Recombinant RNasin® Ribonuclease Inhibitor Protects RNA Without Affecting Analyses
Mark Bratz, Brad Hook and Trista Schagat
Promega Corporation
Publication Date: September 2012
Abstract
Protecting purified RNA samples from ribonuclease (RNase) contamination is essential. Ribonuclease inhibitors are effective at protecting RNA quality but must not inhibit downstream assays. Recombinant RNasin® Ribonuclease Inhibitor and many other commercially available inhibitors are effective at protecting RNA from RNase, but some inhibitors can interfere with downstream applications. Here we report on our evaluation of the effects of Recombinant RNasin® Ribonuclease Inhibitor on four downstream analyses after titrating various amounts of the inhibitor into newly purified RNA samples and again after the samples were stored frozen at –20°C for 6 weeks with repeated freeze-thaw cycles. Recombinant RNasin® Ribonuclease Inhibitor protected RNA integrity and did not affect the downstream analyses, even when the sample contained more than 9% Recombinant RNasin® Ribonuclease Inhibitor.
Introduction
RNA degradation through RNase contamination causes many problems in downstream applications. RNA integrity is extremely important in gene expression studies where degradation can lead to misleading amounts of transcripts (1,2). Preventing this degradation is essential and very simple using commercially available RNase inhibitors; however, some commercially available inhibitors can interfere with downstream applications and analyses after RNA isolation (1,2).
The addition of as little as 1µl of Recombinant RNasin® Ribonuclease Inhibitor inhibits RNases and prevents degradation. Recombinant RNasin® Ribonuclease Inhibitor (Cat.# N2515) effectively protects RNA from RNase contamination in the laboratory (1) over a wide range of temperatures (4, 25, 37 and 50°C) and pH (5–8). Recombinant RNasin® Ribonuclease Inhibitor also offers a distinct advantage over native RNasin® Inhibitor because it is not isolated from human tissue, minimizing the risk of incorporating human nucleic acids into the assay (3).
Upstream gene expression analysis and RNA experiments commonly involve a workflow that includes RNA isolation, quantification, analysis of purity and integrity, and downstream applications such as RT-qPCR. Two common techniques used to quantitate RNA are absorbance using spectrophotometry and fluorescence using an RNA-specific binding dye. Spectrophotometry, using the NanoDrop® instrument, for example, is a simple, quick technique for quantification of RNA; however, it is not specific for RNA and results in inflated concentrations due to contaminants such as protein, DNA and organic compounds. The QuantiFluor® RNA System (Cat.# E3310) is a fluorescent dye-based system that allows fast quantitation over a wide range of RNA concentrations with a lower limit of detection than the NanoDrop® instrument, making it an excellent quantitation system for samples with low RNA concentrations. The Agilent Bioanalyzer, which generates an RNA Integrity Number (RIN) to measure RNA integrity, also has become a popular method for analyzing RNA quality.
All techniques in this RNA workflow require intact RNA. Ideal RNase inhibitors protect against RNases but must not impair downstream assays. In this report, Recombinant RNasin® Ribonuclease Inhibitor was added to RNA samples collected using the Maxwell® 16 LEV simplyRNA Blood Kit. These samples were then tested using the NanoDrop® instrument, Quantifluor® RNA System, Agilent Bioanalzyer and RT-qPCR before and after storage to determine the effect of Recombinant RNasin® Ribonuclease Inhibitor on downstream applications.
Methods
Sample Preparation
Sixteen RNA samples were obtained from a single human donor following the protocol described in the Maxwell® 16 LEV simplyRNA Blood Kit Technical Manual #TM372. The samples were pooled, mixed and redistributed in 50μl aliquots. Recombinant RNasin® Ribonuclease Inhibitor was added to the samples at volumes of 0, 0.5, 1.0, 2.0, or 5.0μl (40U/μl), and nuclease-free water was added to a final volume of 55μl. Four analyses were completed on each of the samples to assess the effects of Recombinant RNasin® Ribonuclease Inhibitor. The RNA was then frozen at –20°C. Between initial and final analysis of the samples, the RNA was thawed, mixed by pipette, and then placed back into storage eight times for each sample over a six-week period to simulate multiple uses. After the samples were stored for six weeks, the analyses were performed again.
QuantiFluor® RNA System
RNA quantitation was performed as described in the QuantiFluor® Dye Systems Technical Manual, #TM346. RNA samples, diluted 1:50 in 1X TE buffer, were mixed with QuantiFluor® RNA dye and incubated, protected from light, for 5 minutes at room temperature. Fluorescence was measured using the QuantiFluor® ST handheld instrument.
