Size Does Matter: NanoLuc® Technologies Advance Virology Research
Kyle Hooper
Promega Corporation
Publication Date: September 2018
t-pub 202
Introduction
In 2012, a new reporter enzyme called NanoLuc® Luciferase (NLuc) and its detection reagent, Nano-Glo® Luciferase Assay System, were introduced. NLuc was molecularly evolved from a 19kDa deep sea shrimp luciferase to increase stability and improve brightness. The natural substrate of the original luciferase, coelenterazine, was chemically evolved to a brighter, more stable substrate, furimazine. Together, NLuc and furimazine produce a luminescent signal >100-fold brighter than either firefly luciferase (FLuc) or Renilla luciferase (RLuc). The bonus is that the NanoLuc® luminescent signal has a long lifetime, exhibiting glow kinetics rather than a rapid flash signal like other small, bright luciferases such as Gaussia luciferase. Additionally, other key features of NLuc made it an ideal reporter. The small 19kDa enzyme was thermally stable, active over a broad pH range and required no post-translational modifications or maturation after translation to make an active enzyme. NLuc produces blue light with maximum emission at 460nm. Details of the creation, characterization and early applications of NLuc and Nano-Glo® Assay System are found in Hall et al. (1) .
While imaging the blue light of NLuc does present a challenge in animal models, the brightness and long lifetime of the signal, as well as its small size, are significant benefits
Luciferase enzymes are an important tool for making recombinant reporter viruses, which are used to further understanding of viral life cycles and lethality in animal models (2) . Though luciferases come with their challenges, viruses with embedded reporters make it easier to follow infection in the same animal over time using bioluminescent imaging. FLuc is most commonly used for in vivo animal imaging due to its emission at ~600nm in live animals, which is better for deep-tissue imaging. At 61kDa, FLuc is a rather large protein, and limitations on viral capsid size mean that many viral genomes cannot incorporate an additional 1,600+ nucleotides of DNA. Viruses are more tolerant of the 34kDa RLuc reporter, but imaging is difficult because its blue light emission is easily scattered within animals. While imaging the blue light of NLuc does present a challenge in animal models, the brightness and long lifetime of the signal, as well as its small size, are significant benefits. Gaussia luciferase is also a small, bright enzyme, but is challenging to work with in vivo due to secretion from the cell and a rapid flash signal.
This article will review two examples of how the NLuc size produced better, more stable recombinant viruses. See Table 1 for a more extensive list of NLuc-containing viruses or virus-like particles that have been cited in peer-reviewed publications. Recently, a short 11-amino acid bioluminescent protein tag was introduced that uses structural complementation to achieve a functional luciferase, which also can be applied to viral research. A review of two early papers using this peptide tag, known as HiBiT, follows the section concerning full-length NLuc.

Viral Studies with Full-Length NanoLuc® Luciferase
Generation of a near-native, replicative reporter Influenza A virus
Generating an influenza reporter virus is difficult for several reasons (3) :
- All the viral genes are critical in vivo, precluding simple gene replacement with a reporter.
- The compact genome does not tolerate large insertions, which are destabilized and lost over time.
- Insertions can severely attenuate replication.
- Insertions at the end of coding regions can disrupt packaging.
An influenza reporter virus would allow real-time analysis of the infection process in live animals rather than cell culture models, which is important for determining the pathogenicity and transmission potential of emerging influenza viruses.
Tran et al. (4) overcame these challenges by using the small, bright NLuc and clever approaches to recombinant virus design. The polymerase subunit gene PA was known to tolerate insertions at the C terminus and was chosen as the integration site. Though PA-NLuc fusions were deemed as active as wild-type PA, the viral construct chosen used the self-cleaving 2A peptide from porcine teschovirus as a linker (ultimately producing separate PA and NLuc proteins) to avoid any problems with other PA functions like trafficking and assembly.
