Quantifying dsDNA and ssDNA Isolated by Manual and Automated Methods Using a Fluorescent Nucleic Acid Dye
Promega Corporation
Publication Date: August 2013
Abstract
The DNA IQ™ System is designed specifically for forensic and paternity labs to isolate DNA for short tandem repeat (STR) analysis and, due to the limited amounts of DNA found in many forensic samples, yields DNA that cannot be accurately quantitated by traditional UV-absorbance methods. To find an alternative to quantitative PCR, which is a more sensitive method commonly used for quantitating trace DNA, we tested the ability of fluorescent dye-based QuantiFluor® dsDNA and ssDNA Systems to quantitate DNA isolated using the DNA IQ™ System. Here we examined samples that had undergone various preprocessing protocols prior to DNA binding and found that the preprocessing temperature and buffer used dictate whether a single-stranded DNA or double-stranded DNA fluorescent dye system should be used for quantitation.
Introduction
The DNA IQ™ System is a DNA isolation system designed specifically for forensic laboratories to extract DNA from a variety of sample types, including liquid samples and liquid stains on solid supports. Often these forensic samples contain very low or limiting amounts of DNA. For forensic laboratories, it can be important to quantitate accurately the amount of DNA present in these samples as downstream applications such as STR amplification require specific amounts of template for proper analysis and identification of the sample source. Fluorescent DNA-binding dye-based systems often are used to quantitate DNA purified from samples with low DNA concentrations as the systems are highly sensitive and can detect picogram amounts of DNA, significantly less than traditional UV absorbance analysis. Thus, for forensic laboratories these fluorescence dye systems represent a sensitive, fast and inexpensive alternative to qPCR for DNA quantitation from trace samples.
The DNA IQ™ System can be preceded by different preprocessing protocols to enhance DNA recovery, depending on the sample type. These preprocessing protocols vary in the buffers and temperatures used for DNA extraction, which can affect the DNA form (i.e., single-stranded [ssDNA] or double-stranded [dsDNA]) that is ultimately purified. Here we determine the appropriate fluorescent ssDNA- or dsDNA-binding dye-based system to quantitate DNA isolated following various DNA IQ™ System preprocessing protocols.
Methods
Blood was collected from a single donor in a K2EDTA Vacutainer® tube (BD Biosciences). Ten microliters of liquid blood and 10µl of blood spotted on a cotton swab were processed according to the DNA IQ™ Casework Pro Kit for Maxwell® 16 Technical Manual #TM332. Ten microliters of blood spotted on cotton swabs also was processed manually according to the DNA IQ™ Systems—Small Sample Casework Protocol Technical Bulletin #TB296, and 10µl of liquid blood samples were processed manually according to DNA IQ™ System—Database Protocol Technical Bulletin #TB297. The preprocessing requirements for each protocol are shown in Table 1. DNA samples purified using the Maxwell® 16 System were eluted in 50µl of Elution Buffer, and DNA samples isolated using the manual protocol were eluted in 100µl of Elution Buffer.
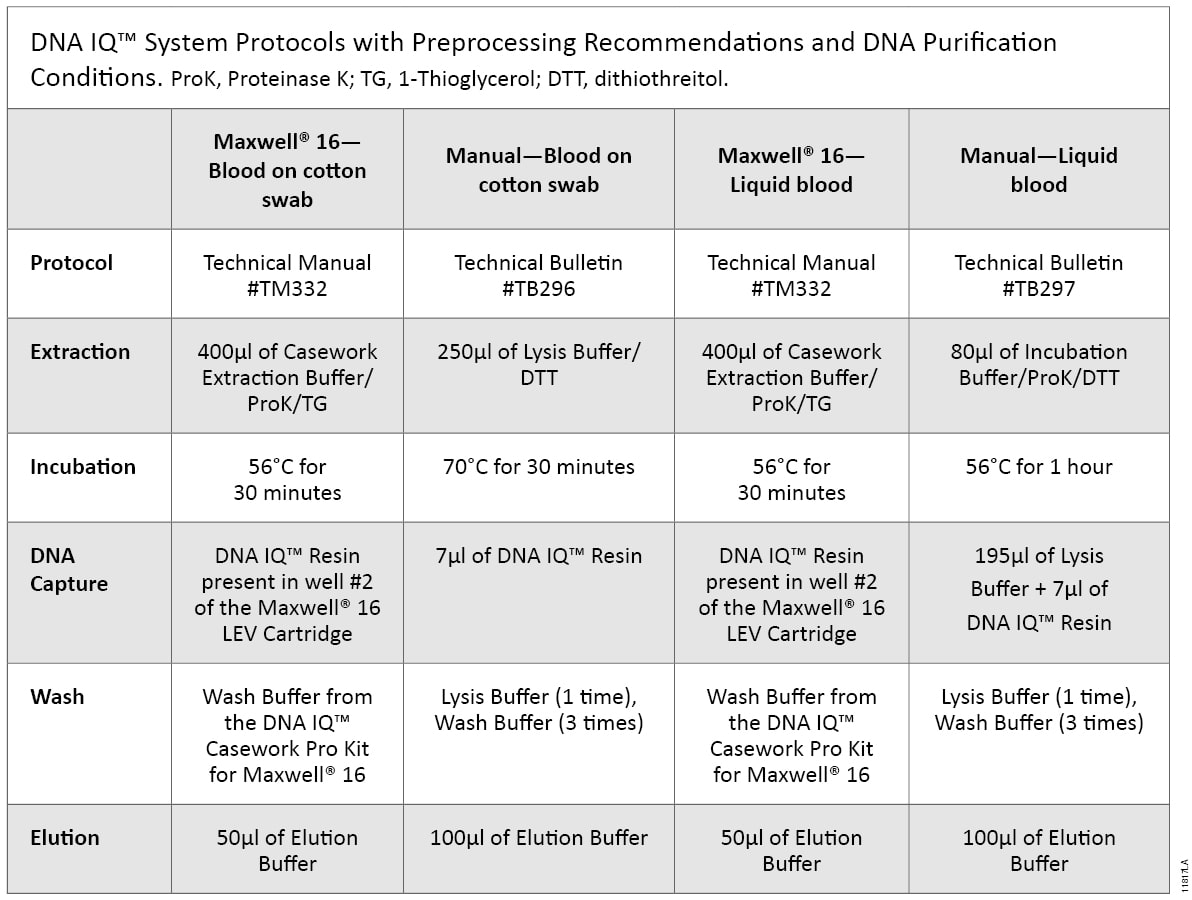
Table 1. DNA IQ™ System Protocols with Preprocessing Recommendations and DNA Purification Conditions. ProK, Proteinase K; TG, 1-Thioglycerol; DTT, dithiothreitol
