Reliable Automated Cytotoxicity Profiling for Chemistry-Focused Laboratories by Combining Multiplexed Cytotoxicity Assays with the CyBi®-Felix Liquid Handling Instrument
Matt Butzler1, Tracy Worzella1, Marlon Hungriano2, Felix Osorio2, Ian Hanegraaff2 and Cris Cowan1
1Promega Corporation, Madison, WI;
2Analytik Jena US, CyBio Product Line
Publication Date: December 2014
Abstract
Understanding the mechanism of toxicity of compounds is important to basic research as well as drug development programs. We developed an automated workflow for determining the mechanism of chemical-mediated cytotoxicity. The mode of cytotoxicity is revealed by multiplexing CellTox™ Green Express Cytotoxicity Assay with either CellTiter-Glo® 2.0 or Caspase-Glo® 3/7 Assay. When used together these assays help distinguish among the common mechanisms of chemical action on cellular health: necrosis, apoptosis, or growth arrest. We performed the profiling experiments using the CyBi®-FeliX liquid handler and GloMax® Discover Multimode Reader instruments, which are simple options for the novice automation user. In addition, reagents included frozen, thaw-and-use HepG2 cells, which removed the burden of growing and maintaining cells in culture. Our automated approach renders cytotoxicity profiling into an easier and straightforward experience for chemistry-focused labs.
Introduction
In vitro cell-based assays are a convenient and cost-effective way of evaluating test compound toxicity especially in drug discovery research programs. A typical workflow involves titration of the test compound, application to cells, and incubation for several days. We used the CyBi®-FeliX (Analytik Jena US, CyBio Product Line) instrument as a stand-alone, fully automated system to perform all cytotoxicity assay-specific liquid handling tasks. .jpg) Effects of the test compounds were quantified through the application of reagents designed to measure biomarkers of interest. In general terms, a compound or treatment that elicits a cytotoxic effect does so by causing dramatic morphological changes, adversely affecting replication rate or leading to a reduction in viable cell count. Information gained from in vitro methods is an important antecedent to flag potential toxic liabilities.
Effects of the test compounds were quantified through the application of reagents designed to measure biomarkers of interest. In general terms, a compound or treatment that elicits a cytotoxic effect does so by causing dramatic morphological changes, adversely affecting replication rate or leading to a reduction in viable cell count. Information gained from in vitro methods is an important antecedent to flag potential toxic liabilities.
In mechanistic cytotoxicity profiling, unique biomarkers are used to profile the health status of cells. In this report, we used a panel of three assays to assess biomarkers associated with changes in membrane integrity (cytotoxicity), cell number and apoptosis. When used together, these biomarkers can help distinguish populations of cells that are necrotic, apoptotic or growth arrested. The assays used for this work are described in Table 1.
Table 1. Description of the Promega Cell Health Assays Used for Automated Cytotoxicity Profiling in This Study
| Assay | Determination | Description |
| CellTox™ Green Express Cytotoxicity Assay | Cytotoxicity or compromised cell membrane | Nonlytic reagent comprised of a DNA binding dye that is permeable to cells with a compromised cell membrane. The dye becomes highly fluorescent when bound to DNA, and the fluorescent signal is proportional to the number of dead cells in a sample. The dye is conveniently added to the cell suspension at the time of cell plating allowing cytotoxicity to be monitored over time. The nonlytic reagent can be multiplexed with other spectrally distinct fluorescent dyes or luminescent assays TM375 |
| CellTiter-Glo® 2.0 Assay | Cell number or viability | Lytic reagent that quantifies the ATP present in a cell population using a luminescent firefly luciferase reaction. Luminescent signal is proportional to ATP content and indicates the number of viable cells. TM403 |
| Caspase-Glo® 3/7 Assay | Apoptosis | Lytic luminescent assay that quantifies caspase-3/7 activity, enzymes active in cells undergoing late-stage apoptosis. A DEVD-proluciferin substrate is cleaved by caspase-3/7 to generate free luciferin that is quantified in a firefly luciferase reaction. Luminescent signal is proportional to caspase-3/7 activity and indicates cells undergoing apoptosis. TB323 |
We multiplexed the CellTox™ Green Assay with either the CellTiter-Glo® 2.0 or Caspase-Glo® 3/7 Assay to minimize sample processing and conserve cell samples. The detection chemistries of CellTiter-Glo® 2.0 and Caspase-Glo® 3/7 are incompatible. Consequently, we assembled duplicate samples to assess cell viability and apoptosis separately.
