In Vivo Evaluation of Regulatory Sequences by Analysis of Luciferase Expression
Nerea Razquin and Puri Fortes*
Division of Gene Therapy and Hepatology, CIMA, University of Navarra, Pio XII 55, 31008 Pamplona, Spain
*Corresponding Author
Publication Date: 2009
Abstract
Promoters and regulatory sequences that allow high expression levels in cell culture are not always efficient in animal models. In this work we have compared performance in vivo of different promoters and regulatory sequences, which have been shown to yield high and prolonged expression in cultured cells. To this aim, we have evaluated luciferase expression in mice liver after hydrodynamic injection of plasmids containing a luciferase reporter and the regulatory sequences. Luciferase activity has been evaluated at different times after plasmid injection using an in vivo imaging system with a cooled charge-coupled device (CCD) camera.
Introduction
High expression levels over long periods of time are required for many gene therapy applications. To obtain high expression, several sequences have been described. These include enhancers and promoters, which increase transcription; introns and mRNA stabilization and transport sequences, which favour processing and accumulation of mRNAs; and translation enhancers, which increase translation efficiency. Additionally, several protocols have been described to improve introduction of exogenous DNA in the nucleus of the cell of interest and increase nuclear DNA stability. Interestingly, methods that allow replication of exogenous DNA upon cell division also have been described. These methods function either by inducing replication of episomal DNA or integration of exogenous DNA into the cell genome. However, most of the tools described to increase expression of exogenous genes have been evaluated in cultured cells. When functionality is assayed in animal models, many of these methods behave differently and do not yield high expression levels for long periods of time. Expression of exogenous genes in animals is hampered by natural systems developed to avoid expression of toxic factors. These systems often find exogenous genes as unwanted and activate machineries developed to inhibit their expression such as the immune system or molecular mechanisms that silence gene expression such as promoter methylation. The aim of our work was to evaluate the efficacy of regulatory sequences in a mouse animal model. We have cloned the regulatory sequences in a plasmid that contains luciferase sequences and evaluated luciferase expression in cultured cells to ensure functionality. The same plasmids have been introduced in mice by hydrodynamic injection, and luciferase activity has been evaluated with a CCD camera at different times after plasmid administration.
Reagents, Materials and Instrumentation
HeLa and HuH7 cells were obtained from the ATCC and treated as recommended (1). Cells were transfected with calcium phosphate (1). Renilla and firefly luciferase activities were measured using the Dual Luciferase® System (Promega) as previously described (1) in a Berthold Luminometer (Lumat LB 9507). Eight week-old Balb/C mice were used in all experiments following the guidelines from the institutional ethical committee. Hydrodynamic injections were performed as described with 5mg of luciferase expressing plasmid and 20mg of an unrelated carrier plasmid (2), and luciferase activity was evaluated using a CCD camera (Xenogen) after intraperitoneal injection of 3mg of luciferin substrate (Promega) (3).
Evaluation of Luciferase Expression from CMV Promoter In Vivo
Cytomegalovirus (CMV) promoters have been widely used to express high levels of exogenous proteins in cultured cells. Similarly, we obtained high expression levels when a plasmid expressing luciferase under a CMV promoter (pCMVLuc, Figure 1, Panel A) was transfected in HeLa cells (data not shown). When the same plasmid was introduced in mice by hydrodynamic injection, we also detected very high luciferase activity with a CCD camera (Figure 2, Panel A). Quantification showed 5 × 1010 luciferase units (photons/second × cm2 × sr) in the liver of treated mice one day after plasmid administration (Figure 2, Panel B). However, luciferase activity decreased rapidly and reached a plateau of 5 × 107 luciferase units 20 days after plasmid administration (Figure 2). Thus, there was a 1000-fold decrease in luciferase expression. This reduction in expression has been previously described, and it has been associated with promoter methylation (2,4,5).