Purified DNA samples were quantitated using the following fluorescent dye-based chemistries, and fluorescence was measured using the GloMax®-Multi+ Detection System:
- QuantiFluor® dsDNA System (Cat.# E2670)
- QuantiFluor® ssDNA System (Cat.# E3190)
Prior to quantitation using the QuantiFluor® ssDNA System, samples were first digested with Shrimp DNase (USB Cat.# 78314) as described in the QuantiFluor® ssDNA System Technical Manual #TM376. Because the QuantiFluor® ssDNA Dye chemistry will also detect dsDNA, this step is important to accurately quantify ssDNA. Shrimp DNase activity is 5,000 times more specific for dsDNA than ssDNA so it efficiently degrades dsDNA and leaves ssDNA intact. The QuantiFluor® dsDNA System is specific for dsDNA thus no pretreatment is necessary for quantitation.
Results
Figure 1 shows the dsDNA and ssDNA concentration obtained from each sample type using the QuantiFluor® dsDNA and ssDNA Systems. Samples processed using the Maxwell® 16 System yielded higher DNA concentrations because DNA was eluted in 50µl of Elution Buffer; samples processed using manual protocols were eluted with 100µl of Elution Buffer and so the DNA was more dilute. A ratio was calculated to compare the concentrations for each DNA form (Table 2). Samples of blood spotted on cotton swabs and processed manually (Manual–Blood on Cotton Swab) yielded two to three times as much ssDNA as dsDNA, while the inverse was true for the other sample types. Manual processing of blood on solid supports requires a 70°C incubation in the presence of Lysis Buffer and DTT (heated lysis protocol), while the other protocols require a 56°C incubation in the presence of Casework Extraction Buffer or Incubation Buffer with Proteinase K and 1-Thioglycerol. Thus, the heated lysis protocol results in a higher proportion of ssDNA than protocols requiring heating in Casework Extraction Buffer or Incubation Buffer.
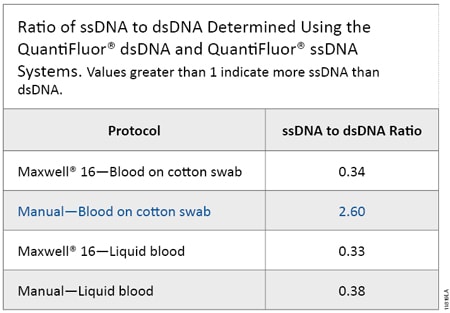
Table 2. Ratio of ssDNA to dsDNA Determined Using the QuantiFluor® dsDNA and QuantiFluor® ssDNA Systems. Values greater than 1 indicate more ssDNA than dsDNA.
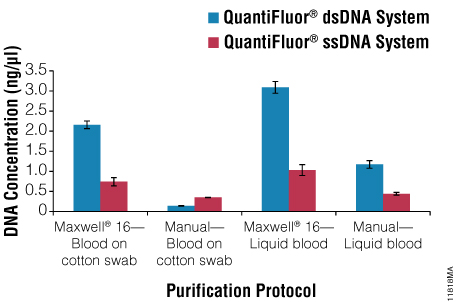
Figure 1. The QuantiFluor® dsDNA System and QuantiFluor® ssDNA System (with shrimp DNase digestion) were used to determine DNA concentrations from automated (DNA IQ™ Casework Pro Kit for Maxwell® 16) or manual protocols for DNA purification from liquid blood or blood on a cotton swab. Samples processed using the Maxwell® 16 System were eluted in 50µl of Elution Buffer, and samples processed manually were eluted in 100µl of Elution Buffer. Error bars represent ± 1 standard deviation. N = 4.
Conclusion
The QuantiFluor® dsDNA and ssDNA Systems can be used to quantitate purified DNA from manual and automated DNA IQ™ protocols. These protocols result in the isolation of dsDNA as well as ssDNA. However, the relative amounts of each form of DNA isolated depend on the preprocessing protocol used. For the DNA IQ™ protocol that includes incubation at 70°C in Lysis Buffer (heated lysis), a higher proportion of ssDNA was observed, and we recommend using the QuantiFluor® ssDNA System. DNA IQ™ protocols that include incubation at 56°C in Incubation Buffer or Casework Extraction Buffer resulted in a higher proportion of dsDNA; therefore, we recommend using the QuantiFluor® dsDNA System. For the most accurate DNA quantitation, samples should be quantitated first with the QuantiFluor® dsDNA System to specifically measure dsDNA, treated with shrimp DNase and then quantitated using the QuantiFluor™ ssDNA System to specifically measure ssDNA.
Learn more about the QuantiFluor® ssDNA System