Frozen, thaw-and-use cells offer a convenient option over cells from continuous culture for this profiling workflow. Such thaw-and-use cells are prepared and plated immediately after thawing, eliminating any prior cell culture activities. Passage number and confluency can impact biological performance of cultured cells, and the use of thaw-and-use cells circumvents these issues.
The profiling workflow is streamlined by using laboratory automation for all experimental steps. Here we automated serial titration and application of test compound(s) to an assay plate, as well as addition of cells and reagents. Incorporating automation ensures consistent experimental setup by eliminating user-to-user variability in pipetting and enhances laboratory efficiency. For this application we used Frozen Instant HepG2 Cells (Cell Culture Service), a human liver cell line, a CyBi®-FeliX instrument for all liquid handling steps and a GloMax® Discover multimode detection system (Promega) for detection of fluorescent and luminescent assay signals.
Adopting this automated profiling process requires less time and attention devoted to experimental steps. The automated process enables testing of up to eight compounds in quadruplicate in 384-well microplate format. Duplicate plates can be assembled when time course studies are desired.
Materials and Methods
Materials
• DMSO (Sigma Aldrich, Cat.# D8418)
• methotrexate (Sigma Aldrich, Cat.# M9929), 10mM in DMSO
• panobinostat (Selleckchem, Cat.# S-1030), 10mM in DMSO
• tamoxifen Citrate (Sigma Aldrich, T9262), 10mM in DMSO
• nocodazole (Sigma Aldrich, Cat.# M1404), 10mM in DMSO
• bortezomib (LC Laboratories, Cat.# B-1408), 10mM in DMSO
• cisplatin (Sigma Aldrich, Cat.# P4394), 10mM in DMSO
• staurosporine (EMD Millipore, Cat.# 569397), 10mM in DMSO
• terfenadine (Sigma Aldrich, Cat.# T9652), 10mM in DMSO
• CellTiter-Glo® 2.0 Assay (Promega, Cat.# G9242)
• CellTox™ Green Express Cytotoxicity Assay (Promega, Cat.# G8731)
• Caspase-Glo® 3/7 Assay (Promega, Cat.# G8093)
• cellgro DMEM, 1X with 4.5 g/ glucose, L-glutamine & sodium pyruvate (Corning Cat.# 10-012-CV) supplemented with 10% FBS
• EMEM with L-Glutamine (ATCC Cat.# 30-2003) supplemented with 10% FBS and 1X Penicillin/Streptomycin
• EMEM with L-Glutamine (ATCC Cat.# 30-2003) supplemented with 10% FBS and 5% DMSO
• Costar® v-bottom polystyrene 96-well assay plate (Corning, Cat.# 3896)
• white 384-well assay plate (Greiner bio-one, Cat.# 781080)
• 8-row partitioned reservoir (Seahorse Bio, Cat.# 201282-100)
• 384 HH, 4X control, pyramid bottom reservoir (Seahorse Bio, Cat.# 201300-100)
• 96 LP pyramid bottom reservoir (Seahorse Bio, Cat.# 201254-100)
• frozen Instant Cells, HepG2 (Cell Culture Service, ID 89)
CyBi®-FeliX Automated Liquid Handler
The CyBi®-FeliX automated liquid handler contains 12 deck positions on two levels, allowing liquid transfer of multiple reagents between multiwell plates, tube racks and reservoirs in a small footprint of 650mm × 450mm (W × D; Figure 1). We used a 96-channel pipetting head (250µl) and an 8-channel liquid handling adapter. The 96-channel head enables parallel processing of 96-well and 384-well multiwell plates when using preracked CyBi®-RoboTipTrays. The 8-channel adapter allows pipetting into single wells and columns. Quick dispensing negates worries of cell settling in the source reservoir and minimizes the time in which cells are exposed to the environment. The pipetting head performed the switching of liquid handling adapters and loaded the pipet tips without manual intervention, allowing walk-away automation of the assays.
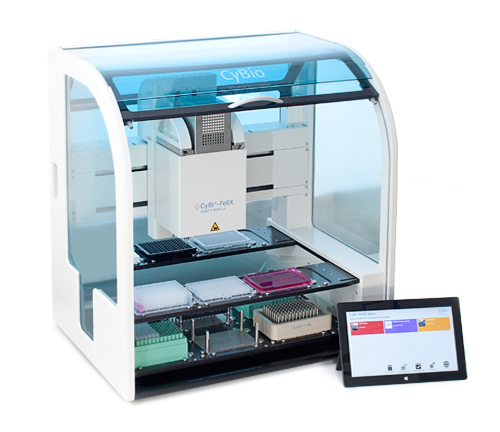

Figure 1. The CyBio CyBi®-FeliX pipetting station. The two-level deck design of the CyBi®-FeliX possesses a small footprint while providing sufficient deck positions for reagents and lab ware consumables. Users may customize the instrument configuration to meet their specific needs.
