Removal of Ethidium Bromide and Calf Intestinal Alkaline Phosphatase Using the Wizard® SV Gel and PCR Clean-Up System
Megan Buros and Natalie Betz
Promega Corporation
Publication Date: 2002
Abstract
Introduction
Methods
Removal of Calf Intestinal Alkaline Phosphatase: One unit of Calf Intestinal Alkaline Phosphatase (Cat.# M1821) was added to 50µl of a 500bp PCR product. Replicate samples of the PCR product with or without CIAP were purified using the Wizard® SV Gel and PCR Clean-Up System or by phenol/chloroform extraction followed by ethanol precipitation. For the Wizard® System, the DNA was eluted with 50µl of Nuclease-Free Water (Cat.# P1193). For the phenol/chloroform extraction and ethanol precipitation, the DNA pellet was resuspended in 50µl Nuclease-Free Water. The unpurified PCR product without CIAP, unpurified PCR product with CIAP, or purified PCR products were then ligated into the pGEM®-T Easy Vector using the pGEM®-T Easy Vector System (Cat.# A1360). Ligation of purified PCR products were performed in triplicate, as described in the pGEM®-T Easy Vector System Technical Manual #TM042. Chemically competent JM109 cells were transformed with 2µl of the ligation reaction and plated onto LB plates containing IPTG, X-gal, and 125µg/ml ampicillin. After overnight growth at 37°C, colonies were counted. Light blue colonies were counted as white colonies since both types were determined to have insert DNA.
Removal of Ethidium Bromide: In a separate experiment, 5µl of Ethidium Bromide (Cat.# H5041) and 20µl of water were added to 25µl of the 1kb DNA Ladder (Cat.# G5711). Two replicates of 50µl each were purified using the Wizard® SV Gel and PCR Clean-Up System as described in the Wizard® SV Gel and PCR Clean-Up System Technical Manual #TB308. Equivalent volumes of the 1kb DNA Ladder before and after purification were separated on a 2% agarose gel and visualized on a UV transilluminator. Photographs were taken before and after ethidium bromide staining of the gel.
Results
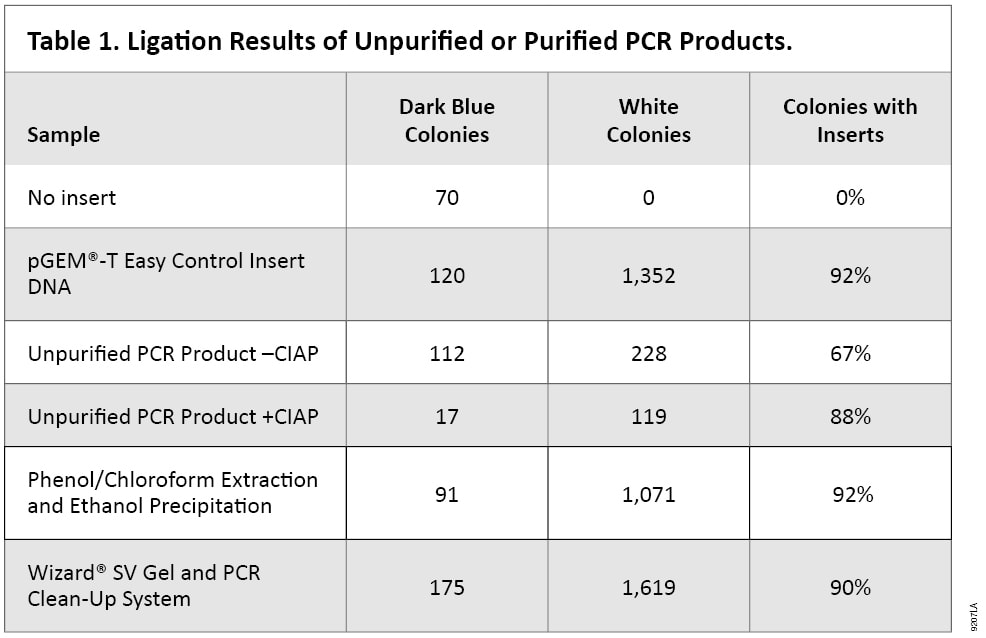
Table 1 shows the results of experiments designed to examine the effectiveness of CIAP removal. Purified or unpurified PCR products were ligated into the pGEM®-T Easy Vector. A ligation with vector but no PCR product was performed as a negative control, and the Control Insert DNA supplied with the pGEM®-T Easy Vector System was used as a positive control for ligation. The number of white colonies, which represent plasmid DNA with insert, is dramatically lower with the unpurified PCR products as compared to those with purified PCR products, regardless of the purification method. Ligation of PCR products purified using the Wizard® SV Gel and PCR Clean-Up resulted in more colonies than phenol/chloroform extraction and ethanol precipitation but gave comparable results for the percentage of colonies that contain insert. Similar results were obtained with DNA purified using the Wizard® SV 96 PCR Clean-Up System (Cat.# A9340, data not shown).
To monitor ethidium bromide removal, purified and unpurified 1kb DNA Ladder were visualized on an agarose gel before and after ethidium bromide staining. Prior to staining, no DNA bands are visible on the gel for the purified 1kb DNA Ladder, indicating removal of the ethidium bromide (see Figure 1, Panel A). To confirm the presence of DNA, the gel was then stained with ethidium bromide and visualized again. DNA fragments are clearly visible (see Figure 1, Panel B). The staining intensities are equivalent for DNA samples before and after purification, indicating efficient recovery of DNA from solutions containing ethidium bromide.
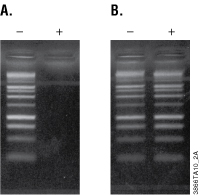
Figure 1. Gel image of purified and unpurified PCR products. Ethidium bromide was added to the 1kb DNA Ladder, which was then purified with the Wizard® SV Gel and PCR Clean-Up System. Panel A. Equivalent volumes of unpurified ladder (–) and purified ladder (+) were loaded in each lane. The gel was photographed prior to ethidium bromide staining with an exposure time of 5 seconds. Panel B. The agarose gel shown in Panel A was then stained with ethidium bromide and photographed with an exposure time of 1/8 second.
Conclusion
The Wizard® SV Gel and PCR Clean-Up System effectively remove both calf intestinal alkaline phosphatase and ethidium bromide from DNA solutions. DNA purified with this system gave comparable ligation efficiency to that of DNA purified by phenol/chloroform extraction and ethanol precipitation. The Wizard® SV Gel and PCR Clean-Up System does not involve the use of hazardous chemicals, unlike phenol/chloroform extraction and ethanol precipitation.
References
- Aktipis, S. and Panayotatos, N. (1976) Mechanism of ethidium bromide inhibition of RNA polymerase. Biochem. Biophys. Res. Comm. 68, 465–70.
Learn more about the Wizard® SV Gel and PCR Clean-Up System