RNA 6000 Nano Kit
RNA quantitation and integrity also was assayed using the RNA 6000 Nano Kit as described in Agilent RNA 6000 Nano Kit Quick Start Guide, #G2938-90034. One microliter of sample was used for analysis.
GoTaq® 1-Step RT-qPCR
RT-qPCR using HPRT1-specific primers was performed using GoTaq® 1-Step RT-qPCR System (Cat.# A6020, TM355). RNA samples were diluted 1:20 in nuclease-free water, and a 10μl aliquot of this dilution was added to the reactions, resulting in approximately 50ng RNA (0.5µl original eluate) in each 50μl reaction.
Results
RNA samples, isolated using the Maxwell® 16 LEV simplyRNA Blood kit, were pooled, and aliquots of 50μl were removed and treated with various amounts of Recombinant RNasin® Ribonuclease Inhibitor. The samples were analyzed using spectrophotometry on the NanoDrop® instrument, by fluorescent-based quantitation with the QuantiFluor® RNA System, on the Agilent Bioanalyzer, and used in RT-qPCR analyses. The samples were then frozen for 6 weeks and subjected to 8 freeze-thaw cycles. All analyses were repeated post-storage.
Using the NanoDrop® instrument to determine concentration and purity, no significant difference in concentration with 0–2µl of Recombinant RNasin® Ribonuclease Inhibitor added was detected (Table 1, “Initial”), and a slight (<5%) increase in concentration at the highest Recombinant RNasin® Ribonuclease Inhibitor addition (5µl) was observed. The A260/A230 and A260/A280 ratios decrease with increased Recombinant RNasin® Ribonuclease Inhibitor addition, but all ratios, except those containing 5.0μl of the inhibitor, are above 1.9, indicating high purity. The small ratio decreases in the 5μl-addition sample are attributed to higher levels of protein and glycerol, the latter present in the storage buffer. The same samples were tested after 6 weeks at –20°C with multiple freeze-thaw cycles. The concentrations and purity ratios were similar to the initial experiment (Table 1, “Final”).
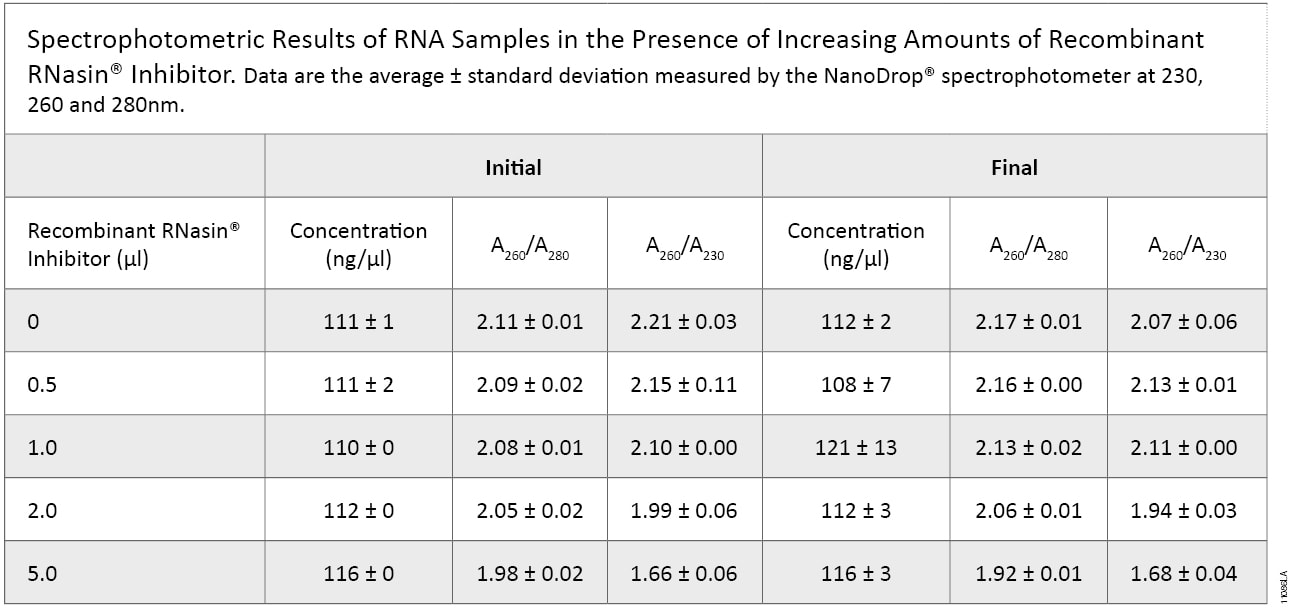
Table 1. Determination of concentration and purity of RNA samples before and after storage. We used the NanoDrop® instrument to determine concentration and purity of samples with increasing amounts of Recombinant RNasin® Ribonuclease Inhibitor before (Initial) and after (Final) six weeks of storage with repeated freeze-thaw cycles. No significant changes were observed in RNA concentration among the samples before and after storage.
