Screening and Profiling Kinase Inhibitors with a Luminescent ADP Detection Platform
Hicham Zegzouti, Juliano Alves, Tracy Worzella, Gediminas Vidugiris, Gregg Cameron, Jolanta Vidugiriene and Said Goueli
Promega Corporation
Publication Date: 2011
Abstract
Selectivity of a kinase inhibitor towards the target of interest is critical for its advancement through the various phases of drug development. This makes kinase profiling of the inhibitor a common practice during the drug discovery phase. A robust and universal kinase assay is desired at this phase to assess inhibitor selectivity and potency on multiple kinases from different classes that often use substrates with different chemical structures. Here we describe a kinase assay platform that uses the universal ADP-Glo™ technology to attain the sensitivity and reproducibility required for kinase inhibitor screening and profiling. Using an improved ultrapure ATP, the sensitivity of the ADP-Glo™ Kinase Assay is significantly enhanced, reducing the amount of enzyme needed in the kinase assay. The ADP-Glo™ technology is optimized for use with a large panel (70 kinases) of Kinase Enzyme Systems (KES) that represent different families of the human kinome. Here we present selectivity profiles of four specific kinase inhibitors, demonstrating that the ADP-Glo™ Kinase Assay provides a universal platform ideal for screening diverse kinase targets and profiling inhibitors against small or large kinase panels.
Introduction
Kinases have become one of the most important and widely studied families of enzymes in biochemical and medical research. Kinases are a class of phosphotransferases that use ATP to catalyze the transfer of phosphate to a wide variety of cellular substrates. Many cellular processes such as cell signaling, cell division and growth, development, differentiation and cell death are orchestrated by kinases (1,2). Under normal physiological conditions, the regulation of kinases is tightly regulated. However, under pathological conditions, kinase activities can be dysregulated, and the disruption of the intracellular signaling networks governed by these enzymes leads to many diseases including cancer and inflammation (3–6). Consequently, kinases have become an important target for drug discovery, resulting in successful kinase inhibitor-based drugs, such as Gleevec® (imatinib) and Tarceva® (erlotinib) (7,8).
Given the importance of kinase inhibitors as potential therapeutics, researchers have focused on developing technologies that monitor changes in kinase activities and their modulation by small-molecule inhibitors (9). Isotopic-based methods for measuring kinase activity have long been considered the gold standard. However, due to safety concerns and waste management issues, alternative approaches have been sought (9,10) . The ideal kinase assay platform should be able to: 1) detect activity of many enzyme classes, 2) use universal substrates, 3) detect activity at low substrate conversion rates, 4) have a large dynamic range and low data variability, 5) be amenable to high-throughput applications and 6) reduce false-positive results. With these goals in mind, we have developed a luminescence-based ADP detection assay, ADP-Glo™ Kinase Assay, that measures kinase activity by quantifying the amount of ADP produced in the kinase reaction (11). This assay can be used to monitor the activity of virtually any ADP-generating enzyme (e.g., kinase or ATPase). Since the ADP-Glo™ Kinase Assay is universal and can be used with any kinase/substrate combination (11–13), we applied it to screening compounds for kinase inhibitor activity and creating inhibitor selectivity profiles for several kinase families spanning the human kinome.
ADP-Glo™ Kinase Assay Fundamentals
The ADP-Glo™ Kinase Assay (Cat.# V9101, V9102, V9103) is performed in two steps. First, after the kinase reaction, an ATP-depletion reagent is added to terminate the kinase reaction and deplete any remaining ATP, leaving only ADP. Second, a detection reagent is added to simultaneously convert ADP to ATP and allow the newly synthesized ATP to be converted to light using a coupled luciferase/luciferin reaction (Figure 1). The luminescent signal generated is proportional to the ADP concentration produced and is correlated with the kinase activity (11). The assay can be performed over a wide range of ATP concentrations (low micromolar to millimolar). This allows detection of small concentrations of ADP in the presence of large amounts of ATP (Figure 2, Panel A), producing very high signal-to-background (SB) ratios (Figure 2, Panel B, and Table 1). The robustness of the ADP-Glo™ Kinase Assay and suitability for high-throughput applications is evidenced by high Z´-factor values reported in previous studies (13).
