Conveniently Perform In-Solution Digestions with Immobilized Trypsin
Laurie Engel, Rod Flemming, Tonny Johnson, Dan Simpson and Marjeta Urh
Promega Corporation
Publication Date: 2009
Abstract
Immobilized Trypsin provides a convenient in-solution method of digesting simple or complex protein mixtures over a wide range of concentrations. Digestion of protein solutions are performed on the bench top at room temperature within 30 minutes. Digested peptides are easily separated from the Immobilized Trypsin via the spin column, reducing potential enzyme interference during mass spectrometry analysis. The percentage of digestion and sequence coverage is comparable to overnight digestion.
Introduction
Mass spectrometry (MS) has become a powerful tool in proteomics for proteome-wide analysis and characterization of proteins from a variety of organisms and cell types. Recent advances in mass spectrometry provide tools for protein identification, protein characterization, relative and absolute quantitation, and the study of post-translational modifications and protein:protein interactions. Proteins are generally digested with proteases to generate peptides for MS analysis followed by sequencing (tandem MS).
Trypsin is the most widely used protease for MS analysis of proteins. Trypsin cleaves specifically at Lysine (Lys) and Arginine (Arg) sites in a protein. Trypsin is used for in-solution and in-gel digestion of proteins. Generally, free trypsin is used in these experiments. However, there are a few limitations in using free trypsin. These limitations are 1) the digestion requires a long time (three hours to overnight), often at high temperature and 2) high concentrations of trypsin cannot be used because of the generation of dominant trypsin autolytic fragments in the digested samples (1–5).
To overcome these limitations, we have immobilized trypsin for in-solution digestion of simple or complex proteins. Immobilized Trypsin reduces digestion time and allows easy removal of trypsin from the digestion reaction (Figure 1). Immobilized Trypsin allows flexibility, accommodating the digestion of 20–500µl of protein simply by changing the amount of resin used in the reaction. This flexibility facilitates the digestion of the protein while decreasing potential trypsin interference in downstream sample analysis.
Immobilized Trypsin has been used for studying protein expression profiling in serum (6), microwave-assisted digestion of proteins (7), phosphopeptide analysis (8,9), analysis of membrane proteins (10) and for 18O/16O labeling of peptides (11).
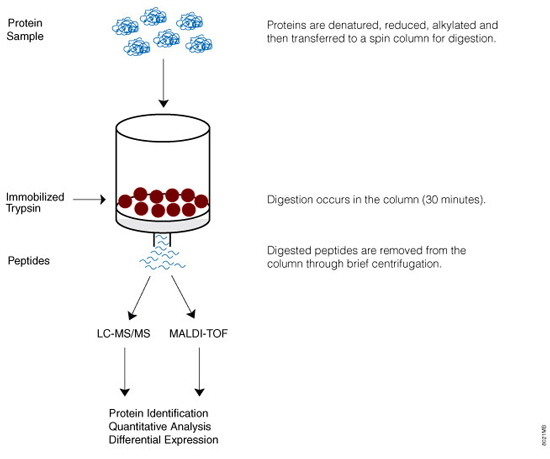
Figure 1. Overview of the Immobilized Trypsin digestion protocol. Proteins are denatured, reduced and alkylated as appropriate before digestion. The protein digestion solution, containing the protein to be digested, is prepared and added to the Immobilized Trypsin resin. The protein digestion solution is incubated with the resin for 30 minutes. Digested peptides are removed by centrifugation and then prepared for downstream analysis.
In this report, we describe the unique features and advantages of our Immobilized Trypsin for in-solution digestion of proteins. Immobilized Trypsin can be used for the digestion of purified as well as complex protein samples.