GloMax® Discover Multimode Detection System
This GloMax® Discover system includes high-performance luminescence, fluorescence, UV-Visible absorbance, BRET, FRET, filtered luminescence and kinetic measurement capabilities (Figure 2). The integrated tablet PC features intuitive software with quick drag-and-drop protocol creation. Quick-read functions and >40 preloaded and validated protocols facilitate fast and easy data generation.

Figure 2. The Promega GloMax® Discover System. The GloMax® Discover is a multimode reader that accepts plate types ranging from 6-well to 384-well multiwell plates. The instrument is operated by using an integrated tablet PC containing enhanced software with an intuitive graphical interface. Quick-read functions and preloaded detection protocols enable fast data generation. Drag-and-drop functionality expedites the creation of customized detection protocols.
Plate Layouts and Reservoir Configuration
We used a 96-well v-bottom plate (Corning) for compound titration because v-shaped wells help minimize dead volume. We then placed compounds into columns 1 and 6 of the dilution plate (Figure 3) and used the 8-channel adapter to add medium to the remaining columns and to perform the serial five-point titration for each column of compounds. The 96-channel adapter is then used to multi-dispense titrated compound to the 384-well assay plate.
An eight-row reagent reservoir helped conserve deck space by allowing both CellTiter-Glo® and Caspase‑Glo® 3/7 detection reagents to be in the same source location (Figure 3). The 96‑channel adapter with CyBi®-RoboTipTray 96/250µl was used to simultaneously transfer both solutions to the assay plate.
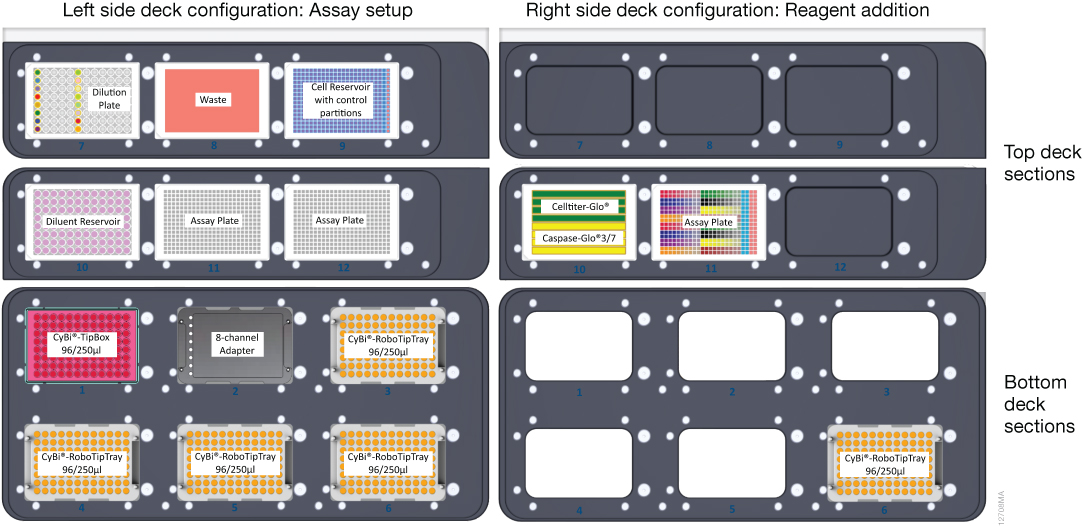
Plate Setup
- HepG2 cells were thawed and suspended in EMEM + 10% FBS + 1X penicillin/streptomycin at a density of 250,000 cells/ml. CellTox™ Green reagent was added to the medium at a 1:800 dilution; after assay assembly the overall dye dilution is 1:1000.
- The test compounds were diluted to 5X concentrations ranging from 125 to 500µM in EMEM + 10% FBS. 100µl of each diluted compound was manually arrayed to columns 1 and 6 of a 96-well plate (Figure 3), and the plate was placed on top of the deck of the CyBi®-FeliX at position 7.
- Diluent consisting of EMEM + 5% DMSO + 10% FBS was poured into a 96-well low profile pyramid-bottomed reservoir.
- The cell suspension was poured into the main partition of a 384-well 4X control pyramid bottomed reservoir. A no-cell control consisting of EMEM + 10% FBS + CellTox™ Green reagent was added to the control partitions of the reservoir (see Figure 3), and the reservoir was placed at position 9.