The ADP-Glo™ Kinase Assay is as sensitive as radioactivity-based methods and more sensitive than fluorescence-based technologies (11–13). In order to lower the background and further improve the sensitivity of the assay, we increased the purity of our ATP to have less ADP contamination. To assess the importance of ATP purity on ADP-Glo™ Kinase Assay sensitivity, we compared the signal-to-background ratios generated in an ADP-Glo™ Kinase Assay using Promega Ultra Pure ATP and ATP from other suppliers. Table 1 shows that the Promega ATP outperforms ATP from other sources (S, T and G) by greatly improving ADP-Glo™ Kinase Assay sensitivity with SB ratios that are 2–3 times higher than those produced using other commercial preparations.
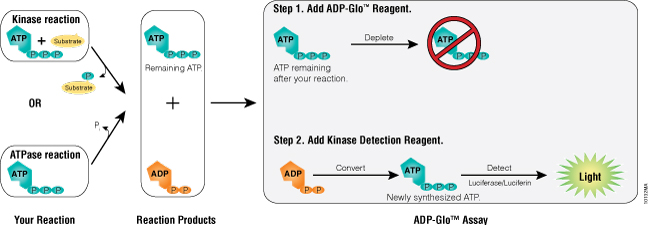
Figure 1. ADP-Glo™ Kinase Assay principle. The assay is composed of two steps. After the kinase or ATPase reaction, the first step is performed by addition of the ADP-Glo™ Reagent that terminates the kinase reaction and depletes any remaining ATP (40-minute incubation time). Addition of a second reagent converts ADP to ATP and generates light from the newly synthesized ATP using a luciferase/luciferin reaction (incubation is 30–60 minutes depending on the ATP concentration used in the kinase reaction). The light generated is proportional to ADP present and, consequently, kinase or ATPase activity. The assay is performed at room temperature and is compatible with automation.
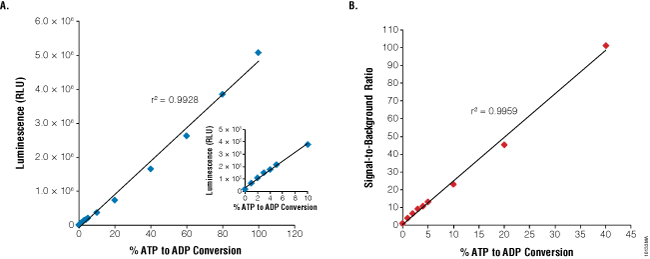
Figure 2. Linearity and sensitivity of the ADP-Glo™ Kinase Assay. The 1mM ATP-to-ADP percent conversion curve (standard curve) was prepared in 1X reaction buffer A (40mM Tris [pH 7.5], 20mM MgCl2, and 0.1mg/ml BSA) without kinase present as described in Technical Manual #TM313. The standards were created by combining the appropriate volumes of ATP and ADP 1mM stock solutions. Five microliters of each ATP+ADP standard was transferred to a white, opaque 384-well plate. The ADP-Glo™ Kinase Assay was performed by adding 5μl of ADP-Glo™ Reagent and 10μl of Kinase Detection Reagent at room temperature to each well. ADP-Glo™ Kinase Assay reagents were dispensed in 384-well plates using Multidrop® Combi nL liquid dispenser (Thermo Fisher Scientific) for this experiment and all subsequent experiments described here. Luminescence values represent the mean of four replicates (RLU = relative light units). Panel A. Linearity of the assay up to 1mM ADP. Panel B. Sensitivity of the assay is shown as signal-to-background ratios (SB) over a wide range of % ATP-to-ADP conversion.