The recombinant NLuc virus replicated with near-native properties in culture and in vivo. Treatment with ribavirin demonstrated that the luminescence from NLuc was due to gene expression and not fortuitous carryover or packaging of NLuc. Cell-based assays comparing plaque forming units versus luminescence correlated well (r2 = 0.99), with the benefit that the NLuc assay takes 8 hours compared to the 48–72 hour plaque assay.
The NLuc virus exhibited pathogenicity and lethality in mice indistinguishable from the parental virus. The virus was used to perform in vivo imaging and to track viral load and spread of infection into mouse lungs. A known mutation that restricts replication of Influenza A strains to avian species was shown to produce the same phenotype with the NLuc virus. Thus, this reporter virus could evaluate emerging viruses as well as assess antivirals in animal models. Later work by the lab also demonstrated contact and airborne transmission of a different NLuc-expressing H1N1 influenza A virus in a ferret model (4) .
Integration of NanoLuc® Luciferase generates stable, replicative reporter alphaviruses
While reporter alphaviruses for tracking viral replication in vitro and in animals do exist, inserting the FLuc reporter gene has several disadvantages, including:
- Potential for rapid transgene loss.
- Replication attenuation.
Sun, et al. (5) sought to address these problems by choosing a different integration site and using NLuc.
Previous work directed reporter fusions to the poorly conserved carboxy-terminal half of nonstructural protein 3 (nsP3). Sun et al. targeted that site in Sindbis, Chikungunya, and both Eastern and Venezuelan equine encephalitis viruses with FLuc (61kDa; 1,650 nucleotides) or NLuc (19kDa; 513 nucleotides). Other sites and strategies were examined as well. An insertion site between the capsid and PE2 protein was used for both FLuc and NLuc with the Trosea asigna virus (TaV) peptide 2A sequence included to create the reporter without protein fusion. Recombinant viruses were also created expressing either NLuc or FLuc from a subgenomic promoter not fused to any viral protein, but still part of the viral genome.
Inserts with FLuc were rapidly lost, no matter which insertion method used, as judged by diminished luminescence over cell passages and Western blots demonstrating loss of expressed protein. The NLuc constructs were very stable. The nsP3 fusion, whether to FLuc or NLuc, showed lower expression compared to the nsP3 gene alone. Lethality studies confirmed that the TaV-NLuc constructs were just as lethal as the parent viruses. Imaging of animal models verified that the NLuc-expressing viruses replicated in vivo and spread throughout the animal whereas luminescence from FLuc-expressing viruses was localized to the site of injection due to loss of the FLuc insert after repeated replication.
Studies with HiBiT Bioluminescent Protein Tag
In 2016, we introduced a protein structural complementation technology called NanoLuc Binary Technology or NanoBiT (6). The initial peer-reviewed paper focused on how NLuc was divided between amino acids 156 and 157 to produce a 156aa subunit and a 13aa subunit. The larger subunit was molecularly evolved to improve stability and expression, yielding the LgBiT subunit. LgBiT has little inherent enzymatic activity. Over 350 peptide variants of the 13aa subunit were made in vitro and tested for both binding properties to LgBiT and structural complementation of LgBiT to produce a new functional enzyme (NanoBiT® Luciferase). Two peptides stood out: One that had minimal affinity for LgBiT, termed SmBiT; and one that had extremely high affinity for LgBiT, termed HiBiT. LgBiT and SmBiT subunits are used to measure protein:protein interactions where the affinity of the proteins under study brings LgBiT and SmBiT into close enough proximity to interact. This intramolecular study of protein:protein interactions is performed using the NanoBiT® Protein:Protein Interaction System.