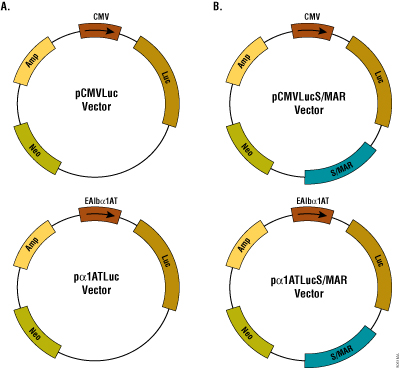
Figure 1. Schematic of the plasmids used in this study. Panel A. Plasmid pCMVLuc and pα1ATLuc express luciferase from CMV promoter and the α1AT promoter fused to the mouse albumin enhancer, respectively. Panel B. A S/MAR sequence was cloned in the plasmids described in A to generate pCMVLucS/MAR and pα1ATLucS/MAR. Amp and Neo denote ampicillin and neomycin resistance genes, respectively.
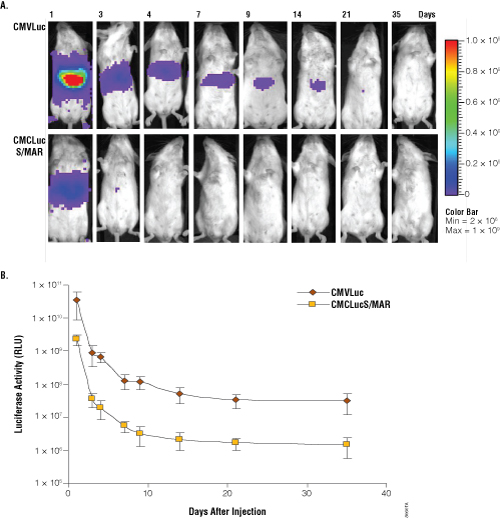
Figure 2. Analysis of luciferase activity in mice at different times after administration of pCMVLuc and pCMVLucS/MAR. Panel A. Imaging of light emission. The color gradient is indicated to the right. Panel B. Quantification of the data shown in Panel A. Luciferase units are photons/second × cm2 × sr.
Evaluation of the Effect of S/MAR Sequences in Luciferase Expression from CMV Promoter In Vivo
In cell culture, silencing of the CMV promoter by methylation has been prevented by the presence of scaffold/matrix attachment regions (S/MARs) (6). S/MARs are sequences found in all eukaryotes, often associated with chromosomal origins of bidirectional replication. Cis-acting S/MAR sequences have been shown to allow plasmid replication and therefore plasmid maintenance in several tissue culture cells (7). S/MAR sequences can bind the scaffold attachment factor A protein, which is the principal component of chromatin and chromosomes (8), and this interaction could bring the A/T-rich S/MAR sequence, which tends to be single stranded, close to the replication machinery. Therefore, plasmids containing a S/MAR sequence are replicated once per cell cycle, resulting in episomal stability. Interestingly, S/MAR functionality has been described recently to allow episomal maintenance in primary hematopoietic cells derived from transfected human cord blood CD34(+) cells (9) and to allow generation of genetically modified pigs (10).
