Preparation of Low Background Sugar-Nucleotide Donors for Use in the UDP-Glo™ Glycosyltransferase Assay
M. Osman Sheikh and Lance Wells
Complex Carbohydrate Research Center, University of Georgia, Athens, GA 30602 USA
Publication Date: April 2016
Abstract
The UDP-Glo™ Glycosyltransferase Assay is a sensitive bioluminescent assay that relies on the detection of released uridine diphosphate (UDP) upon transfer of the carbohydrate from the sugar-nucleotide donor to the acceptor molecule. Assay sensitivity depends on ultrapure sugar-nucleotide donors, available from Promega, that have less than 0.005% UDP contamination. To use sugar-nucleotide donors that are currently not available as ultrapure grade, we have developed a simple protocol to remove free UDP in sugar-nucleotide donor preparations to reduce background levels and maintain sensitivity of the UDP-Glo™ Glycosyltransferase Assay.
Introduction
Glycosyltransferases are enzymes that catalyze the transfer of ‘activated’ monosaccharides, usually a nucleoside diphosphate sugar, to a substrate protein or existing glycoconjugate, followed by the release of the nucleoside diphosphate product (Figure 1). The UDP-Glo™ Glycosyltransferase Assay (Cat.# V6961) is a simple and highly sensitive bioluminescent assay for the detection of glycosyltransferase activity that relies on the detection of released uridine diphosphate (UDP) upon transfer of the carbohydrate to the target molecule (1). However, ultrapure sugar-nucleotide donors are required to minimize background levels of signal caused by free nucleotide diphosphates that may be present in many commercial preparations of the sugar nucleotide.
Furthermore, the corresponding nucleoside diphosphates are inhibitors of glycosyltransferases (2). Here, we describe a simple method to prepare sugar-nucleotide donors for use in the UDP-Glo™ Glycosyltransferase Assay with low background levels comparable to ultrapure varieties. This approach may be useful in applications where either an ultrapure sugar-nucleotide donor is not available (such as UDP-Xylose) or where unnatural, modified sugar-nucleotide donors (i.e., donors covalently linked with chemical tags) are desired for assay.
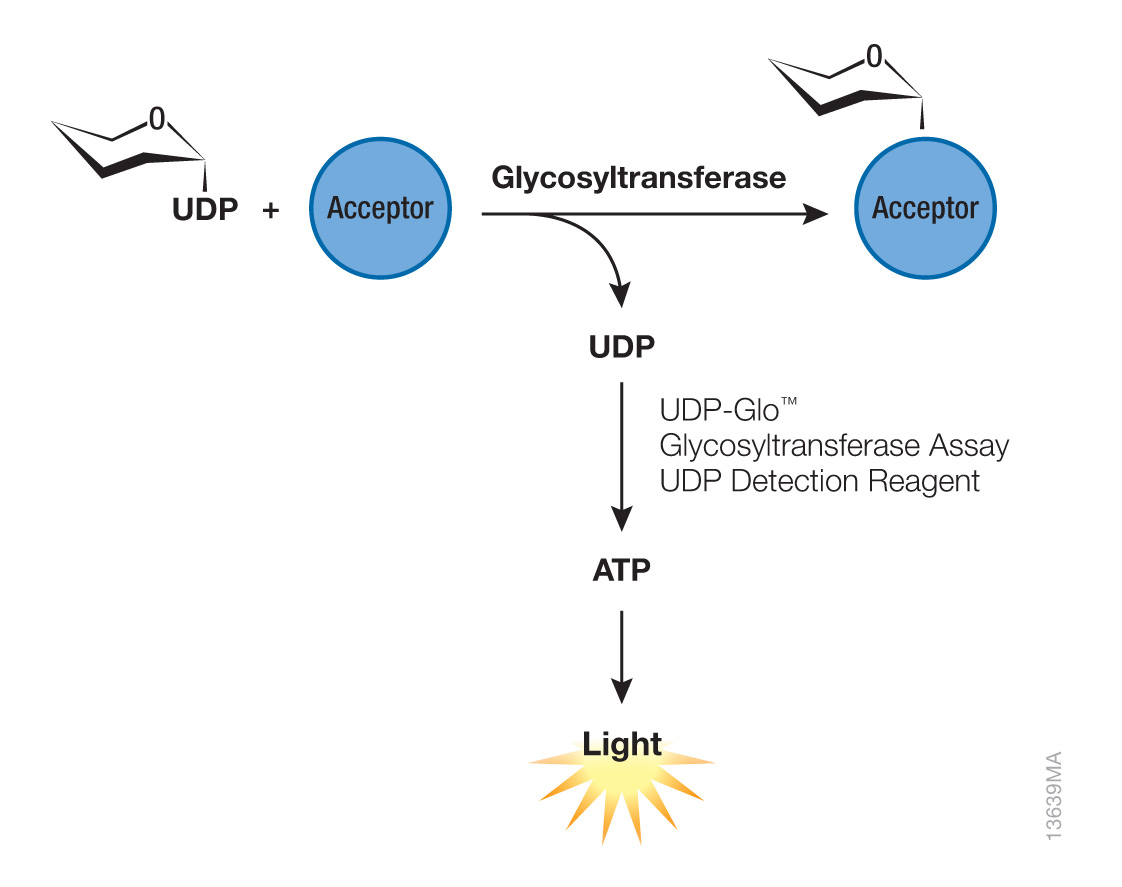
