A TEV Protease Compatible with Inhibitory Compounds from Protein Purification
Alisha Truman and Brad Hook
Promega Corporation
Publication Date: November 2013; tpub_136
Abstract
ProTEV Plus, an engineered form of TEV protease, is a highly specific proteolytic enzyme that cleaves fusion proteins at a specific TEV linkage site. ProTEV Plus functions over a broad pH and temperature range. In this study we test the enzymatic activity of ProTEV Plus in the presence of various compounds commonly found in protein purification protocols.
Introduction
ProTEV Plus (Cat.# V6101) is an engineered form of Tobacco Etch Virus (TEV) protease, a highly specific proteolytic enzyme that cleaves within a seven-amino-acid sequence. It can be used to cleave protein fusions that have been engineered to contain this seven-amino-acid sequence at the desired cleavage site. ProTEV Plus also contains a metal affinity tag, HQ tag, located at the N-terminus of the protein. The HQ tag consists of 6 amino acids (HQHQHQ), and allows the protease to be immobilized on nickle affinity resins and removed from the cleavage reaction. ProTEV Plus protease performs over a broad pH and temperature range, and has been shown to be highly specific in cleaving fusion proteins with a TEV recognition site both in solution as well as when the fusion protein is bound to affinity resins (1) . Here we show that ProTEV Plus can be used to cleave fusion proteins in the presence of various compounds common to protein purification protocols.
Methods
ProTEV Plus Digestions of HaloTag® Standard Protein
HaloTag® Standard Protein (Cat.# G4491), a ~61kDa fusion protein containing HaloTag® (34kDa) and glutathione-S-transferase (GST; 26kDa) separated by an optimized TEV cleavage site, was cleaved with ProTEV Plus protease. Protease reactions were assembled as shown in Table 1. A master mix was prepared with all components except the ProTEV Plus and test compound. The master mix was dispensed to reaction tubes, and the appropriate compounds (Table 2) were added before finally dispensing ProTEV Plus. Two control reactions were used: 5% Sodium Dodecyl Sulfate (SDS), a known inhibitor of protease, was used to block cleavage, and water was used as a no inhibitor control.
Table 1. Protease Reaction Components
|
Reagent
|
Volume per Reaction
|
|---|---|
|
ProTEV Buffer, 20X
|
2.5µl
|
|
DDT, 100mM
|
0.5µl
|
|
HaloTag® Standard Protein (10µg)
|
2.5µl
|
|
ProTEV Plus (5 units)
|
1µl
|
|
Water
|
19.5µl
|
|
2X Compound
|
25µl
|
|
Final Volume
|
50µl
|
|
Final Reaction Concentration
|
Test Compound
|
|---|---|
|
Imidazole
|
500mM, 50mM, 5mM, 0.5mM
|
|
Sodium Chloride (NaCl)
|
1M, 125mM, 50mM, 10mM
|
|
Protease Inhibitor Cocktail
|
1X, 0.05X
|
|
Urea
|
500mM, 125mM
|
|
Guanidine-HCl
|
500mM, 1mM
|
|
Glutathione
|
2.5mM, 0.01mM
|
|
Magnesium Chloride (MgCl2)
|
100mM, 0.1mM
|
|
Glycerol
|
50%, 1%
|
|
IGEPAL® CA 630
|
1%, 0.05%
|
|
Tween™ 20
|
1%, 0.05%
|
|
Triton™ X-100
|
1%, 0.05%
|
|
SDS
|
5%, 1%
|
Protease reactions were incubated at 30°C for 30 minutes. Following digestion, 30µl of each reaction was added to 10µl of 4X SDS gel-loading buffer, heated to 95°C for 5 minutes, and 20µl was loaded onto a 4–20% precast SDS gel (Bio-Rad, Cat.# 345-0033). The gels were run for 1 hour at 200V and stained with SimplyBlue™ SafeStain (LifeTech, Cat.# LC6060). The SDS gels were visualized using a Molecular Imager® Gel Doc™ XR+ System and quantitated with Image Lab™ Software.
Results
We determined the percentage of HaloTag® Standard Protein, which was comprised of the 34kDa HaloTag® protein and 26kDa GST protein, cleaved by quantitating and comparing the bands of uncleaved HaloTag® Standard Protein in each experimental reaction to those present in the water control reactions (Figure 1). Bands were quantitated using Image Lab™ Software. The presence of NaCl (>10mM), imidazole (>5mM), glutathione (>1mM) and MgCl2 (>1mM) decreased ProTEV Plus cleavage efficiency to less than 50%. ProTEV Plus cleavage efficiency remained at ≥94% in the presence of Tween™ 20, Triton™ X-100, IGEPAL®, glycerol, urea, Protease Inhibitor Cocktail and guanidine for all concentrations tested (Figure 2).
