Value-Added Human Flexi® ORF Clones with HaloTag® Labeling Technology
Takahiro Nagase, Hisashi Yamakawa and Osamu Ohara
Kazusa DNA Research Institute
Publication Date: 2009
Abstract
Flexi® ORF clones have been constructed using human cDNA resources, including genes originally identified as long, difficult to clone cDNAs that encode large proteins. A new versatile reagent for the advancement of molecular biology and medical science has been created by providing these ORFs as genetic fusions to the HaloTag® labeling and immobilization technology.
Introduction
Expression-ready ORF (Open Reading Frame, i.e., protein coding regions) clones are one of the most critical tools for functional characterization of the gene product in vitro or in cultured cells. ORF clones are widely used for reverse-proteomics analyses in addition to biochemical studies. Protein fusion tags have been developed to facilitate detection, purification and functional studies of the fused proteins, and they have furthered the study and manipulation of structurally and chemically diverse proteins in a general and systematic manner. However, the use of different protein fusion tags best suited for the respective experimental purpose demands construction of multiple expression clones. The HaloTag® technology, providing both protein capture and labeling, allowed us to carry out various proteomic applications with a single fusion tag. Because the Flexi® Vector cloning method is suitable for the preparation of a large number of ORF clones and allows seamless implementation of HaloTag® interchangeable technology, we selected this system to construct a set of sequence-defined protein-expression clones for human genes. Initial human ORF clones were constructed using the pF1K Flexi® Vector, since it has very low non-induced expression levels. Over 5,000 pF1K ORF clones were created, and 3,000 of these ORFs have been transferred into a HaloTag® Flexi® Vector (pFN21A) to create N-terminal HaloTag® fusions. Moreover, we demonstrated that most human proteins were efficiently expressed as HaloTag® fusions in a biochemically active form in both an in vitro coupled protein expression system and in cultured mammalian cells. These fusion proteins will be directly used for functional studies.
"Over 5,000 pF1K ORF clones were created, and 3,000 of these ORFs have been transferred into a HaloTag® Flexi® Vector (pFN21A) to create N-terminal HaloTag® fusions."
Constructing and Characterizing the Expression-Ready ORF Clones
Construction of Flexi® ORF Clones
The Flexi® Vector cloning system is designed for easy cloning and transfer of ORFs between compatible vectors, with the correct orientation and reading frame, by using rare-cutting restriction enzymes SgfI and PmeI. Those 8bp restriction sites are generally added to the flanking 5´ and 3´ ends of the ORFs, respectively. The Flexi® clones were constructed by the following three methods: 1) PCR cloning; 2) ORF Trap Cloning, and 3) Restriction enzyme-based transfer from previously prepared Gateway® ORF clones. The first two methods produce Flexi® Vector-type clones, where the SgfI site is 1bp upstream of the translation start codon for the ORF and the PmeI site is placed such that a single valine codon, followed by the translation stop codon, is appended to the ORF (Figure 1, Panel A). The third method produces Flexi® Vector-RBS-type clones since remnants of the Gateway cloning sites are included 19bp upstream of the translational start codon for the ORF and two additional codons (6bp) at the end of the ORF (Figure 1, Panel A).
For PCR cloning the ORF, flanked by SgfI and PmeI sites, was amplified by PCR and cloned into the pF1K Flexi® Vector. Resultant clones were entirely sequenced in the ORF region to eliminate clones with potential PCR errors. ORF trap cloning was used to prevent artificial introduction of mutations in long ORFs by PCR. The sequence-defined ORF was transferred into pF1K-based vectors in a high-fidelity manner based on homologous recombination in Escherichia coli (JC8679), according to the method previously described.
Transferring ORFs from previously prepared Gateway ORF clones was also done in a high-fidelity manner using BstBI and SnaBI restriction endonucleases, whose recognition sites were uniquely located upstream and downstream of the ORF, respectively. The DNA fragments were inserted between the BstBI-SnaBI sites of an intermediate vector, which contains SgfI and PmeI sites upstream and downstream of the BstBI and SnaBI sites, respectively. The ORF was cut again with SgfI and PmeI restriction endonucleases and then recloned between the SgfI and PmeI sites of the pF1K-based vector. The resultant Flexi® clones are discriminated as Flexi® Vector-RBS-type from the former clones termed Flexi® Vector-type since Flexi® Vector-RBS-type clone has a ribosome-binding site (RBS) upstream of the ORF. The flanking sequences of both types of ORF are shown in Figure 1, Panel A. ORF sequences recovered by SgfI-PmeI digestion are easily transferred to Flexi® Vector pFN21A to construct N-terminal HaloTag® fusions (Figure 1, Panel B).
