Measuring Gene Expression from Mammalian Brain Tissue
Promega Corporation
Publication Date: 1999
Abstract
These experimental procedures cover four major steps from RNA isolation through T-vector cloning of the PCR amplification product. RNA was isolated from mouse and rat brain using the SV Total RNA Isolation System (Cat.# Z3100) and the PolyATtract® System 1000 (Cat.# Z5420). Denaturing agarose gel electrophoresis and Northern blot analysis of isolated RNA was performed. RT-PCR analysis was performed on total RNA from mouse and rat brain, as well as rat poly(A)+ RNA. T-vector cloning of beta-actin RT-PCR product was performed followed by colony screening for positive recombinants. These experiments were originally developed as part of the test experiments for use in a course offered by the BioPharmaceutical Technology Center Institute.
Introduction
The isolation and characterization of nucleic acids entails basic procedures in the molecular biology repertoire. In neurobiology, though, the overwhelming tissue source is brain, which can present special problems such as the presence of high amounts of lipids and the fact that many messages of CNS origin are rare. The ability to easily and effectively obtain nucleic acids---DNA and RNA---from brain tissue is critically important in many neurobiology experiments. In this brief report, it is demonstrated that the isolation of total and messenger RNA (mRNA) from brain tissue can be accomplished with straightforward procedures and reagents, which are available commercially.
RNA Isolation
Total RNA was isolated from mouse and rat brain using the SV Total RNA Isolation System. Poly(A)+ RNA was isolated using the PolyATtract® System 1000. For total RNA isolation, 175µl of SV RNA Lysis Buffer per 60mg of brain were added to a 50ml conical tube. Brain tissue was placed into the tube and homogenized to completely disrupt the tissue. RNA was isolated from 175µl of the generated lysate following the provided spin format SV Total RNA Isolation System protocol (#TM048). Poly (A)+ RNA was isolated using the PolyATtract® System 1000 following the recommended protocol (#TM228). Typical yields and purity are listed in Table 1.
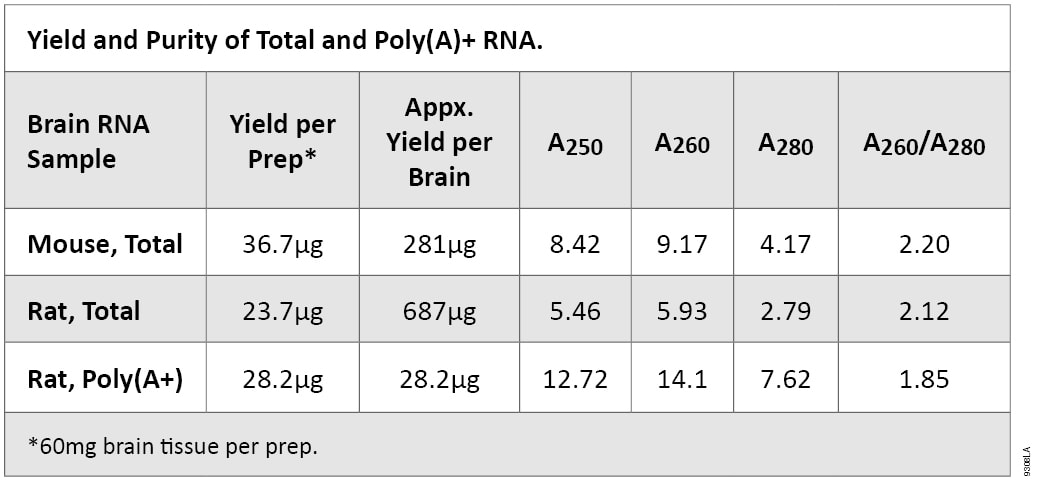 Table 1. Yield and Purity of Total and Poly(A)+ RNA.
Table 1. Yield and Purity of Total and Poly(A)+ RNA. Electrophoretic Analysis of RNA
Mouse and rat total RNA was analyzed by denaturing agarose gel electrophoresis. In this procedure, 2.5µg of RNA was denatured and resolved in an agarose gel containing formaldehyde. After electrophoresis, the resolved RNA samples were visualized by ethidium bromide staining. As evident in Figure 1, Panel A, bands representing ribosomal RNA (rRNA) subunits (18s and 28s) present in the total RNA samples were clearly visible. If the total RNA sample is undegraded, the 28s rRNA subunit band will appear approximately twice as intense as the 18s rRNA subunit band.
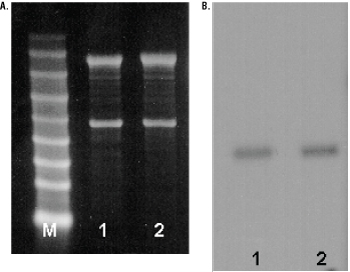 Figure 1. Analysis of total RNA from mouse and rat brains.
Figure 1. Analysis of total RNA from mouse and rat brains.
Panel A. Total RNA was resolved by denaturing agarose gel electrophoresis. Lane 1, mouse total RNA (2.5µg); lane 2, rat total RNA (2.5µg); lane M, RNA Markers, 0.28–6.58kb (Cat.# G3191). Panel B. Northern blot of gel depicted in Panel A blotted and probed for G3PDH mRNA. (Marker lane not shown.)
The RNA was transferred to a membrane by upward capillary transfer and the resulting Northern blot was analyzed for specific message by detection with an RNA probe (Riboprobe® System-T7, Cat.# P1440) for mouse glyceraldehyde-3-phosphate dehydrogenase (G3PDH) mRNA. A specific band of the appropriate size (~1.2kb) was detected, as shown in Figure 1, Panel B.