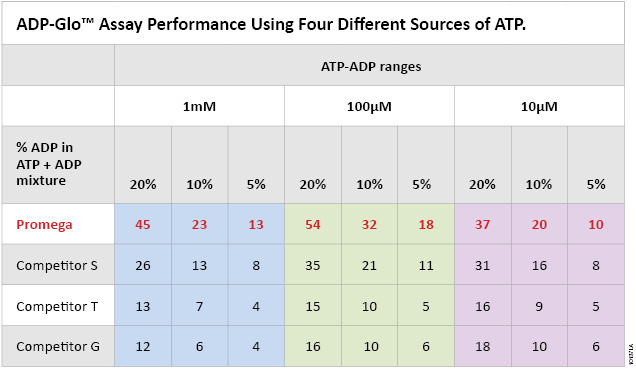
Table 1. ADP-Glo™ Kinase Assay performance using four different sources of ATP. Each ATP-to-ADP percent conversion was prepared in 1X reaction buffer A (40mM Tris, [pH 7.5], 20mM MgCl2, and 0.1mg/ml BSA) at the indicated ATP-ADP concentration ranges (10µM, 100µM and 1,000µM) by mixing the indicated percentages of ADP to ATP to prepare 100% total nucleotide. Five microliters of each % conversion mix and the 0% control consisting of 100% ATP were dispensed in a white, opaque 384-well plate. The ADP-Glo™ Kinase Assay was performed by adding 5μl of ADP-Glo™ Reagent and 10μl of Kinase Detection Reagent at room temperature to each well as described in Technical Manual #TM313. Sensitivity of the Promega Ultra Pure ATP compared to the other ATP suppliers (G, T and S) is shown as signal-to-background ratios of the assay at each % ATP-to-ADP conversion level.
Screening Kinase Inhibitors and Compound Interference
We wanted to ensure that screening chemical libraries with the ADP-Glo™ Kinase Assay would not generate a high rate of compound interference and, subsequently, false responses, since multiple enzymes are used as components of the ADP-Glo™ Kinase Assay. To address this issue, we performed two small-scale screenings of the Library of Pharmacologically Active Compounds (LOPAC, which contains 1,280 compounds) at 10µM each, using two kinases, epidermal growth factor receptor (EGFR; Cat.# V3831) and lymphocyte-specific protein tyrosine kinase (LCK; Cat.# V2691). For comparison, an additional LOPAC screen was performed with the ADP-Glo™ reagents using a mixture of 20% ADP and 80% ATP at 30µM total nucleotide concentration without EGFR or LCK present. The results show that only the specific inhibitory effects of the compounds on both kinases can be detected using this assay (Figure 3, Panels B and C), while no effect was observed on the performance of the ADP-Glo™ Kinase Assay in the presence of the LOPAC chemical compounds (Figure 3, Panel A). Using the ADP-Glo™ Kinase Assay to screen for kinase inhibitors generates specific compound-inhibitory data with less compound interference. These data also confirm what has been shown by others regarding the reproducibility and resistance to chemical interference of the combination of Ultra-Glo® Recombinant Luciferase used in the ADP-Glo™ assay and the formulations of the assay reagents (14,15).