We introduced a S/MAR sequence in the plasmid expressing luciferase under the CMV promoter (pCMVLucS/MAR, Figure 1, Panel B). We hypothesized that this construct might lead to stable expression levels due to the lack of silencing of the CMV promoter and to stable plasmid maintenance by episomal replication. To assay the functionality of the vector, we first evaluated luciferase activity after transfection into HeLa cells. High expression levels of luciferase were detected after transfection, but luciferase activity decreased gradually in the cells after cell passage in the absence of selection (data not shown). However, luciferase activity was stable after selection with G418, which requires neomycin expression and selects cells that contain the plasmid (data not shown). This pattern was observed in pools or in isolated clones. To evaluate whether these clones had the plasmid integrated in their genome or in an episomic fraction, a Southern-blot analysis was performed. Plasmid DNA was found in the episomal fraction, and this DNA could be recovered after transformation in bacteria (data not shown). These data indicate that the S/MAR sequence was functional in episomal plasmid replication in our system. Thus, we introduced pCMVLuc as a control and pCMVLucS/MAR by hydrodynamic injection in mice, and we evaluated luciferase activity in vivo with a CCD camera. Surprisingly, luciferase activity was lower one day after injection of pCMVLucS/MAR (3 × 109 luciferase units) than after treatment with pCMVLuc (5 × 1010 luciferase units) (Figure 2). Additionally, luciferase activity in pCMVLucS/MAR- treated animals decreased very fast after plasmid administration, reaching a plateau of 3 × 106 luciferase units 15–20 days after hydrodynamic injection. The presence of S/MAR sequences did not avoid the 1000-fold decrease in expression observed with the CMV promoter. The decrease in expression could indicate CMV promoter silencing even in the presence of S/MAR sequences. When the promoter is silenced, transcription is impaired, and S/MAR sequences cannot activate replication (11). Therefore we decided to analyze the effect of S/MAR sequences with a promoter that allows stable expression of transgenes.
Evaluation of Luciferase Expression from Albumin Enhancer Fused to alpha1AT Promoter In Vivo
We previously showed that combination of the mouse albumin enhancer with the α1antitrypsin (α1AT) promoter results in liver-specific stable expression of α1AT (2). Therefore, we decided to test expression of luciferase from this promoter in vivo. Thus, we constructed plasmids pα1ATLuc and pα1ATLucS/MAR, with or without S/MAR sequences, respectively (Figure 1). HuH7 human liver cells transfected with these plasmids showed strong luciferase activity (data not shown). Luciferase activity in cells transfected with pα1ATLuc decreased rapidly upon cell passage but was more stable in cells transfected with pα1ATLucS/MAR, suggesting S/MAR functionality (data not shown). These plasmids were introduced by hydrodynamic injection in mice, and we evaluated luciferase activity in vivo with a CCD camera. The results show that luciferase activity is very high one day after transfection (8 × 109 luciferase units) and similar for both constructs (Figure 3). Then, there is a reproducible increase in luciferase activity in the animals treated with pα1ATLucS/MAR that could account for S/MAR functionality. However, this increase is transient. After this transient increase, luciferase activity is maintained in animals injected with pα1ATLuc and pα1ATLucS/MAR for longer than a month (Figure 3, Panels A and B). In order to detect plasmid replication, we forced hepatocyte cell division by performing a hepatectomy of 70% of the liver of mice injected with pα1ATLuc and pα1ATLucS/MAR. Hepatocyte regeneration should stimulate S/MAR-mediated replication, and therefore higher expression levels should be observed in animals treated with pα1ATLucS/MAR than in animals injected with pα1ATLuc. However luciferase activity was similar in hepatectomized pα1ATLuc and pα1ATLucS/MAR- treated mice (Figure 3, Panel C). Similar results were obtained in animals injected with 100-fold lower dose of the plasmids, indicating that the negative result is not due to saturation of the replication initiation machinery (data not shown). So far we cannot conclude that S/MAR sequences induce plasmid replication in mouse liver after hydrodynamic injection.