Immobilized Trypsin Decreases Digestion Time
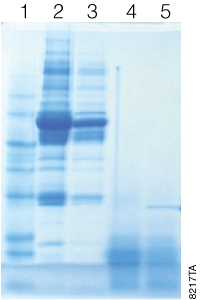
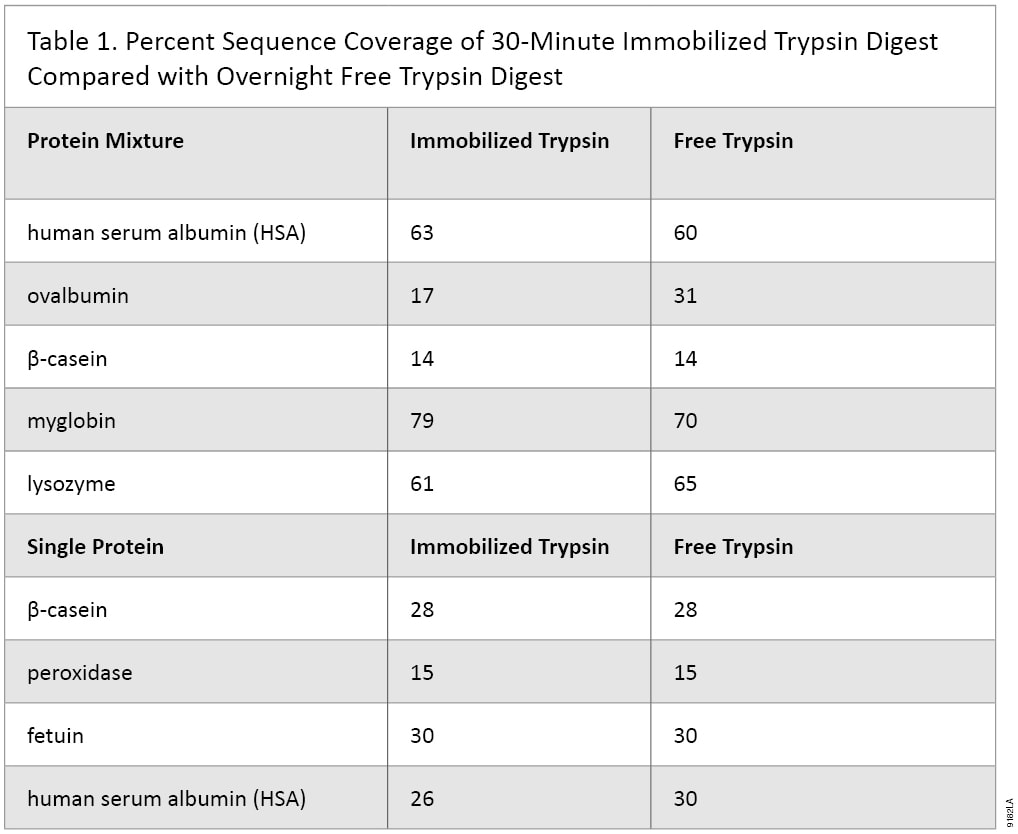
Immobilized trypsin particles contain a high concentration trypsin, which results in faster digestion time. Protein digestion can be achieved within 30 minutes at room temperature. To demonstrate this feature a depleted bovine serum sample was digested using the in-solution overnight method, and we compared it to a 30-minute Immobilized Trypsin digestion (Figure 2). The results show that digestion of complex protein samples such as serum can be digested within 30 minutes using Immobilized Trypsin. In addition MS/MS analyses of simple and complex protein samples have shown that the sequence coverage of protein digested with immobilized trypsin (30 minutes digestion) is comparable to overnight digestion using free trypsin (Table 1).
Immobilized Trypsin Reduces the Amount of Trypsin in the Final Peptide Solution
Regardless of the ratio of enzyme to protein that is used in the digestion reaction, the presence of trypsin or trypsin autolytic fragments in the final peptide solution is minimal due to the immobilization of trypsin to the resin. Trypsin is irreversibly attached to the resin, decreasing the potential of incidental removal. This reduces the potential interference of excess enzyme during MS analysis, and has enabled the development of quantitative proteomics approaches (12).
Immobilized Trypsin Reduces Tryptic Autolysis Peaks
Like other proteases, trypsin undergoes autolysis, which may reduce its protein digestion efficiency. Immobilized Trypsin has been treated with: 1) benzamidine to block the active site during storage; 2) N-p-tosyl-l-phenylalanine chloromethyl ketone (TPCK) to block the chymotrypsin activity; and 3) formaldehyde to block amine groups on the trypsin and decrease autolysis. These treatments reduce the presence of trypsin autolytic peptides that may interfere with sample analysis and other downstream applications.
Digest 20 to 500 Micrograms of Protein
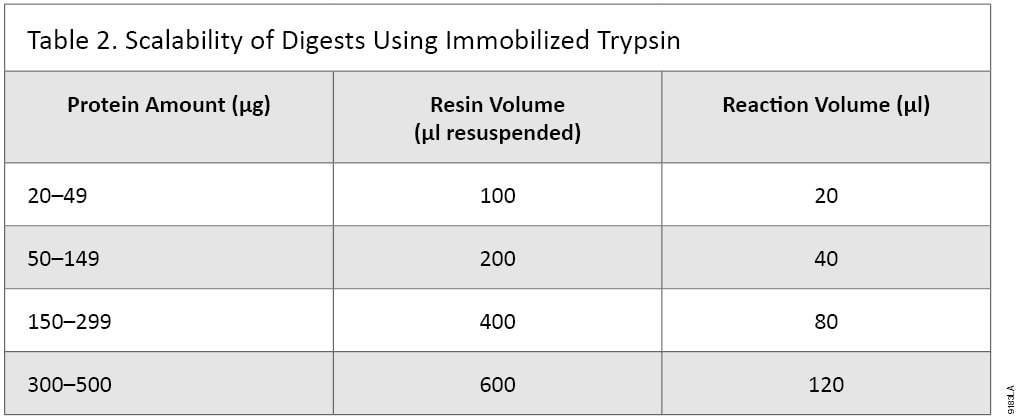
Immobilized Trypsin is designed to digest 20–500µg of protein. The amount of Immobilized Trypsin resin and digestion buffer must be determined to achieve optimal digestion (Table 2). It may be possible to digest lower or higher than the recommended concentrations of proteins, but digestion conditions will need to be adjusted.
