Sample Preparation Method for Accurate Analysis of Nonenzymatic PTMs in Biotherapeutic Proteins
Gary Kobs
Promega Corporation
Publication Date: August 2017
Abstract
Introduction
Nonenzymatic posttranslational modifications (PTMs) can spontaneously occur in biotherapeutic proteins during manufacturing and storage, affecting efficacy and stability of these proteins. The major nonenzymatic PTMs are deamidation, disulfide bond scrambling and oxidation (Figure 1).

Peptide mapping is the primary analytical tool used to monitor nonenzymatic PTMs and other protein primary structural features (Figure 2).

Unfortunately, the sample preparation steps involved in peptide mapping are also sources of these modifications. Alkaline pH, used during all steps of sample preparation, induces deamidation and disulfide bond scrambling. These sample preparation-induced modifications are particularly prominent during the proteolytic step of the peptide mapping process.
Artificial PTMs compromise the analysis of authentic nonenzymatic PTMs. The problem can be resolved by shifting the pH of the sample preparation steps into the acidic range. The chemical agents and proteases involved in the sample preparation process, however require alkaline pH.
Development of a kit that provided reproducible reduction, alkylation and digestion under low pH conditions required optimization of reaction conditions as well as the selection of corresponding reagents. Details relating to the development process are described here.

Methods and Results
Reduction and Alkylation at Low pH
Common reducing and alkylating agents favor alkaline pH. Because alkaline pH induces deamidation and disulfide bond scrambling, we modified the reduction and alkylation procedure to be compatible with low pH. With the AccuMAP® Kit, reduction and alkylation steps are performed at pH 5.6–5.8. To ensure efficient reduction at low pH we used TCEP (Tris(2-carboxyethyl)phosphine), which maintains high reducing activity at low pH. Alkylation is performed with IAM (iodoacetamide). Activity of this reagent is decreased at low pH. To compensate for decreased IAM activity at low pH, alkylation is allowed to proceed through the digestion reaction.
Protein Digestion at Low pH