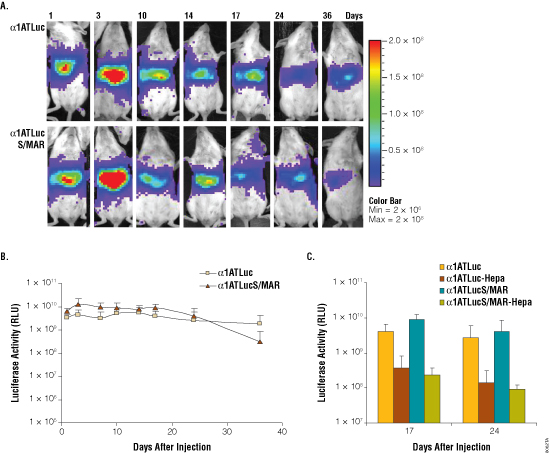
Figure 3. Analysis of luciferase activity in mice at different times after administration of p-alpha-ATLuc and p-alpha-ATLucSMAR. Panel A. Imaging of light emission. The color gradient is indicated to the right. Panel B. Quantification of the data shown in Panel A. Panel C. Quantification of luciferase activity at the indicated days post-hydrodynamic injection of sham operated and hepatectomized (Hepa) animals. Hepatectomy was performed at day 8 post-hydrodynamic injection. Luciferase units are photons/second × cm2 × sr.
Summary
We have not been able to detect replication of S/MAR-containing plasmids in mouse liver after hydrodynamic injection. While preparing this article, another work was published showing similar results (12). However, we have obtained high and stable luciferase expression levels in vivo with the combination of the mouse albumin enhancer and the α1AT promoter. These results have been obtained by harmless, fast and easy detection of luciferase expression. We highly recommend this method over others available in the market. Unlike evaluation of chloramphenicol acetyltransferase, luciferase measurement does not require the sacrifice of the animals at different times post-treatment and, unlike secreted alkaline phosphatase or α1AT, luciferase does not require stressful blood extractions. Green fluorescent protein also can be detected in vivo, but detection requires excitation light, is less sensitive and shows a high background and, therefore, a low dynamic range. Thus we recommend evaluation of luciferase expression to analyze in vivo expression and for other applications such as biodistribution analysis of viral vectors or tumor cells expressing luciferase.
References
- Aparicio, O. et al. (2006) Adenovirus virus-associated RNA is processed to functional interfering RNAs involved in virus production. J. Virol. 80, 1376–84.
- Kramer, M.G. et al. (2003) In vitro and in vivo comparative study of chimeric liver-specific promoters. Mol. Ther. 7, 375–85.
- Vera, M. et al. (2007) Liver transduction with a simian virus 40 vector encoding insulin-like growth factor I reduces hepatic damage and the development of liver cirrhosis. Gene Ther. 14, 203–10.
- Guo, Z.S. et al. (1996) Evaluation of promoter strength for hepatic gene expression in vivo following adenovirus-mediated gene transfer. Gene Ther. 3, 802–10.
- Löser, P. et al. (1998) Reactivation of the previously silenced cytomegalovirus major immediate-early promoter in the mouse liver: Involvement of NFkappaB. J. Virol. 72, 180–90.
- Jenke, A.C. et al. (2004) Expression of a transgene encoded on a non-viral episomal vector is not subject to epigenetic silencing by cytosine methylation. Mol. Biol. Rep. 31, 85–90.
- Piechaczek, C. et al. (1999) A vector based on the SV40 origin of replication and chromosomal S/MARs replicates episomally in CHO cells. Nucleic Acids Res. 27, 426–8.
- Jenke, B.H. et al. (2002) An episomally replicating vector binds to the nuclear matrix protein SAF-A in vivo. EMBO Rep. 3, 349–54.
- Papapetrou, E.P. et al. (2006) Gene transfer into human hematopoietic progenitor cells with an episomal vector carrying an S/MAR element. Gene Ther. 13, 40–51.
- Manzini, S. et al. (2006) Genetically modified pigs produced with a nonviral episomal vector. Proc. Natl. Acad. Sci. USA 103, 17672–7.
- Stehle, I.M. et al. (2003) Exploiting a minimal system to study the epigenetic control of DNA replication: The interplay between transcription and replication. Chromosome Res. 11, 413–21.
- Argyros, O. et al. (2008) Persistent episomal transgene expression in liver following delivery of a scaffold/matrix attachment region containing non-viral vector. Gene Ther. 15, 1593–605.
Dual-Luciferase® Reporter Assay System