RT-PCR Analysis of Total and Poly(A)+ RNA
RNA was next analyzed by reverse transcriptase-PCR (RT-PCR) using the Access RT-PCR System (Cat.# A1250). One microgram of mouse total RNA, rat total RNA or 10ng of rat poly(A+) RNA were amplified using β-Actin or NGF Primer Pairs as indicated. (A separate β-actin primer pair [B2] distinct from the β-Actin Primer Pair previously mentioned was also used.) Expected product size was 189bp for the NGF Primer Pair, 285bp for the β-Actin Primer Pair and 511bp for the B2 primer pair.
Reverse transcription was performed for 45 minutes at 48°C. The cDNA amplification program consisted of an initial denaturation step (94°C for 2 minutes) followed by 40 amplification cycles of denaturation (94°C for 30 seconds), annealing (60°C for 1 minute) and extension (68°C for 1 minute). A final extension at 68°C for 7 minutes was included. Equivalent volumes of each amplification reaction were resolved on a 1.5% agarose gel as evident in Figure 2. Negative control reactions that contained NGF or β-Actin primers with no RNA resulted in no product (lanes 7 and 8).
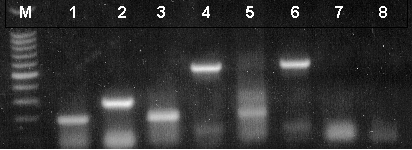 Figure 2. RT-PCR analysis of mouse and rat total RNA and rat poly(A)+ RNA.
Figure 2. RT-PCR analysis of mouse and rat total RNA and rat poly(A)+ RNA.
Lanes 1 and 2, mouse total RNA; lanes 3 and 4, rat total RNA; lanes 5 and 6, rat poly(A)+ RNA, lanes 7 and 8, no RNA. β-Actin Primer Pair was used for reactions represented in lanes 2 and 8. NGF Primer Pair was used for reactions represented in lanes 1, 3, 5 and 7. β-actin [B2] primer pair was used for reactions represented in lanes 4 and 6 (511bp expected product). Lane M, Promega 100bp DNA Ladder (Cat.# G2101).
Cloning PCR products in pGEM®-T Easy Vector
T-vector cloning was performed using the pGEM®-T Easy Vector System (Cat.# A1380) and mouse RT-PCR product generated using the β-Actin Primer Pair (reaction represented in lane 2 of Figure 2). Ligation reactions of the PCR product and pGEM®-T Easy Vector were transformed into JM109 competent bacteria and plated onto LB plates containing ampicillin (100µg/ml), IPTG (0.5mM) and X-gal (80µg/ml) for blue-white color screening. After overnight growth at 37°C, blue, pale blue and white colonies (presumed to contain β-actin insert) were rapidly screened by PCR using Tfl DNA Polymerase (Cat.# M1941). Each colony was picked and placed into 30µl of LB broth + ampicillin. Two microliters of each culture were then placed directly into PCR mixes that contained the β-actin primer pairs and amplified for 30 cycles under the following conditions: initial denaturation at 94°C for 2 minutes, followed by 30 cycles of denaturation (94°C for 30 seconds), annealing (60°C for 1 minute) and extension (68°C for 1 minute). A final extension at 68°C for 7 minutes was included. Equivalent volumes of products were resolved on a 2% agarose gel and visualized by ethidium bromide staining.
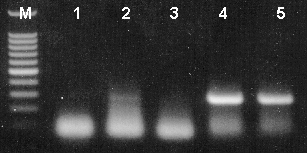 Figure 3. Rapid screening of T-vector cloning recombinants by PCR amplification.
Figure 3. Rapid screening of T-vector cloning recombinants by PCR amplification.
Lane 1, blue colony; lanes 2 and 3, pale blue colonies; lanes 4 and 5, white colonies (285bp expected product). Lane M, Promega 100bp DNA Ladder (Cat.# G2101).
Reaction setup for PCR and RT-PCR
| Component | Volume |
| Sample in water | 32µl |
| Downstream Primer | 1µl |
| Upstream Primer | 1µl |
| AMV/Tfl 5X Buffer | 10µl |
| dNTP Mix (10mM each) | 1µl |
| 25mM MgSO4 | 2µl |
| Tfl DNA Polymerase | 2µl |
| *For RT-PCR, add AMV RT | 1µl |
| Total | 50µl |
Conclusions
These experiments demonstrate the integration of many core Promega products for the isolation and analysis of nucleic acids in neurobiology. The protocols used were as recommended in the Technical Bulletins or Manuals for each product. The reagent amounts for RT-PCR are listed below. These procedures have been used successfully in the training course, Molecular Approaches to Neuroscience.
Related Resources
LabFact #2
RT-PCR should be performed in the absence as well as presence of the reverse transcriptase, to assess DNA contamination in the template RNA. In addition, "no-template" negative control reactions should always be performed.
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Kephart, D. Measuring Gene Expression from Mammalian Brain Tissue. [Internet] 1999. [cited: year, month, date]. Available from: https://www.promega.com/resources/pubhub/enotes/measuring-gene-expression-from-mammalian-brain-tissue/
American Medical Association, Manual of Style, 10th edition, 2007
Kephart, D. Measuring Gene Expression from Mammalian Brain Tissue. Promega Corporation Web site. https://www.promega.com/resources/pubhub/enotes/measuring-gene-expression-from-mammalian-brain-tissue/ Updated 1999. Accessed Month Day, Year.
pGEM, PolyATtract and Riboprobe are registered trademarks of Promega Corporation.
Products may be covered by pending or issued patents or may have certain limitations. Please visit our Web site for more information.
 T-vector cloning of PCR products made easy.
T-vector cloning of PCR products made easy.