Trypsin and other proteases commonly used in peptide mapping sample preparation favor alkaline pH to efficiently digest proteins. To avoid artificial nonenzymatic PTMs induced at these conditions, we developed protocols for trypsin digestion at low pH. (With the AccuMAP® Kit pH is shifted to 5.2–5.4 at the digestion step.) Our studies show that low pH primarily inhibits tryptic cleavages at lysine sites whereas arginine cleavage sites are still efficiently digested. To restore trypsin cleavage efficiency at lysine sites, we supplemented trypsin with a special, low pH resistant recombinant Lys-C (rLys-C) protease. By supplementing trypsin with low pH resistant rLys-C, we achieved efficient tryptic digestion at low pH (Figure 4).

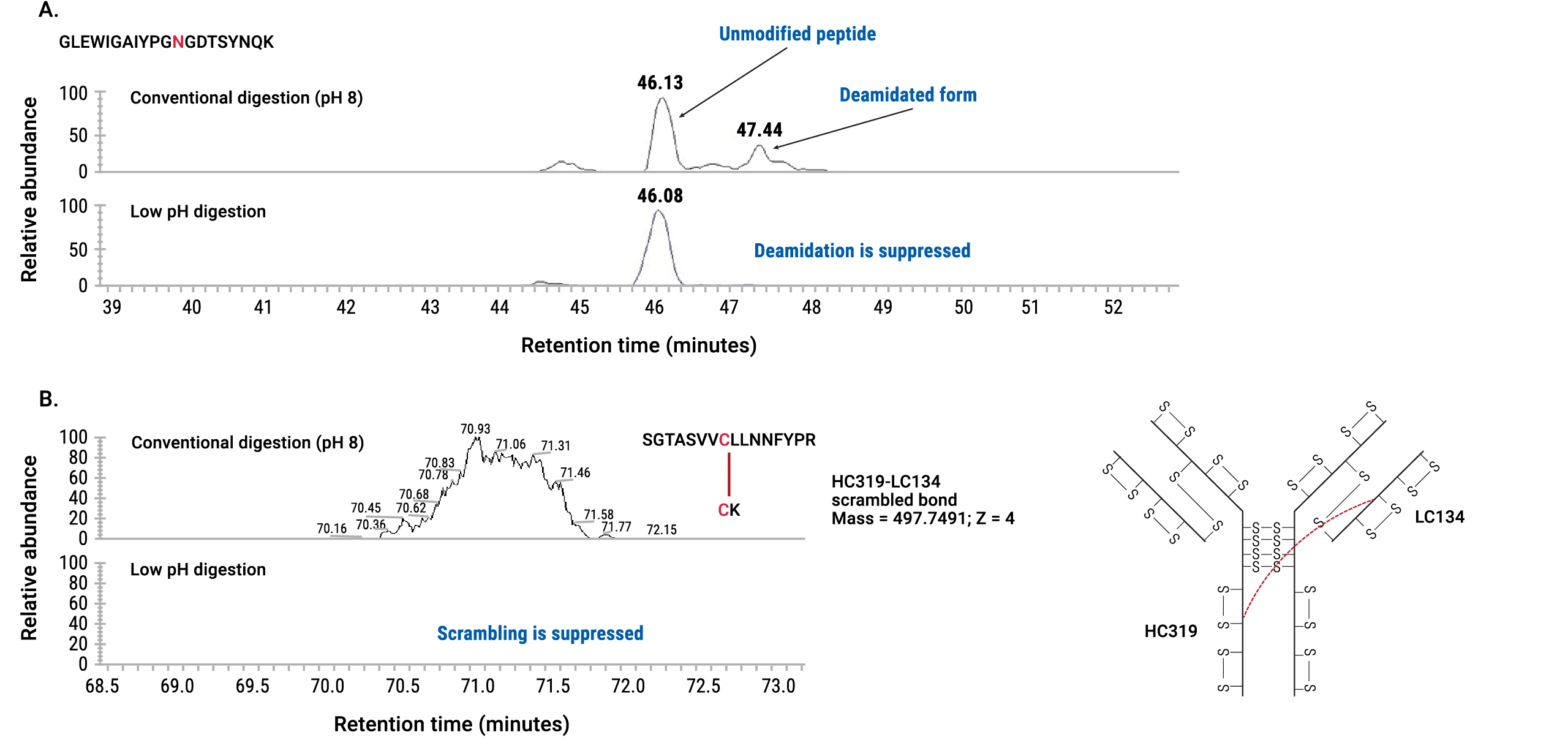
Under the conditions established in this protocol, artificial deamidation and disulfide bond scrambling were completely suppressed (Figure 5).
Predigestion Step
Often, certain protein regions are resistant to proteolysis even after reducing disulfide bonds. This resistance is caused by the tightly folded conformation of these regions. Tight folding prevents protease access to internal cleavage sites. A common approach to overcome proteolytic resistance of tightly folded proteins or protein domains is predigestion with Lys-C protease under denaturing conditions. Digestion under denaturing conditions is possible due to the unique ability of Lys-C proteases to tolerate denaturing conditions. We used this tolerance to denaturing conditions to ensure efficient digestion of proteolytically resistant domains. As with all other steps in this protocol, predigestion is performed at low pH. Subsequently a denaturing agent is diluted and digestion is completed with trypsin or, optionally, rLys-C alone.
Reduction of Baseline Noise in Tryptic Digests
Baseline noise is a common problem in tryptic digests. The major cause of baseline noise is trypsin overdigestion, which results in the generation of semi-tryptic peptides. Semi-tryptic peptides are the products of specific cleavage at lysine or arginine residue at one terminus and nonspecific cleavage (primarily at an aromatic residue) at the other terminus.
Semi-tryptic peptides were evident in overnight trypsin digestions at 1:20 or lower trypsin:protein ratio in our study. We offer two potential solutions to this problem. One solution is to remove trypsin from a reaction and digest a protein with the AccuMAP® Low pH Resistant rLys-C alone. In contrast to trypsin, AccuMAP® Low pH Resistant rLys-C does not generate semi-tryptic peptides.