Figure 1. Glycosylation is mediated by enzymes called glycosyltransferases. Glycosyltransferases catalyze the transfer of an ‘activated’ monosaccharide, usually a nucleoside diphosphate sugar, such as uridine diphosphate (UDP), to an acceptor molecule (e.g., peptide, protein or lipid) or existing oligosaccharide. The UDP-Glo™ Glycosyltransferase Assay (Cat.# V6961) detects released UDP upon transfer of the carbohydrate to the acceptor molecule, converting UDP to ATP and then light in a luciferase reaction, which can be measured by a luminometer. Using a standard curve, luminescence is directly proportional to UDP concentration, and thus, activity of the glycosyltransferase being investigated.
Preparation of UDP-Xylose
We wanted to use the UDP-Glo™ Assay to study the xylosyltransferase activity of the glycosyltransferase LARGE(3); however, ultrapure UDP-Xylose is not available. The principle behind minimizing background levels is removal of free UDP (or ATP) that may be present in sugar-nucleotide preparations(1, 4). To accomplish this, we incubated UDP-Xylose with Calf Intestinal Alkaline Phosphatase (CIAP, Cat.# M1821) to degrade any free, contaminating nucleotide diphosphates (Figure 2). Since the downstream UDP-Glo™ Glycosyltransferase Assay converts free UDP to ATP, then ATP to light (Figure 1), it is imperative to remove all of the CIAP from the sugar-nucleotide preparation, as CIAP will cleave ATP and interfere with detection of luminescence. Therefore, following incubation of UDP-Xylose with CIAP, we used a 10kDa microcentrifuge concentrator (EMD Millipore) to filter out CIAP (~140–160kDa dimer). Using the following protocol, we were able to reproducibly ‘clean up’ UDP-Xylose (and conceivably any sugar-nucleotide donor) by degrading free UDP to uridine to use it in the UDP-Glo™ Glycosyltransferase Assay with low, comparable background relative light unit (RLU) levels compared to ultra-pure varieties (Figure 3).
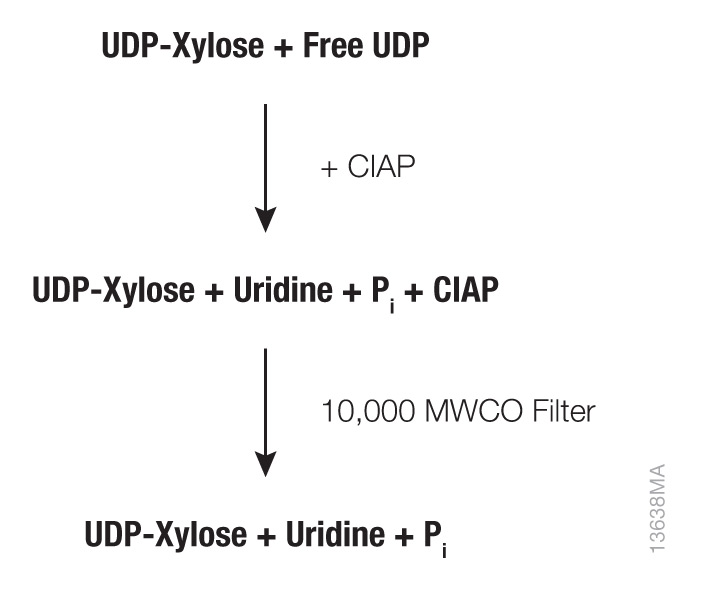

Figure 2. Sugar-nucleotide preparation workflow. Calf Intestinal Alkaline Phosphatase (CIAP, Cat.# M1821) is incubated with UDP-Xylose to degrade any free, contaminating nucleotide diphosphates. Since CIAP exists as a 140–160kDa dimer, a 10kDa molecular weight cut-off (MWCO) microcentrifuge concentrator is used to filter out CIAP.