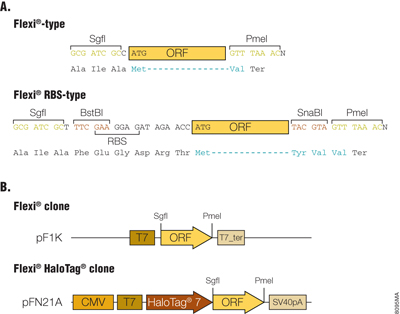
Figure 1. Flanking sequences of ORF and functional elements in Flexi® ORF clones. Panel A. The flanking sequences of ORF in the Flexi®-type and Flexi®-RBS type clones are shown. The SgfI site is placed either 1bp or 19bp upstream of the initiation codon for the Flexi® or Flexi®-RBS type clones, respectively, which allows production of recombinant proteins with the native translational initiation site or with N-terminal tags using appropriate Flexi® Vectors. The PmeI site was placed just after protein-coding sequence or 6bp downstream, which resulted in attachment of the Val in Flexi®-type or the Tyr-Val-Val in Flexi®-RBS type clones to the carboxy end of authentic ORF. When an ORF sequence flanked by SgfI and PmeI is cloned at SgfI and EcoICRI sites of a C-terminal tag fusion expression Flexi® Vector, a translational stop codon in PmeI site is destroyed and the protein can be expressed as a C-terminal fusion. RBS: ribosome binding site. Panel B. Functional elements of Flexi® ORF clones are shown. The pF1K T7 Flexi® Vector (Cat.# C8451) contains a T7 RNA polymerase promoter and terminator. The pFN21A HaloTag® CMV Flexi® Vector (Cat.# G2821) contains a cytomegalovirus (CMV) immediate-early enhancer/promoter, T7 promoter and SV40 late polyadenylation signal.
Effect of Linker Sequence Containing RBS on HaloTag® Fusion Proteins
N-terminal HaloTag® protein fusions in the pFN21A HaloTag® CMV Flexi® Vector contain an optimized peptide linker separating the ORF-encoded protein from the HaloTag® label. Because the cloning strategy employed to create Flexi® Vector-RBS-type clones inserted an additional 19bp sequence (5′-TTTCGAAGGAGATAGAACC-3′ containing an RBS sequence) between the SgfI site and the ORF sequence, additional amino acids are now appended to the peptide linker between ORF-encoded protein and HaloTag® label (Figure 2, Panel A). To elucidate whether these additional peptide linker residues in Flexi® Vector-RBS-type clones affect functionalities of HaloTag® fusions, we constructed HaloTag®-luciferase (luc2) expression clones with the Flexi® Vector-RBS-type linker sequence (HT-luc2(RBS)) or original Flexi® Vector-type linker sequence (HT-luc2) and compared them. We compared their expression levels, their HaloLink™ Resin-binding activity, their luciferase activity and their susceptibility to TEV Protease using the HaloTag® fusions expressed in vitro. HT-luc2 and HT-luc2(RBS) proteins produced in TnT® Rabbit Reticulocyte Lysate System (in vitro cell-free expression) were purified with HaloLink™ Resin, and their luciferase activities in supernatant or precipitated fractions were measured (Figure 2, Panel B). In addition, HT-luc2 and HT-luc2(RBS) proteins were incubated with or without ProTEV Protease for the time indicated in Figure 2, Panel C, and the HaloTag® fusions visualized on SDS-polyacrylamide gel electrophoresis (PAGE) after labeling with HaloTag® TMR Ligand. As results show, there were no significant differences in the abilities of either the HaloTag®-luc2 fusions produced from Flexi® Vector and Flexi® Vector-RBS-type clones in our experiments.