Another common method for RNA quantitation is the QuantiFluor® RNA System, which uses a fluorescent RNA-binding dye to quantitate RNA in samples. The increase in Recombinant RNasin® Ribonuclease Inhibitor did not lead to a difference between the samples (Table 2). A difference in concentrations between the NanoDrop® instrument and the QuantiFluor® RNA Systems is expected due to differences in the method of detection and sensitivity levels between an absorbance measurement (NanoDrop® instrument) and a fluorescent measurement (QuantiFluor® RNA Systems).
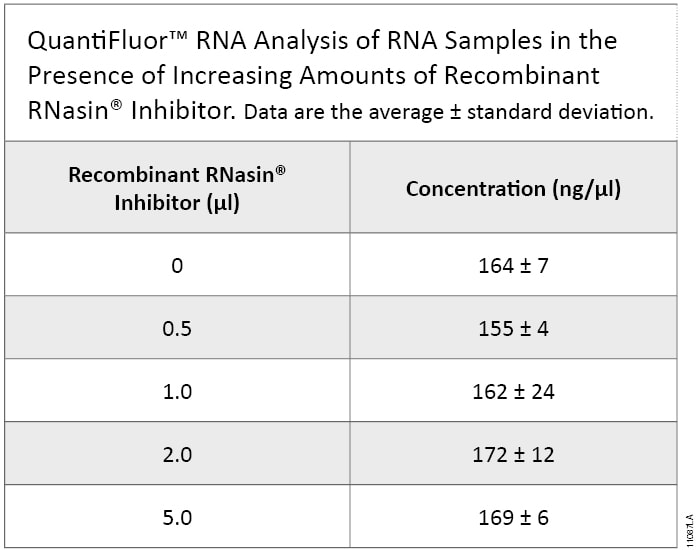
Table 2. Quantitation of RNA samples containing increasing amounts of Recombinant RNasin® Inhibitor. Quantitation of RNA using QuantiFluor® RNA binding dye. Increasing amounts of Recombinant RNasin® Ribonuclease Inhibitor did not lead to a significant difference among RNA samples tested.
A bioanalyzer is commonly used to determine the integrity of the RNA. We used the Agilent RNA 6000 Nano Kit and a 2100 Bioanalyzer to determine the RNA Integrity Numbers (RIN) for our samples with and without Recombinant RNasin® Inhibitor. The average RINs for all sample types were ≥8 indicating highly intact RNA (Figure 1). Adding 2–5µl Recombinant RNasin® Inhibitor tended to give slightly higher RINs than adding 0–1.0µl of inhibitor, especially after six weeks of storage with repeated freeze-thaw cycles (final RIN). Gel images for both the initial and final tests illustrate that the samples have very little degradation (Figure 2).
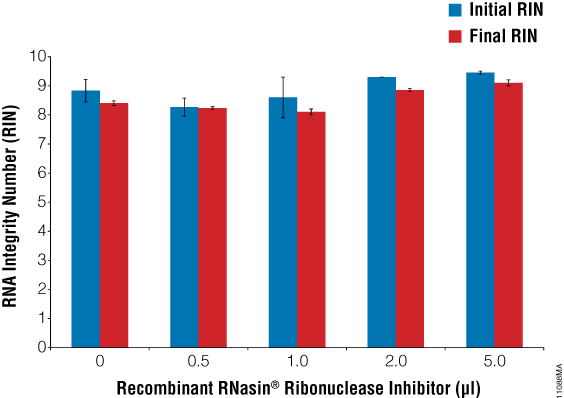
Figure 1. Initial and final RINs for the samples illustrating no significant change in RIN over the six-week storage period. We used the Agilent RNA 6000 Nano Kit and a 2100 Bioanalyzer to determine RNA Integrity Numbers (RIN) for samples with Recombinant RNasin® Inhibitor before (initial) and after (final) six weeks of storage with repeated freeze-thaw cycles. The average RIN for all samples was greater than or equal to 8, indicating intact RNA.