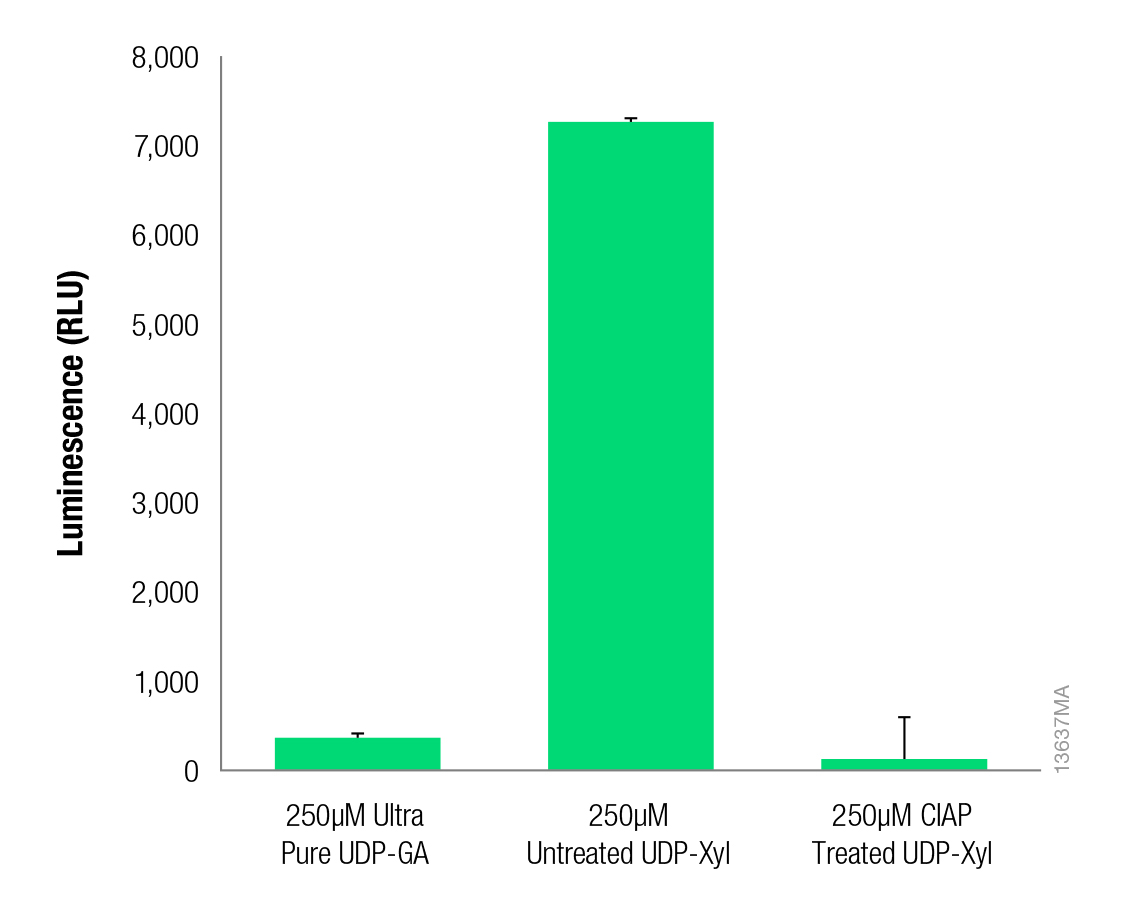
Figure 3. Background luminescence is reduced upon treatment of UDP-xylose with CIAP. Ultra Pure UDP-Glucuronic Acid (UDP-GA, Cat.# V7321), untreated UDP-Xylose (Carbosource) or CIAP-treated UDP-Xylose (250µM) was incubated for 1 hour with the UDP-Glo™ Glycosyltransferase Assay UDP Detection Reagent (Cat.# V6961). Luminescence was measured in triplicate using a GloMax®-Multi+ Luminescence Module (Cat.# E8041). Data are represented as the average of three trials, and error bars represent standard deviation.