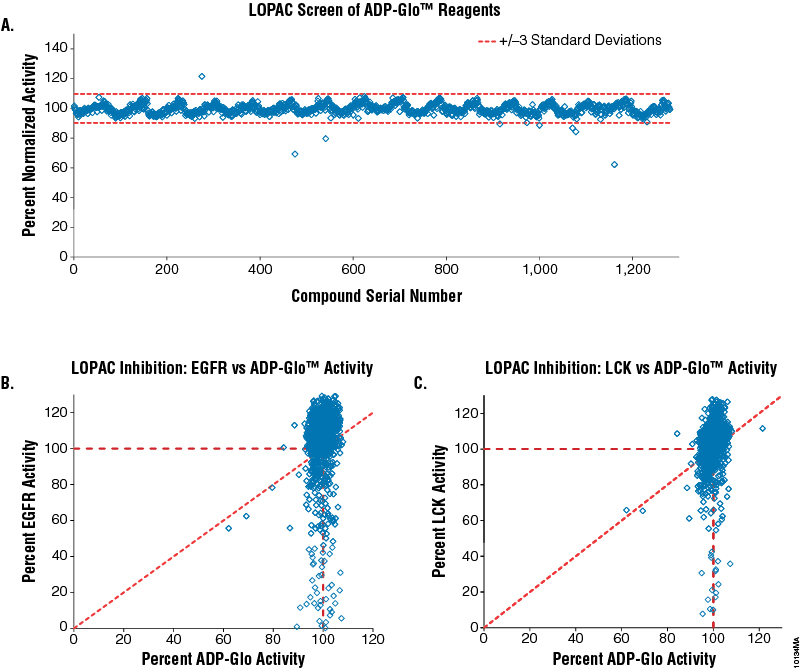
Figure 3. Screening the LOPAC library for kinase inhibitors and compound interference in the ADP-Glo™ Kinase Assay. Five nanoliters of each of the 1,280 compounds of the LOPAC library were dispensed (EDC Biosystems ATS Acoustic Liquid Dispenser) in 384-well plates (total of 16 plates) at 10mM concentration (final concentration will be 10µM) in four replicates. Panel A. The effect of the LOPAC compounds on the ADP-Glo™ reagents was tested by assaying a solution of 20% ADP in a 30µM ADP-ATP mixture in the presence of the compounds (no kinase present). As a control, 30µM ATP was used as 0% activity, and any signal subtracted to normalize each compound result. Panel B. Correlation between LOPAC inhibition of EGFR kinase (Cat.# V3831) and ADP-Glo™ Kinase Assay activities. Panel C. Correlation between LOPAC inhibition of LCK kinase (Cat.# V2691) and ADP-Glo™ Kinase Assay activities. Both kinase reactions were performed in 1X reaction buffer A supplemented with 50µM DTT and 2mM MnCl2 as described in their respective protocols (Product Information #9PIV383 and #9PIV269) with the following exceptions: 2.5μl of the kinase in buffer was added first to the compounds in the plates and incubated for 20 minutes, followed with the addition of ATP/substrate solution to initiate the reaction. The EGFR reaction contained 0.2µg/µl PolyE4Y1 (Glu4, Tyr1) substrate, 3.75ng enzyme and 5µM ATP. The LCK reaction contained 0.2µg/µl PolyE4Y1 (Glu4, Tyr1) substrate, 1.5ng enzyme and 30µM ATP. Both reactions were incubated for 60 minutes. Positive controls contained no compound and were used to calculate 100% kinase activity. Negative controls were used to calculate 0% kinase activity since they contained neither compound nor enzyme. The ADP-Glo™ Kinase Assay reagents were dispensed according to Technical Manual #TM313.
Selectivity Profiling of Kinase Inhibitors
The universal product of all kinase reactions is ADP. This makes the ADP-Glo™ Kinase Assay applicable to a variety of kinase reactions regardless of the nature of their substrates (peptides, proteins, alcohols, lipids and sugars) with no need for substrate modification. We have shown that the assay detects the activity of several kinases with high sensitivity and can be used to generate small selectivity profiles of compounds using kinases from different families (11,16). To expand upon this application using a larger set of kinases, we optimized the ADP-Glo™ Kinase Assay protocol for 70 Kinase Enzyme Systems (KES) that represent all families of the human protein kinome. Individual KES available from Promega contain a purified kinase, substrate, reaction buffer and reaction buffer components. The data described in Table 2 show that all kinases were detected with high sensitivity. This demonstrates that the ADP-Glo™ Kinase Assay is ideal for performing kinase assays using low amounts of enzymes and shorter enzyme reaction times (e.g., <1 hour). These features are critical for accurate determination of kinase inhibitor IC50 values.