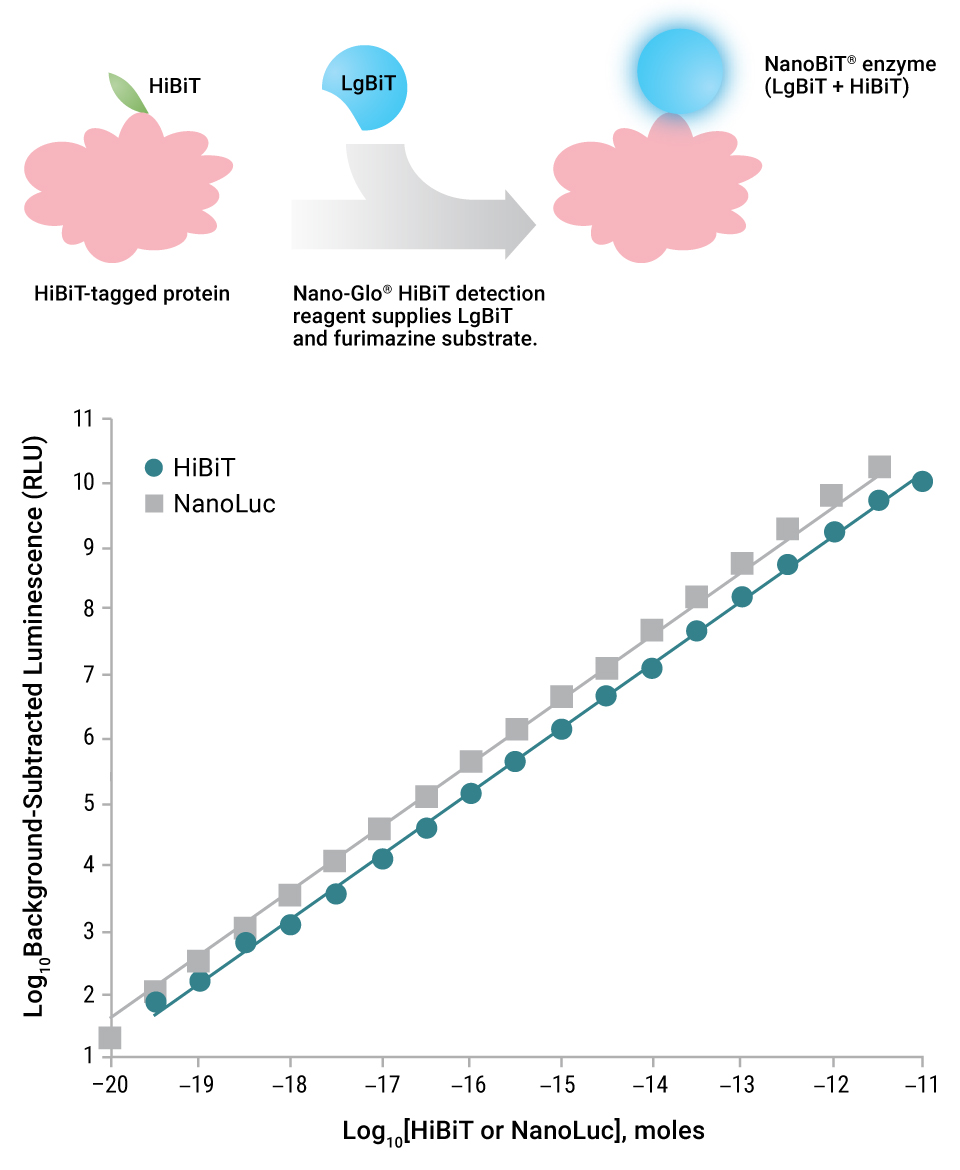
HiBiT could spontaneously complement LgBiT to form a NanoBiT® luciferase enzyme that is nearly as bright as NLuc. Work by Schwinn, et al. (7) examined the utility of the HiBiT subunit as a bioluminescent protein tag. The HiBiT tag (11 amino acids) can be appended to a protein, and this fusion can be recognized by reagents containing the LgBiT subunit and furimazine substrate. Three detection reagents for HiBiT-tagged proteins are available: Nano-Glo® HiBiT Lytic Detection System, Nano-Glo® HiBiT Extracellular Detection System and a unique reagent for detecting HiBiT-tagged proteins on Western blots, the Nano-Glo® HiBiT Blotting System.