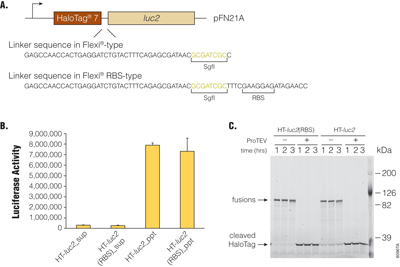
Figure 2. Effects of linker sequence in Flexi®-RBS type clones on activities of HaloTag®-fused luciferase. The HaloTag®-fused luciferases were produced using the TnT® Quick Coupled Transcription/Translation System (Cat.# L1170). Panel A. HaloTag® 7-fused luciferase (luc2)-coding region in pFN21A HaloTag® CMV Flexi® Vector is schematically represented and linker sequences between HaloTag® and luc2 in Flexi® type (HT-luc2) and Flexi®-RBS type (HT-luc2(RBS)) are shown. Panel B. Luciferase activities of HT-luc2 and HT-luc2(RBS) recovered by HaloLink™ Resin are indicated. The activities of luciferase remained in supernatant after HaloLink™ Resin- (Cat.# G1911) binding (sup) and covalently bound to HaloLink™ Resin (ppt) were measured by the Dual-Glo® Luciferase Assay System (Cat.# E2920). Panel C. Susceptibility of HaloTag®-fused luciferases to the ProTEV Protease (Cat.# V6051) are analyzed. HT-luc2 and HT-luc2(RBS) incubated with or without ProTEV Protease for the indicated time period were labeled by the HaloTag® TMR Ligand (Cat.# G8251) and detected by SDS-polyacrylamide gel electrophoresis (PAGE; 5–15%).
Characterization of HaloTag® Flexi® Clones
The ORF sequences of Flexi® clones were verified in their entirety when the original ORFs were amplified by PCR. To ensure that clones contained full-length ORFs, the N and C-terminal junctions were sequenced by single-pass sequencing. In addition, the size of each clone’s ORF was estimated by running agarose gel electrophoresis of SgfI and PmeI DNA fragments against DNA standards of known length. The expression of N-terminal HaloTag® fusions was verified by transient transfection of HEK293 cells with pFN21A-ORF clones in HEK293 cells, and HaloTag® fusion proteins were detected by HaloTag® TMR Ligand labeling. Labeled proteins were resolved on SDS-PAGE against protein markers of known size to verify the expected size of the HaloTag® fusion proteins (Figure 3, Panels A and B, respectively). Almost all pFN21A clones express their coding HaloTag® fusion proteins in HEK293 cells. This enabled us to use HaloTag® fusions transfected into HEK293 cells to produce high-nanogram to low-microgram quantities of the HaloTag® fusion proteins from 200μl of HEK293 cells. We can now analyze in vivo-produced human proteins in a high-throughput, systematic manner.
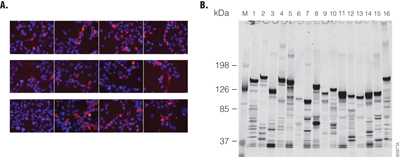
Figure 3. N-terminal HaloTag® fusions expressed in HEK293 cells. Panel A. HEK293 cells were transfected with the pFN21A HaloTag® Flexi® clones in an 8-chambered glass slide. The expressed N-terminal HaloTag® fusions were labeled by HaloTag® TMR Ligand (Cat.# G8251) and observed (red) in living cells by the BioZero fluorescent microscope (KEYENCE, Osaka, Japan) and recoded. DNA in nuclei is labeled by Hoechst33342 (blue). Panel B. After examination by microscope, the cells were lysed and extracts separated by SDS-PAGE (5–15%). The HaloTag® fusions were detected using a fluorescent image analyzer FLA-3000GF (Fujifilm, Tokyo, Japan).
Summary
Using our collection of human cDNAs we have created a collection of more than 5,000 Flexi® clones, 3,000 of which are already available with their encoded proteins fused to HaloTag® ligands. These clones can be used directly for functional studies but can also be used as donors to transfer their ORFs to other Flexi® Vectors that provide either alternate tags, e.g., a C-terminal HaloTag® or access to other expression systems. These clones have been validated for expression in cell-based expression systems.
Learn more about the Flexi® Cloning System