Immobilized Trypsin is Compatible with Many Denaturants
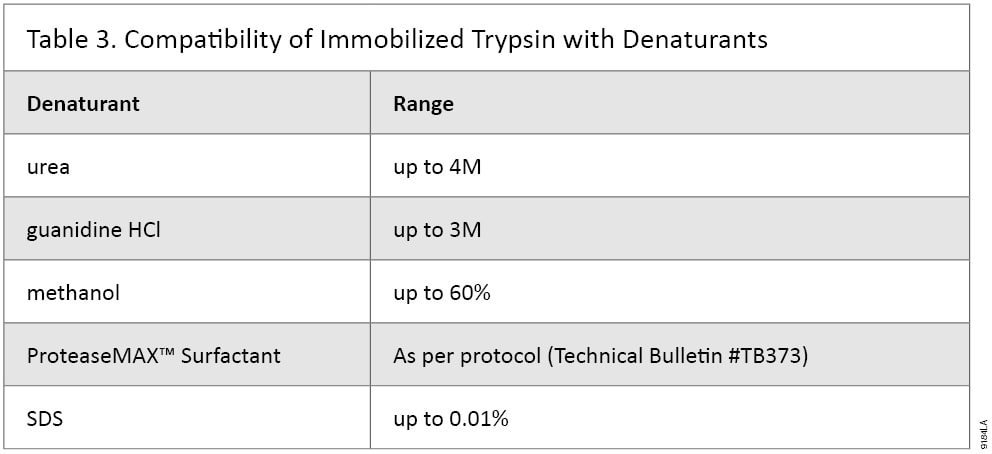
Some proteins require very stringent denaturants while others may need none. We recommend comparing multiple denaturing methods to achieve the best results. Immobilized Trypsin has been tested and shown to be compatible with urea, guanidine HCl, ProteaseMAX™ Surfactant, acetonitrile, ethanol and methanol (Table 3). Immobilized Trypsin has also been tested with various reducing and alkylation reagents to ensure compatibility. When using alternative reagents in the digestion process, be sure to remove them from the sample before MS analysis.
Buffer Formulation Reduces Nonspecific Peptide Binding
Highly hydrophobic peptides can sometimes nonspecifically bind to various resins. Our unique buffer formulation prevents nonspecific binding of digested peptides to resins. This allows better peptide recovery and peptide identification.
Summary
Immobilized Trypsin provides a format for in-solution protein digestion that is rapid and easy to use. The system is designed to reduce potential interference in sample analysis caused by autolytic fragments. When using Immobilized Trypsin, very high concentrations of trypsin can be used, aiding in the digestion of proteins. Immobilized Trypsin is removed from the final peptide solution by use of the spin column to reduce potential interference in the downstream analysis of the peptides. It also provides a scalable method of digesting proteins.
References
- López-Ferrer, D. et al. (2009) Evaluation of a high-intensity focused ultra sound-immobilized trypsin digestion and 18 O labeling method for quantitative proteomics. Anal. Chem. 81, 6272–7.
- Park, Z.-Y. and Russell, D.H. (2000) Thermal denaturation: A useful technique in peptide mass mapping. Anal. Chem. 72, 2667–70.
- Freije, J.R. et al. (2005) Chemically modified, immobilized trypsin reactor with improved digestion efficiency. J. Proteome Res. 4, 1805–13.
- Henzel, W., Watanabe, C. and Stults, J. (2003) Protein identification. The origins of peptide mass fingerprinting. J. Am. Soc. Mass Spectrom. 14, 931–42.
- Li, Y. et al. (2007) On-plate digestion of proteins using novel trypsin–immobilzed magnetic nanospheres for MALDI-TOF-MS analysis. Proteomics 7, 3661–71.
- Karbassi, I.D. et al. (2009) Proteomic expression profiling and identification of serum proteins using immobilized trypsin beads with MALDI-TOF/TOF. J. Proteome Res. 9, 4182–92.
- Hahn, H.W. et al. (2009) Ultrafast microwave-assisted in-tip digestion of proteins. J. Proteome Res. 8, 4225–30.
- Rush, J. et al. (2005) Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat. Biotechnol. 23, 94–101.
- Park, O., Swaisgood, H.E. and Allen, J.C. (1998) Calcium binding of phosphopeptides derived from hydrolysis of αs-casein or β-casein using immobilized trypsin. J. Dairy Sci. 81, 2850–7.
- Durr, E. et al. (2004) Direct proteomic mapping of the lung microvascular endothelial cell surface in vivo and in cell culture. Nat. Biotechnol. 22, 985–92.
- Cox, H.D. et al. (2008) Efficient digestion and mass spectral analysis of vesicular glutamate transporter 1: A recombinant membrane protein expressed in yeast. J. Proteome Res. 7, 570–8.
- Mirza, S.P., Greene, A.S. and Olivier, M. (2008) 18 O labeling over a coffee break: A rapid strategy for quantitative proteomics. J. Proteome Res. 7, 3042–8.
Learn more about the ProteaseMAX™ Surfactant