- Two 384-well assay plates were placed on the instrument deck at positions 11 and 12, and the bottom deck was filled with a CyBi®-TipBox 96/250µl, an 8-channel liquid handling adapter, and four CyBi®-RoboTipTrays 96/250µl.
- The automated protocol was run, directing the CyBi®-FeliX to perform the following liquid handling operations: 20µl of cell suspension and no-cell controls were dispensed into each well of the assay plates. A 5-point 1:10 serial titration of the compounds was performed in 5% DMSO in EMEM + 10% FBS diluent. Next, 5µl of each drug titration was added in quadruplicate to the cells in the assay plate. Columns 23 and 24 of the assay plate contained no cells in order to account for background signal (Figure 4).

Figure 4. Assay plate layout showing titrated compounds and controls. Using 384-well plates, we replicated the same compounds and dispense pattern in Rows A–H and Rows I–P. Key: A = bortezomib, B = cisplatin, C = methotrexate, D = nocodazole, E = panobinostat, F = staurosporine, G = terfenadine, H = tamoxifen, I = 1% DMSO vehicle control, J = no-cell control.
Reagent Addition and Signal Detection
- Assay plates were incubated for 24 or 48 hours in a cell culture incubator, 37°C and 5% CO2. Following each time point the assay plate was cooled to ambient temperature for 15 minutes, then shaken for 1 minute on an off-line electromagnetic shaker (Union Scientific).
- The plate was then placed into the GloMax® Discover to measure fluorescence changes from the CellTox™ Green dye using the blue filter pair [470nm (excitation)/500–550nm (emission)].
- Rows 1–4 of an 8-row partitioned reservoir were each filled with 5ml of CellTiter-Glo® 2.0 reagent, and rows 4–8 were each filled with 5ml Caspase-Glo® 3/7 reagent (Figure 3). The reagent reservoir and the assay plate were placed on the top deck of the CyBi®-FeliX instrument.
- The CyBi®-FeliX transferred 25µl of both reagents simultaneously to the assay plate.
- The plate was shaken for 2 minutes and incubated at ambient temperature for 60 minutes.
- Following the incubation, luminescence was measured on the GloMax® Discover. Results for cytotoxicity (CellTox™ Green Express Assay), cell viability (CellTiter-Glo® 2.0 Assay) and apoptosis (Caspase-Glo® 3/7 Assay) were graphed using GraphPad Prism® software.
Results
We assessed three mechanistic cytotoxicity profiles using three toxic compounds known to result in unique and specific mechanisms of cell death. Panobinostat is a non-selective histone deacetylase inhibitor and shows a hallmark dose-dependent increase in caspase-3/7 activity with corresponding decrease in the cell viability marker, ATP. In this study, we detected a small increase in the cytotoxicity biomarker, indicating that these apoptotic cells are beginning to undergo secondary necrosis (Figure 5, Panel A).
Tamoxifen is an estrogen receptor antagonist and in this study provides example results of cells undergoing secondary necrosis at the time of detection, which occurred after the induction of apoptosis. As seen in Figure 5, Panel B, at 24 hours the highest concentration of tamoxifen tested (100µM) produced a cytotoxic profile consisting of membrane integrity changes with profound negative effect on cell viability. However, some caspase-3/7 biomarker is detected. Previous studies have shown that this concentration of tamoxifen does induce apoptosis within a few hours of treatment (1), but due to biomarker decay, the amount of caspase activity detected decreases over time. Our low-level caspase activity detection, combined with membrane breakdown and large decrease in cell viability, suggests that apoptosis did occur much earlier and that the 24-hour time point is outside the window of maximum detection for apoptosis.
Methotrexate is a dihydrofolate reductase inhibitor that ultimately prevents synthesis of DNA and RNA. As seen at 24-hour exposure in Figure 5, Panel C, the caspase and membrane integrity biomarkers remain unchanged while the cell viability biomarker shows a potent dose-dependent effect. Because the cell membrane biomarker remains unchanged, the resulting decrease in signal compared to control suggests an anti-proliferative, rather than cytotoxic, effect.