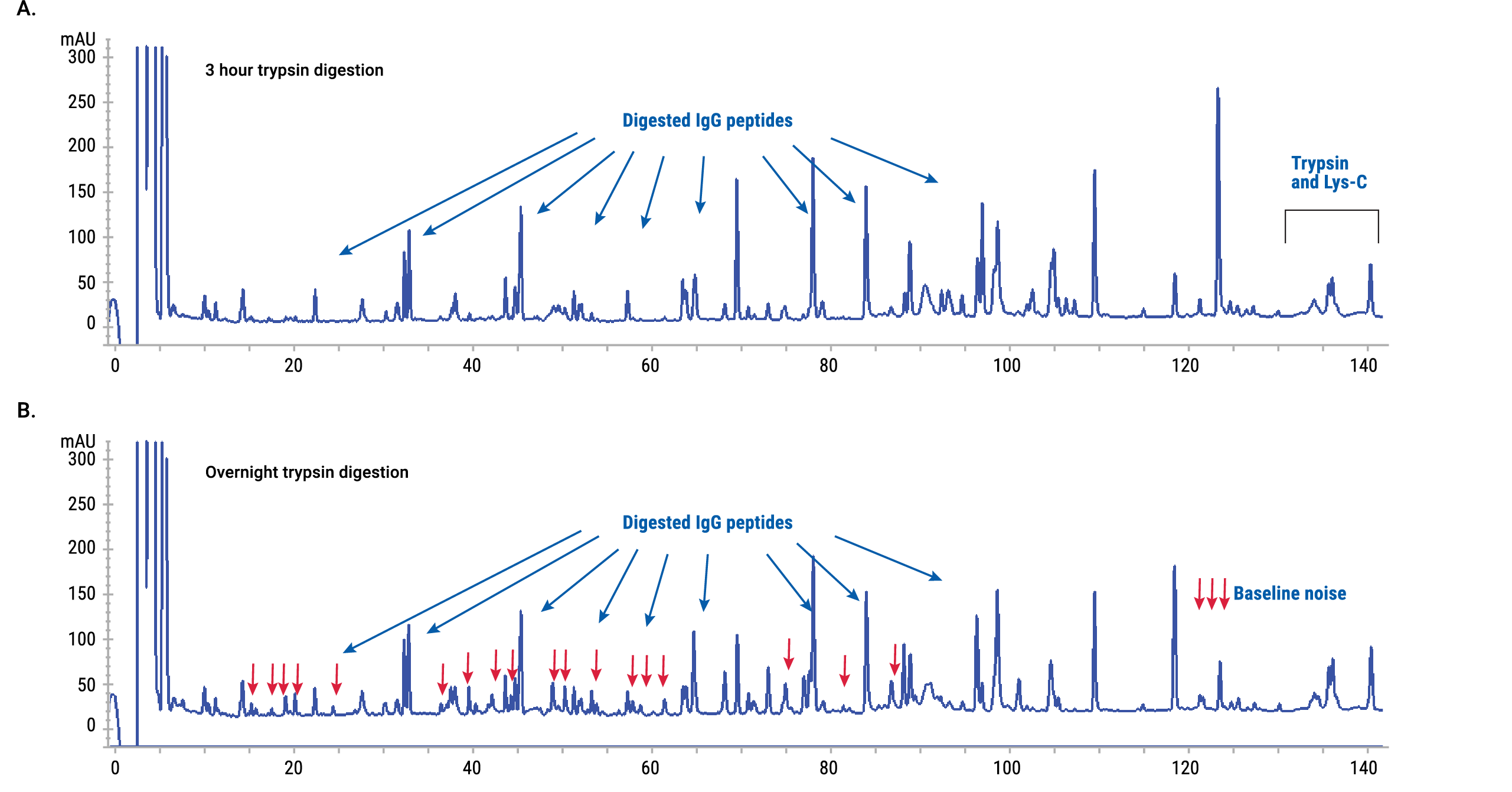
An alternative solution is based on optimization of the trypsin:protein ratio and digestion time. Optimal digestion was achieved at a 1:5 trypsin:protein ratio and 3-hour incubation. Baseline noise was minimal and digestion was efficient at these conditions (Figure 6).
Oxidation Suppression
Certain excipients and impurities have protein oxidation activity. For example, polysorbates, which are popular surfactants used to improve solubility of biotherapeutic proteins, decompose upon storage into hydrogen peroxide, which oxidizes proteins. This oxidation increases during proteolytic digestion due to the increased temperature. To suppress oxidation we recommend adding the optional AccuMAP® Oxidation Suppressant (included in the kit) to a reaction.
Conclusion
Nonenzymatic PTMs can be introduced during protein sample preparation for peptide mapping under conventional conditions. The AccuMAP® Low pH Protein Digestion Kit is designed to suppress artificial nonenzymatic PTMs. The entire procedure (i.e., reduction, alkylation and digestion) has been optimized for use at low pH. The digestion procedure is based on the use of trypsin and Low pH Resistant rLys-C. The combination of these two proteases provides maximum sequence coverage while minimizing baseline noise.
Learn more about the AccuMAP® Low pH Protein Digestion Kit