Just as NLuc accelerated virology research due to its small size, the effect of HiBiT looks promising. Rather than contend with a 169aa (~500 nucleotides) reporter like NLuc, a small 11aa (33 nucleotide) insertion is now sufficient. Early examples of how the HiBiT tag could be used in virology have already been published.
HiBiT tagging simplifies research on West Nile Virus entry and release
Flaviviruses such as West Nile virus typically require biocontainment facilities to do conventional virology work. Investigations into the attachment of the virus to target cells and release of viral particles do not require the whole genome of the virus. Simply providing a cell with a plasmid containing the structural proteins results in budding of viral-like particles from the endoplasmic reticulum and secretion from the cell. These subviral particles (SVP) can be used to study the attachment of the virus to cells without the need of a biocontainment facility.
Sasaki et al. (8) have developed a system to look for inhibitors of West Nile virus entry. These experiments depended upon an ELISA-based assay to quantitate the number of viral particles. Under the best conditions, the assay takes at least 2 hours with a limited range of quantitation (~twofold). HiBiT was incorporated into the intra-SVP loop of the E protein and used to make HiBiT-subviral particles (SVP-HiBiT). Plasmids containing the viral M protein and the HiBiT-E protein were transfected into 293T cells and the appearance of HiBiT-SVP in samples of the culture medium over a 48-hour period could be monitored using the Nano-Glo® HiBiT Extracellular Detection System. The Nano-Glo® HiBiT Assay took only 10 minutes to perform and showed a large linear quantitation range (~fivefold). The presence of the HiBiT sequence was confirmed by Western blot using the Nano-Glo® HiBiT Blotting System. An siRNA knockdown of the Rab11 G-protein greatly reduces secretion of conventional West Nile SVP and likewise inhibited release of the HiBiT-SVP.
HiBiT technology allows sensitive detection of viruses and subviral particles
Including the West Nile virus capsid protein with E and M proteins, along with a construct to make a subgenomic RNA with the packaging sequences, creates West Nile virus virus-like particles (VLPs). These VLPs can deliver a gene to target cells for expression. In addition, VLPs allow examination of cell attachment, and the subgenomic RNA can contain a reporter to demonstrate viral internalization. The HiBiT sequence was added to the C-terminus of the capsid protein and attachment and internalization of VLP-HiBiT was monitored using a Vero cell line stably expressing the LgBiT protein. The formation of the NanoBiT® enzyme that occurs when VLP-HiBiT was internalized was monitored using the Nano-Glo® Live Cell Assay System. The NLuc-VLP was used to demonstrate how the system could be used to screen for inhibitors of virus entry—in this case, an antibody to the West Nile virus E protein or inhibitors of clathrin-coated pit formation.
HiBiT Technology applied to other virus-cell interaction model systems
Yamamoto, et al. (9) used a similar HiBiT strategy to measure gp41-dependent cell fusion of HIV-1 VLPs with an engineered HEK 293 cell line expressing CD4, CCR5 and a HaloTag®-LgBiT fusion. The assay could monitor cell fusion and allow site-directed mutagenesis of the gp41 sequence to identify critical residues.
Torriani, et al. (10) developed a cellular assay to mimic arenavirus-target cell fusion in the endosome. Two VeroE6 cells cell lines were created expressing an arena virus GP protein. One expressed GFP and LgBiT and the other expressed RFP and HiBiT. The two cells were mixed and upon a drop in extracellular pH, the two cell lines fused. Fusion was confirmed by fluorescent imaging and formation of NanoBiT® Luciferase. The fusion could be blocked by the compound TRAM-34, a KCa3.1 calcium-activated potassium channel blocker. The TRAM-34 treatment could block fusion mediated by either the Machupo virus or the lymphocytic choriomeningitis virus GP proteins.