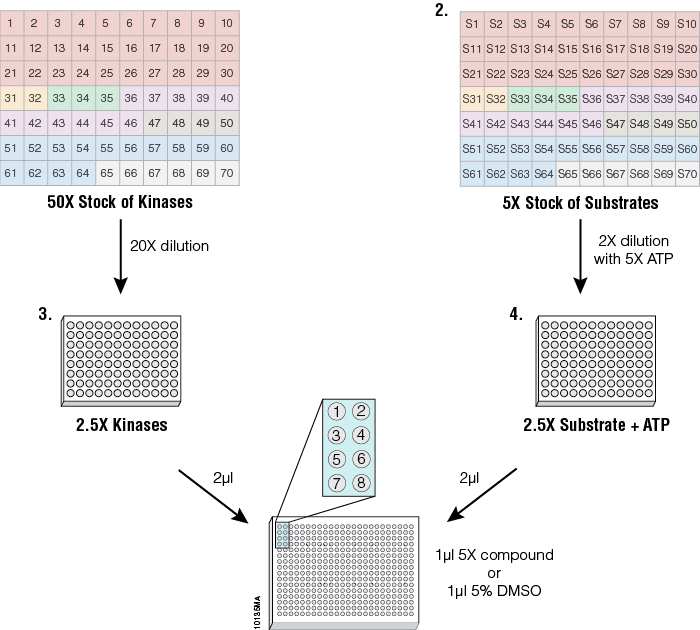
Figure 4. Streamlined profiling protocol using the ADP-Glo™ Kinase Assay platform. To standardize kinase performance, the 70 kinases were diluted in a storage buffer (50mM Tris [pH 7.5], 150mM NaCl, 0.25mM DTT and 25% glycerol) to 50 times the concentration that generates a 10-fold change between the control (0% activity) and signal (100% activity) (Plate 1) and dispensed in a 96-well plate. Substrates (5X) were dispensed in a separate 96-well plate at the corresponding positions to their kinase (Plate 2). Starting from the concentrated stocks, kinases were diluted 20-fold in 1X reaction buffer, as described in the corresponding KES product information sheet, (Plate 3) to generate 2.5X concentrated kinase solutions. The substrate stock solutions were diluted 2-fold into 1X reaction buffer containing 5X ATP (Plate 4) to generate 2.5X concentrated ATP/substrate solutions. The inset of wells 1–8 represents the position where the different reactions and controls are arranged in the 384-well plates for a single kinase. Eight wells were used per enzyme: wells 5–8 contained either replicates of two different concentrations of one compound or replicates of two compounds at a single concentration, wells 3 and 4 contained replicates of the enzyme reaction with no compounds, wells 1 and 2 contained replicates of ATP only with no enzyme and no compound to act as a negative control. One microliter of 5X compound or 5% DMSO in a 1X buffer were arrayed in 384-well plates using the Multidrop® Combi nL liquid dispenser. Then 2µl of 2.5X kinase solution was added to all compound wells. The kinase solution also was added to positive control wells containing DMSO only (signal generated from these wells corresponded to 100% activity), leaving negative control wells where only buffer was added (signal generated from these wells corresponded to 0% activity). After a 20-minute incubation, the kinase reaction was started by adding 2µl of the ATP/substrate solutions to every well. After a brief mixing (30–60 seconds) the plates were incubated for 1 hour at 23°C. After the kinase reaction, the ADP-Glo™ Kinase Assay reagents were dispensed according to Technical Manual #TM313 for a 384-well plate. Enzyme and substrate solutions were transferred from 96-well plates to 384-well assay plates either manually or using the Biomek® FX Liquid Handling System (Beckman Coulter).
Four inhibitors were profiled against the 70 kinases using the ADP-Glo™ Kinase Assay (Figure 5) and the streamlined profiling protocol (Figure 4). Gefitinib is a specific EGFR inhibitor, Lapatinib is a HER kinase subfamily inhibitor, SB203580 is p38 MAP kinase inhibitor, and GDC-0941 is an inhibitor of PI3 Kinase. All four compounds showed a strong inhibitory effect against their corresponding kinase, while the other kinases were not inhibited. These data show that, as a universal assay, the ADP-Glo™ Kinase Assay is ideal for profiling inhibitors against large kinase panels for drug discovery.