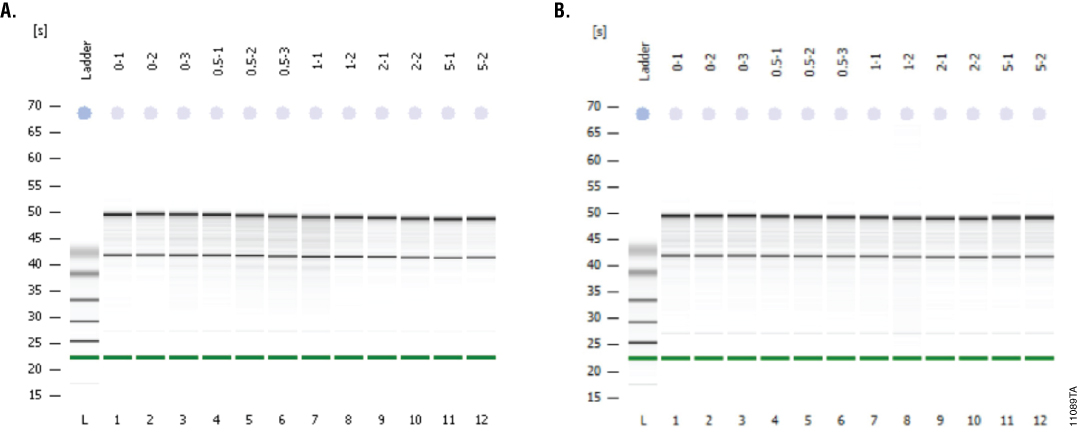
Figure 2. Gel images of RNA samples created by the Agilent Bioanalyzer system. We analyzed samples with varying concentrations of Recombinant RNasin® Ribonuclease Inhibitor before and after six weeks of storage with repeated freeze-thaw cycles. Panel A represents samples before storage. Panel B represents samples after the six-week period. The gel models demonstrate that the samples show little RNA degradation.
Purified RNA is commonly used for RT-qPCR. We tested our samples for the ability to amplify a gene using the GoTaq® 1-Step RT-qPCR System. Using HRPT-specific primers, all samples yielded similar Cq values regardless of the amount of Recombinant RNasin® Inhibitor added (see Table 3). Amplification and melt curves were similar between the initial experiment and the final experiment with samples stored for six weeks, indicating little change in amplifiable RNA (Figure 3).
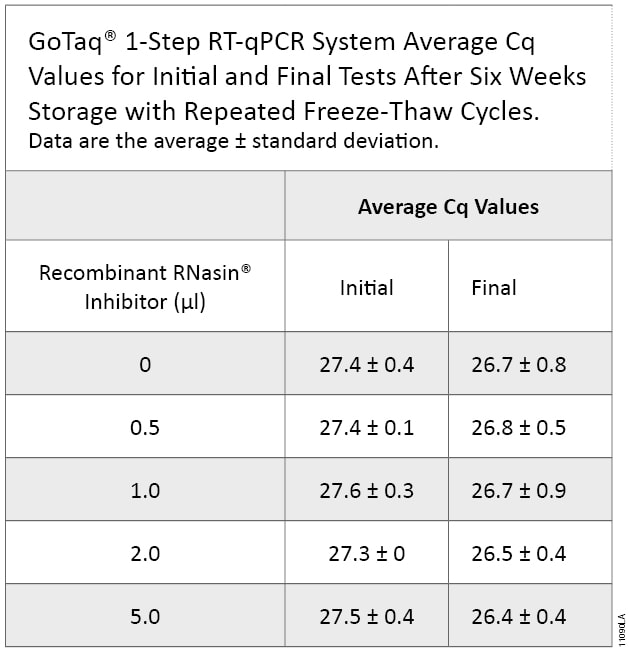
Table 3. Performance of RNA samples treated with Recombinant RNasin® Ribonuclease Inhibitor in RT-qPCR. We used RNA samples treated with increasing amounts of Recombinant RNasin® Ribonuclease Inhibitor to amplify a gene using the GoTaq® 1-Step RT-qPCR System. Average Cq values were compared among samples before (Initial) and after (Final) the six-week storage period with repeated freeze-thaw cycles. All samples yielded similar Cq values.