Xylosyltransferase Assay for LARGE
To test whether our CIAP-treated UDP-Xylose works in a glycosyltransferase reaction and can be used in the UDP-Glo™ Glycosyltransferase Assay, we conducted a reaction using the bifunctional glycosyltransferase, LARGE, which possesses xylosyltransferase and glucuronyltransferase activity towards small molecule acceptors(3). Using para-nitrophenyl-β-Xyl-β4-GlcA as the acceptor and CIAP-treated UDP-Xylose as the sugar-nucleotide donor(5), LARGE activity was assayed using the UDP-Glo™ Glycosyltransferase Assay (Figure 4). Robust luminescence was detected in an enzyme concentration-dependent manner with low background. Thus, CIAP-treated UDP-Xylose is competent for use in the UDP-Glo™ Glycosyltransferase Assay.
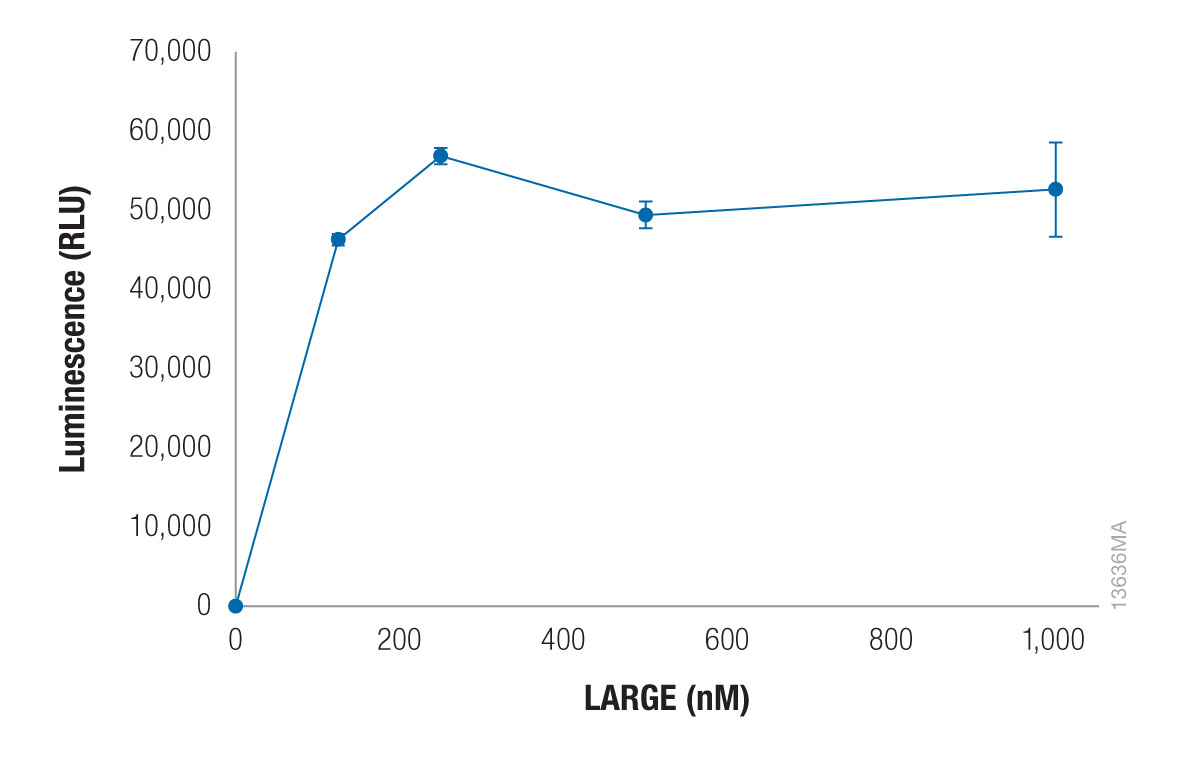
Figure 4. Xylosyltransferase activity of LARGE can be detected using CIAP-treated UDP-xylose. Glycosyltransferase reactions consisting of 1mM para-nitrophenyl-β-Xyl-β4-GlcA (acceptor), 50µM CIAP-treated UDP-Xylose (sugar-nucleotide donor) and increasing concentrations of LARGE were incubated at 37°C for 16 hours in 0.1M MES (pH 6.0), 10mM MgCl2 and 10mM MnCl2. The glycosyltransferase reactions were incubated for 1 hour with the UDP-Glo™ Glycosylation Assay UDP Detection Reagent (Cat.# V6961). Luminescence was measured in triplicate using a GloMax®-Multi+ Luminescence Module (Cat.# E8041). Data are the average of three trials, and error bars represent standard deviation. In the absence of acceptor, RLU levels were ~2,500 at 1,000nM LARGE (data not shown).