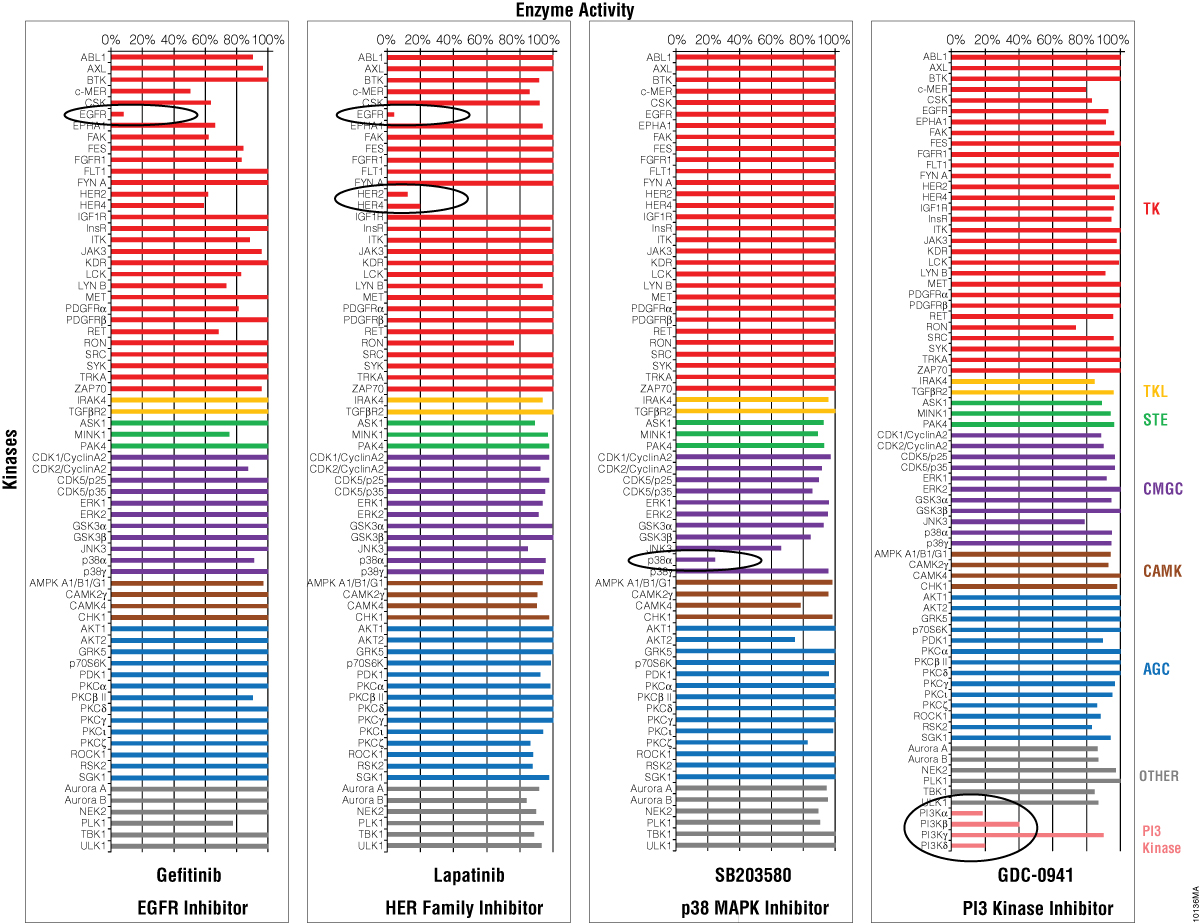
Figure 5. Selectivity profiles of four kinase inhibitors. Four kinase inhibitors were profiled against 70 kinases (color coding corresponds to kinase families found at Promega Human Kinome Map for KES products) following the streamlined protocol described in Figure 4. The GDC-0941 compound was screened against additional PI3 kinases. The luminescent signals generated from the ADP-Glo™ Kinase Assay were converted to percent activity by subtracting the values of the negative control (0%) from all points and using the enzyme-only control as the 100% activity point.