HiBiT tagging produces better reporter flaviviruses with near-native replication
Reporter viruses are useful for determining the viral life cycle and discovering required host cell proteins. A recombinant NLuc-hepatitis C virus (HCV) helped Puig-Basagoiti et al. (11) determine the importance of host apolipoprotein E in secretion of the virus. The lab was concerned about the difference in replication of the virus that contained the full-length NLuc. Therefore, they tested the HiBiT tag to obtain better replication dynamics (12) . The lab produced HiBiT-tagged HCV, Dengue (DENV), Japanese encephalitis (JEV) and bovine viral diarrhea (BVDV) viruses and recovered stable HiBiT reporter viruses with better replication, significantly higher than viruses containing full-length reporters.
In vitro work to determine recombinant viral infectivity and replication rates was performed either in cells stably expressing the LgBiT or using the Nano-Glo® HiBiT Lytic Detection Reagent. The HiBiT-HCV was able to confirm the importance of apolipoprotein E to viral secretion. The HiBiT-JEV replicated in a variety of host cells including hamster (BHK21), human (HeLa, A549, Huh7), monkey (Vero E6) and mosquito (C6/36). The HiBiT-DENV was tested only in human and mosquito cells and found to replicate. With BVDV, HiBiT was fused to an envelope protein and the lab was concerned that the HiBiT fusion would alter viral particle formation. However, the fusion was found to have no effect in comparison to wild-type virus.
All four HiBiT viruses were tested for susceptibility to known flavivirus inhibitors and behaved like the parental viruses. HiBiT-HCV and HiBiT-JEV were used to screen a panel of 69 protease inhibitors and identified inhibitory compounds using the luciferase readout, rather than viral titers. The screen identified known HCV inhibitors and turned up a compound that was effective against HiBiT-JEV that, in turn, was effective against HiBiT-DENV. The HiBiT-HCV replicated in a human liver chimeric mouse model and responded to HCV inhibitor treatments.
Studies of Viral Protein:Host Protein Interactions
NanoBiT® Protein:Protein Interaction Assays
As mentioned earlier, the NanoBiT® Protein:Protein Interaction (PPI) Assay is built upon the LgBiT subunit and the low affinity interacting peptide called SmBiT (6). The interaction of SmBiT with LgBiT will not happen spontaneously but requires assistance to bring the two into close enough proximity to interact which makes the pair ideal for studying PPI. The pair, like HiBiT:LgBiT, form a bright NanoBiT® Luciferase.
Sekiba et al. (13, 14) used the NanoBiT® Assay to find inhibitors of the interaction of the Hepatitis B viral protein, HBx, with the host protein DDB1 in both HEK 293 and HepG2 cells. The assay was used to screen a collection of 800+ FDA-approved compounds and identified 5 compounds offering >40% inhibition of the interaction. The strongest inhibitor, nitazoxanide, was chosen for further study.
Rawle et al. (15) were looking for inhibitors of the HIV-1 reverse transcription complex which includes both viral and host proteins. The focus was on disrupting the viral p66: host eEF1A interaction with inhibitors that bound to the p66 and not the eEF1A. A NanoBiT® PPI Assay was designed in HEK 293T cells to screen >1,200 scaffold for >40% disruption while maintaining >70% cell viability. Nine candidate compounds were identified for further testing. Incidently, Rawle et al also constructed a positive control NanoBiT® PPI pair using Zika virus NS2B and NS3 proteins.
Karposi’s sarcoma-associated herpesvirus (KSHV), also know as human herpes virus 8, has been linked to several cancers in humans. The KSHV latency-associated nuclear antigen (LANA) is critical for establishment of the KSHV latent infection. An important portion of LANA is the host SUMO2 interaction domain. Ding et al. (16) sought small molecule disruptors of the LANA-SUMO2 interaction from a natural products library based on herbal medicines. A NanoBiT® Assay was designed in HEK 293 cells. The screen identified a potent and effective inhibitor from a tropical evergreen tree. The inhibitor blocked primary infection and latency in cell models.