Protocol
Reagents
- UDP-Xylose (Carbosource, University of Georgia; Note: Theoretically, this can be any other sugar-nucleotide of interest.)
- Calf Intestinal Alkaline Phosphatase (CIAP), 1 unit/µl (Cat.# M1821)
- 10X CIAP Buffer (Cat.# M1833)
- 0.2µm microcentrifuge filter (Pall/Nanosep Cat.# ODM02C35)
- Microcon-10kDa Centrifugal Filter Unit with Ultracel-10 membrane (EMD Millipore Cat.# MRCPRT010)
Equipment
- 37°C incubator shaker
- Microcentrifuge
Procedure
- Prepare a reaction in 1X CIAP Buffer with 3 units of CIAP per 10µmol of UDP-Xylose in a 1.5ml microcentrifuge tube. Promega instructions say, “One unit is defined as the amount of enzyme required to catalyze the hydrolysis of 1µmol of 4-nitrophenyl phosphate per minute at 37°C in 1M diethanolamine, 10.9mM 4-nitrophenyl phosphate, 0.5mM MgCl2 (pH 9.8).”
Note: The ratio of CIAP to UDP-Xylose may need to be optimized; however, these values worked well in our studies. We prepared 1,000µl of 10mM UDP-Xylose with 3 units of CIAP in 1X CIAP Buffer. - Incubate for 14–16 hours at 37°C with gentle shaking (~100–150rpm).
- Filter sample through a 0.2µm microcentrifuge filter at 14,000 × g for 1 minute to remove precipitates. Save the filtrate.
- Apply the filtrate from Step 3 through a Microcon-10kDa Centrifugal Filter Unit with Ultracel-10 membrane at 14,000 × g for 10–15 minutes to remove CIAP. Save the filtrate. The UDP-Xylose is ready for use or can be stored at –20°C.
- Continue downstream glycosyltransferase activity assay as described for the UDP-Glo™ Glycosyltransferase Assay using the CIAP-treated sugar-nucleotide prepared in Step 4.
Conclusions
Here, we report the use of CIAP to hydrolyze contaminating nucleotide diphosphates in sugar-nucleotide preparations for use in the UDP-Glo™ Glycosyltransferase Assay when ultrapure sugar-nucleotide varieties are unavailable. Although untested in our laboratory, this principle may be extended to the GDP-Glo™ and UMP/CMP-Glo™ Glycosyltransferase Assays (currently in development; contact Michael Curtin, Product Manager, for details). In addition to soluble CIAP, we have tested the feasibility of using CIAP conjugated to agarose beads (Sigma Insoluble Enzymes) to aid in removing the phosphatase. However, 0.2µm filtration of the CIAP-conjugated beads alone was not sufficient to remove the CIAP, presumably due to residual amounts of unconjugated CIAP in the preparation. Thus, filtering using MW cut-off filters is suggested. We use unconjugated, soluble CIAP in our protocol presented above. In conclusion, we expect that this methodology will be particularly useful in applications where ultrapure-grade standard and synthesized unnatural sugar-nucleotide donors are desired to be assayed using the bioluminescent glycosyltransferase assays.
References
- Zegzouti H. (11/2014) Detection of glycosyltransferase activities with homogeneous bioluminescent UDP and GDP detection assays. Glycobiology 24 (11), 1051.
- Thayer, D.A. and Wong, C.H. (2007) Enzymatic synthesis of glycopeptides and glycoproteins. In: Glycopeptides and Glycoproteins: Synthesis, Structure and Application 267, 37–63.
- Yoshida-Moriguchi, T. and Campbell, K.P. (2015) Matriglycan: a novel polysaccharide that links dystroglycan to the basement membrane. Glycobiology 25, 702–13.
- UDP-Glo™ Glycosyltransferase Assay Technical Manual TM413, Promega Corporation
- Praissman, J.L., Live, D.H., Wang, S., Ramiah, A., Chinoy, Z.S., Boons, G.J., Moremen, K.W., Wells, L. (2014) B4GAT1 is the priming enzyme for the LARGE-dependent functional glycosylation of α-dystroglycan. eLife 3.
Learn more about the UDP-Glo™ Glycosyltransferase Assay