Conclusion
Although several kinase assay technologies have been developed in the last few years, most suffer from limitations that make it difficult to perform 1) kinase inhibitor screening, 2) compound mode of action (MOA) studies, and 3) kinase selectivity profiling using a single platform during the drug discovery process. The ADP-Glo™ Kinase Assay already has been tested for inhibitor screening and inhibitor mode of action studies (15,17,18). Here we demonstrate that a single kinase assay platform can address all the needs of kinase research over a wide range of kinase families as demonstrated with the Promega Kinase Enzyme Systems.
The ADP-Glo™ Kinase Assay can be used to detect the activity of kinases with high ATP Km values based on the assay’s ability to generate high signal strength at low ATP conversion rates and a tolerance for high ATP concentrations (up to 1mM). These assay features are ideal for primary and secondary compound screening as well as profiling of lead therapeutic candidates with kinase inhibitor activity.
References
- Hunter, T. (2000) Signaling-2000 and beyond. Cell 100, 113–27.
- Manning, G. et al. (2002) The protein kinase complement of the human genome. Science 298, 1912–34.
- Mazitschek, R. et al. (2004) Inhibitors of angiogenesis and cancer related receptor tyrosine kinases. Curr. Opin. Chem. Biol. 8, 432–41.
- Resnick, L. et al. (2004) Targeting JNK3 for the treatment of neurodegenerative disorders. Drug Discov. Today 9, 932–9.
- Fabbro, D. et al. (2004) Targeting protein kinases in cancer therapy. Curr. Opin. Drug Discov. Dev. 5, 701–12.
- Myers, M. R. et al. (1997) Inhibitors of tyrosine kinases involved in inflammation and autoimmune disease. Curr. Pharm. Des. 3, 473–502.
- Cohen, P. (2002) Protein kinases—the major drug targets of the twenty-first century? Nat. Rev. Drug Discov. 1, 309–15.
- Zhang, J. et al. (2009) Targeting cancer with small molecule kinase inhibitors. Nat. Rev. Drug Discov. 9, 28–39.
- Jia, Y. et al. (2008) Current in vitro kinase assay technologies: The quest for a universal format. Curr. Drug Discov. Technol. 5, 59–69.
- Ma, H. et al. (2008) The challenge of selecting protein kinase assays for lead discovery optimization. Expert Opin. Drug Discov. 3, 607–21.
- Zegzouti, H. et al. (2009) ADP-Glo: A bioluminescent and homogeneous ADP monitoring assay for kinases. Assay Drug Dev. Technol. 7, 560–72.
- Vidugiriene, J. et al. (2009) Evaluating the utility of a bioluminescent ADP detecting assay for lipid kinases. Assay Drug Dev Technol. 7, 585–97.
- Tai, A.W. et al. (2011) A homogeneous and nonisotopic assay for phosphatidylinositol 4-kinases. Anal. Biochem. 417, 97–102.
- Auld, D.S. et al. (2009) A basis for reduced chemical library inhibition of firefly luciferase obtained from directed evolution. J. Med. Chem. 52, 1450–8.
- Tanega, C. et al. (2009) Comparison of bioluminescent kinase assays using substrate depletion and product formation. Assay Drug. Dev. Technol. 7, 606–14.
- Larson, B. et al. (2009) A simple and robust automated kinase profiling platform using luminescent ADP accumulation technology. Assay Drug Dev. Technol. 7, 573–584.
- Li, H. et al. (2009) Evaluation of an antibody-free ADP detection assay: ADP-Glo. Assay Drug Dev. Technol. 7, 598–605.
- Santaguida, S. et al. (2011) Evidence that Aurora B is implicated in spindle checkpoint signaling independently of error correction. J. EMBO 30, 1508–19.