NanoBRET® Protein:Protein Interaction Assays
Bioluminescent resonance energy transfer (BRET) is a commonly used technique to monitor protein:protein interactions. With BRET, energy is transferred from luciferase to a fluorescent acceptor if the pair are in close proximity to one another (<10nm). Many BRET systems use a RLuc-based donor and a fluorescent protein as the acceptor. In the case of a NanoBRET® Assay (17), the bright NLuc is the donor and the acceptor is a fluorescent dye covalently bound to the self-labeling HaloTag® Protein.
Nishi et al. (18) had found that the interaction of the hepatitis B virus core protein (HBc) with the host protein Pin1 was critical for viral pathogenesis. The interaction required phosphorylation of HBc. Curious as to whether a phosphatase existed that could disrupt this interaction, the lab utilized a NanoBRET® Assay in HEK 293 cells to investigate the interaction of NLuc-HBc individually against 150 human phosphatases tagged with HaloTag® (HT) Protein. The screen picked up two candidates and one, pyruvate dehydrogenase phosphatase catalytic subunit, could indeed dephosphorylate HBc.
Miyakawa et al. (19) noted that interaction of the HIV-2 viral protein X (Vpx) with a host protein SMHD1 allowed HIV-2 infections to procede in non-dividing cells. SMHD1 is capable of hydrolyzing all cellular dNTPs making replication in a non-dividing cell difficult. The Vpx:SMHD1 interaction leads to polyubiquitination by a ubiquitin ligase and degradation of SMHD1 by proteosomal degradation. Vpx is known to be a phosphoprotein. The lab found Vpx phosphorylation necessary in the SMHD1 interaction. An in vitro survey of 412 kinases identified 50 candidate kinases for phosphorylating Vpx. These suspected kinases where tested for interaction individually in HEK 293 cels using NLuc-Vpx and each individual HT-kinase. Three kinases gave strong NanoBRET® signals, two of which were members of the PIM kinase family. Further work was performed with PIM 1, 2 and 3. Interestingly, the NanoBRET® Assay showed that a mutant Vpx lacking the serine for PIM phosphorylation had seriously impaired interaction with SMHD1 but retained interaction with a component of a ubiquitin ligase.
NLuc is a small, bright luciferase with a minimal effect on genomic size for many viruses of interest, making it an important virolooy research tool
Conclusions
Virus research is necessary to understand how viral diseases emerge and spread. Recombinant reporter viruses can help researchers uncover the inner workings of viruses. Viruses, however, have evolved minimal genomes to infect, replicate and spread, and do not tolerate extra genomic baggage very well. NLuc is a small, bright luciferase with a minimal effect on genomic size for many viruses of interest, making it an important research tool. The HiBiT bioluminescent protein tag further decreases the burden that must be carried by a recombinant reporter virus and offers a luminescent signal approaching NLuc when paired with its LgBiT binding partner in detection arrangements. The NanoBiT® PPI Assay and NanoBRET® Assays offers a means to study viral:host interactions at the protein level and have been used to screen for inhibitors of interactions and for other host proteins that could influence an interaction.
Table 1. Recombinant Virus or Virus-Like Particles Created with NanoLuc® Luciferase.
References
- Hall, M.P. et al. (2012) Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazopyrazinone substrate. ACS Chem. Biol. 7, 1848–57.
- Mehle, A. (2015) Fiat Luc: Biolumiescence imaging reveals in vivo viral replication dynamics. PLoS Pathog. 11, e1005081. PMID: 26356297.
- Tran, V. et al. (2013) Highly sensitive real-time in vivo imaging of an influenza reporter virus reveals dynamics of replication and spread. J. Virol. 87, 13321–9.
- Karlsson, E.A., et al. (2015) Visualizing real-time influenza virus infection, transmission and protection in ferrets. Nat. Comm. 6, 6378. PMID: 25744559
- Sun, C. et al. (2014) Stable, high-level expression of reporter proteins from improved alphavirus expression vectors to track replication and dissemination during encephalitic and arthritogenic disease. J. Virol. 88, 2035–46.
