reactions allows a rapid and sensitive assay of Gαs-coupled receptors or phosphodiesterase activity in cultured cells.
A Unique Beacon for Biology
Bioluminescent chemistries find their greatest potential within the enormous complexity characteristic of living systems. The reaction constituents may exist in vanishingly small amounts enveloped within an intricate biochemical milieu, and yet they deliver clear and quantitative signals. This capability is achieved by being the only photonic players in an otherwise dark environment. The only photons present are created by this unique chemistry, imparting an essential singularity within the diverse environment.
Consequently, among photon-emitting chemistries, bioluminescence stands alone as a beacon for revealing the secrets of molecular processes.
References
- Fan, F. and Wood, K. (2007) Bioluminescent assays for high-throughput screening. Assay Drug Dev. Technol. 5, 127–36.
Learn more about Cell Health Assays.