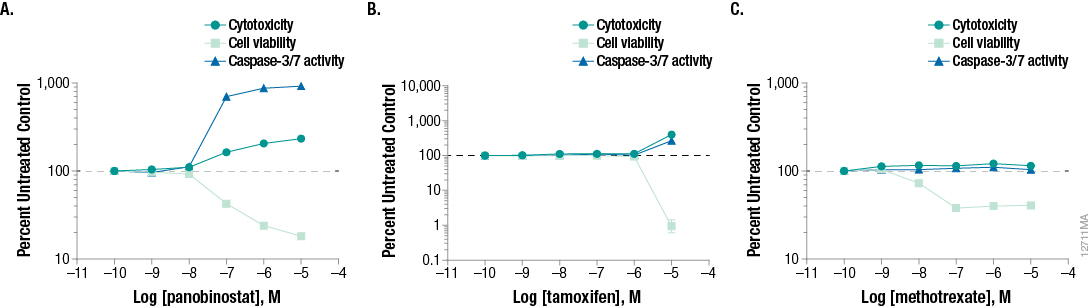
Figure 5. Representative mechanistic cytotoxicity profiles demonstrating (Panel A) apoptosis, (Panel B) necrosis, and (Panel C) growth arrest at 24 hours post treatment. We subtracted medium-only control assay background from the experimental results and plotted data as fold change relative to the untreated control on the assay plate.
Exposure time is a critical parameter when assessing mechanistic cytotoxicity. Figure 6 shows the effects of nocodazole, an inhibitor of microtubule polymerization. No change in any of the assayed biomarkers is detected 24 hours post-treatment, suggesting that cells remain viable in culture. A biologically different biomarker profile is observed 48 hours post-treatment. Dose-dependent decreases in cell viability with concordant increases in caspase-3/7 activity and cytotoxicity are observed, indicating cells are progressing through apoptosis and secondary necrosis. Therefore, concentrating testing efforts on 24 hours only could lead to the false conclusion that the test compound has no effect on cell health, while continued exposure demonstrates a cytotoxic effect.
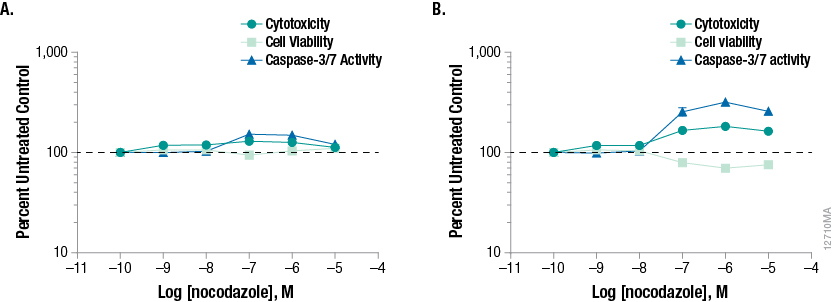
Figure 6. Time and dose-dependent effects of nocodazole, an anti-mitotic agent that disrupts microtubule formation. Overall low %CV was achieved across each titration of compound as shown by small error bars on our data graphs. The %CV was typically 5% or lower among the four replicates and reflects precise liquid handling for all additions to the well. We found the 8-channel adapter performed accurate titration of test compound as supported by the expected potency of response with our model test compounds.
Summary
We successfully implemented an automated method for distinguishing apoptosis, necrosis and growth arrest. Our automated method included using thaw-and-use HepG2 cells as our biological model system, commercial bioluminescent and fluorescent assays to assess cytotoxicity, the CyBi®-FeliX for liquid manipulation, and GloMax® Discover for readout. Three biomarkers were used to evaluate whether the chemical treatment induced necrosis, apoptosis, or growth arrest. Representative compound profiles showed expected data patterns for each unique mechanism of toxicity: panobinostat for apoptosis, tamoxifen for secondary necrosis with some apoptosis, and methotrexate for growth arrest. We also demonstrated the importance of compound exposure time when determining the mechanism of cytotoxicity(1). The typical twenty-four hour exposure time might be too early to detect some apoptotic mechanisms (e.g., the nocodazole assays).
Overall, we found the CyBi®-FeliX liquid handler to be a well-suited platform for conducting the mechanistic cytotoxicity experiments described here. The liquid handler allowed test compounds to be serially titrated and applied to the cells; it is a flexible, competitively priced instrument capable of conducting necessary assay setup and reagent delivery steps of a profiling workflow. Incorporating thaw-and-use cells provides a reproducible source of cells and removes the burden on dedicating time to cell culture, and using the CyBi®-FeliX liquid handler removed the burdensome and more variable manual liquid handling. Such an automated workflow provides researchers with walk-away convenience while still generating consistent and reproducible results. The pairing of homogeneous “add-mix-measure” cell health assays and the GloMax® Discover from Promega ensures sensitive and optimized detection for meaningful cell cytotoxicity profiling data.
References
- Riss, T.L. and Moravec, R.A. (2004) Use of multiple assay endpoints to investigate the effects of incubation time, dose of toxin, and plating density in cell-based cytotoxicity assays Assay Drug Dev. Technol. 2, 51–62.
Learn more about the CellTiter-Glo® 2.0 Assay