- Dixon, A.S. et al. (2016) NanoLuc complementation reporter optimized for accurate measurement of protein interactions in cells. ACS Chem. Biol. 11, 400–8.
- Schwinn, M. et al. (2018) CRISPR-mediated tagging of endogenous proteins with a luminescent peptide. ACS Chem. Biol. 13, 467–74.
- Sasaki, M. et al. (2018) Development of a rapid and quantitative method for the analysis of viral entry and release using a NanoLuc luciferase complementation assay. Virus Res. 243, 69–74.
- Yamamoto, M., et al. (2019) Cell-cell fusion and virus-cell fusion assay-based analyses of alanine insertion mutants in the distal α9 portion of JRFL gp41 subunit from HIV-1. J. Biol. Chem. 294, 5677-87.
- Torriani, G., et al. (2019) Identification of clotrimazole derivatives as specific inhibitors of arenavirus fusion. J. Virol. 93, e01744-18.
- Puig-Basagoiti, F. et al. (2016) Human cathelicidin compensates for the role of apolipoproteins in hepatitis C virus infectious particle formation. J. Virol. 90, 8464–77.
- Tamura, T. et al. (2018) Characterization of recombinant Flaviviridae viruses possessing a small reporter tag.J. Virol. 92, e01582–17.
- Sekiba, K., et al. (2019) Inhibition of HBV transcription from cccDNA with nitazoxanide by targeting the HBx-DDB1 interaction. Cell. Mol. Gastroenterol. Hepatol. 7, 297-312.
- Sekiba, K., Otsuka, M.,and Koike, K. (2019) Identifying inhibitors of the HBx-DDB1 interaction using a split luciferase assay system. J. Vis. Exp. 154, e60652.
- Rawle, D.J., et al. (2019) Oxazole-benzenesulfonamide derivatives inhibit HIV-1 reverse transcriptase interaction with cellular eEF1A and reduce viral replication. J. Virol. 93, e00239-19.
- Ding, L., et al. (2019) Identification of viral SIM-SUMO2-interaction inhibitors for treating primary effusion lymphoma. PLoS Pathog. 15, e1008174.
- Machleidt, T., et al. (2015) NanoBRET-A novel BRET platform for the analysis of protein-protein interactions. ACS Chem. Biol. 10, 1797-804.
- Nishi, M., et al. (2020) Prolyl isomerase Pin1 regulates the stability of hepatitis B virus core protein. Front. Cell Dev. Biol. 8, 26.
- Miyakawa, K., et al. (2019) PIM kinases facilitate lentiviral evasion from SAMHD1 restriction via Vpx phosphorylation. Nat. Comm. 10, 1844.
Related Products
NanoBiT® PPI Starter Systems
Detects interactions at low expression levels and allows real-time kinetic analysis in live cells
NanoBiT PPI systems are based on reversible protein complementation reporter technology. These systems can be used to detect interactions at low expression levels and allow real-time kinetic analysis in live cells.
HiBiT Protein Tagging System
HiBiT simplifies protein detection, providing a streamlined, antibody-free protocol for detecting tagged proteins using a convenient, bioluminescence based method.
NanoBRET® PPI Starter Systems
BRET-based technology to detect protein interactions in real time using full-length proteins or fragments. The reversible assay technology allows you to study both induction and inhibition of protein interactions. Assays can be performed in 96- or 384-well formats.
NanoGlo® Luciferase Assay Systems
The NanoGlo® Luciferase Assay System is a single-addition reagent that generates a glow-type signal in the presence of NanoLuc® luciferase with a half-life of approximately 120 minutes in commonly used tissue culture media. The Nano-Glo® Live Cell Assay System is used to measure NanoBiT® or NanoLuc® luminescence from living cells.