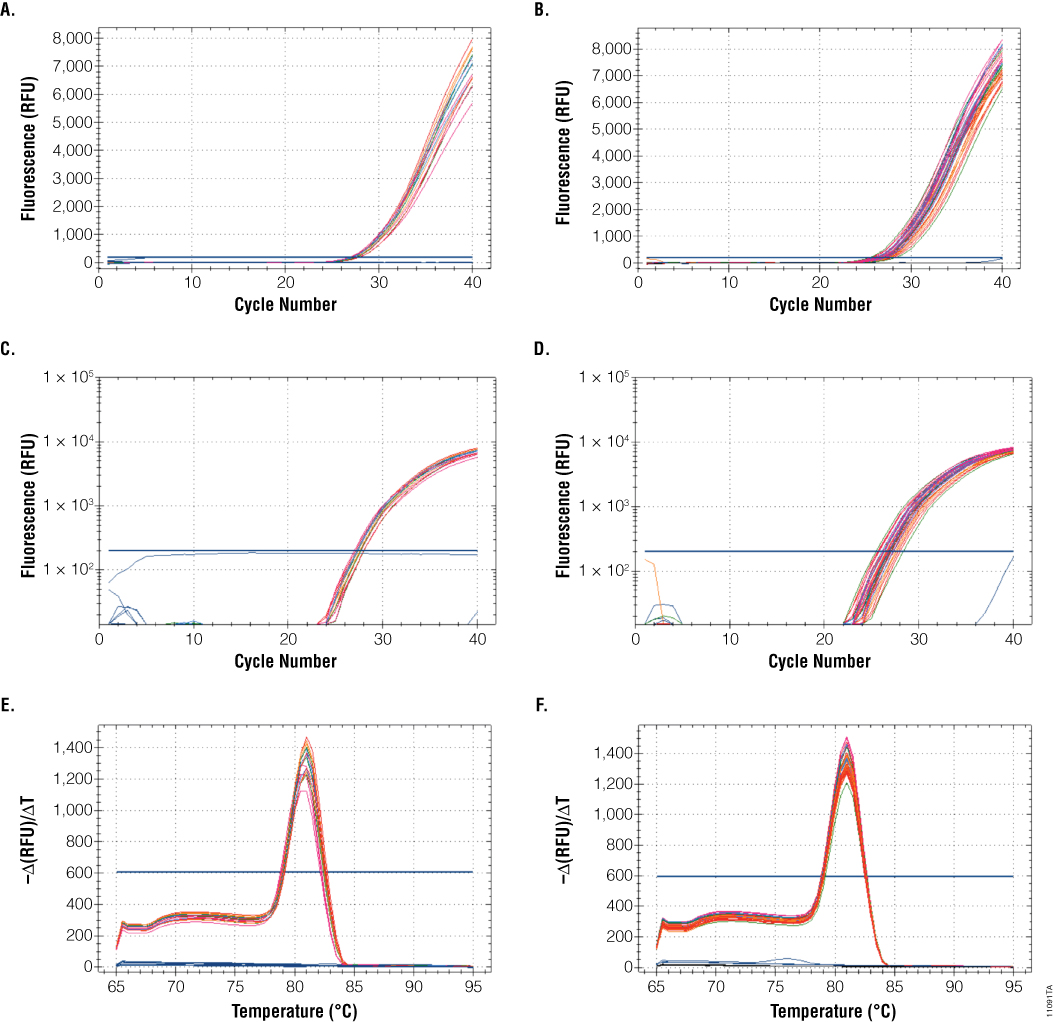
Figure 3. GoTaq® 1-Step RT-qPCR amplification and melt curves for the RNA samples before and after six weeks of storage with repeated freeze-thaw cycles. We tested RNA samples, before and after the six-week storage period, for the ability to amplify a gene using the GoTaq® 1-Step RT-qPCR System with HRPT-specific primers. Panels A and B show linear amplification curves; Panels C and D show log-scale amplification curves; Panels E and F show DNA melt curves. Panels A,C and E are the initial samples (before storage); Panels B, D and F are the final samples (after storage). Red, 0µl RNasin® Inhibitor; Orange, 0.5µl RNasin® Inhibitor; Green, 1.0µl RNasin® Inhibitor; Blue, 2.0µ RNasin® Inhibitor; Pink, 5.0µl RNasin® Inhibitor.
Summary
RNase contamination can occur during RNA purification and anytime during downstream processing, from sample handling, addition of water or buffers, etc. (2). We found that multiple additions of Recombinant RNasin® Ribonuclease Inhibitor to purified RNA samples does not adversely affect downstream applications, making it a quick, reliable and cost-effective way to protect your RNA samples from RNase-related degradation.
References
- Hendricksen, A., Hook, B. and Schagat, T. (2010) RNase contamination happens; Recombinant RNasin® Inhibitor can safeguard your samples. Promega PubHub.
- Hook, B., Hendricksen, A. and Schagat, T. (2011) RNase inhibition: A head-to-head comparison of RNasin to RNaseOUT. Promega PubHub.
- Promega Corporation (1989) Recombinant RNasin® Ribonuclease Inhibitor. Promega Notes 19, 1–2.