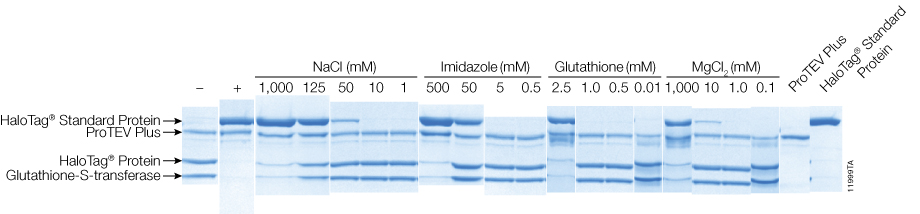
Figure 1. ProTEV Plus digestions containing NaCl, imidazole, glutathione or MgCl2 resulted in a variable amount of cleavage inhibition. HT-GST protein (10µg) was digested with 5 units of ProTEV Plus in 50µl reactions for 30 minutes at 30°C. Reactions were separated using 4–20% SDS-PAGE at 200V for 1 hour. Complete inhibition controls (+) contained 5% SDS, and no-inhibition controls (–) contained water.
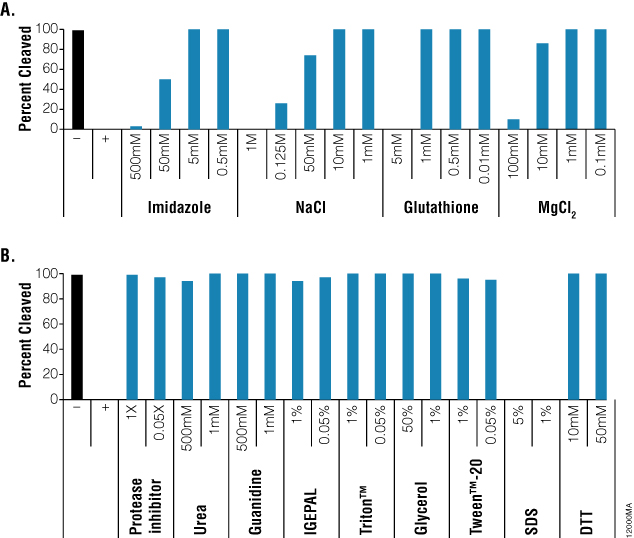
Figure 2. Percent cleavage of HaloTag® Standard Protein using ProTEV Plus in the presence of various compounds. Percent cleavage was calculated by Image Lab™ Software by comparing the bands of the partially digested HaloTag® Standard Protein to the band of undigested HaloTag® Standard Protein. Panel A. Imidazole, NaCl, Glutathione and MgCl2. Panel B. Protease Inhibitors, Urea, Guanidine, IGEPAL®, Triton™ X, Glycerol, Tween™ 20, SDS and DTT.
Summary
We have determined that a variety of compounds commonly used with protein purification protocols do not inhibit ProTEV Plus cleavage reactions. Table 3 lists the compounds tested and the concentrations at which ProTEV Plus shows ≥94% cleavage. The concentration limits provided are general guidelines because we did not test the compound concentrations in small increments. For some of the compounds analyzed, the highest acceptable concentration may be higher than indicated.
|
ProTEV Plus-Compatible Compounds
|
Acceptable Concentrations
|
|---|---|
|
Protease Inhibitor Cocktail
|
≤1X*
|
|
Urea
|
≤500mM*
|
|
Guanidine-HCl
|
≤500mM*
|
|
IGEPAL® CA-630, Tween™ 20, Triton™ X-100
|
≤1%*
|
|
Glycerol
|
≤50%*
|
|
DTT
|
≤10mM*
|
|
Imidazole
|
≤5mM
|
|
Sodium Chloride (NaCl)
|
≤10mM
|
|
Glutathione
|
≤1mM
|
|
Magnesium Chloride (MgCl2)
|
≤1mM
|
|
SDS
|
All tested concentrations resulted in 0% cleavage.
|
|
*Indicates the maximum concentration tested for a given compound.
|
References
Anderson, J. et al. (2008) ProTEV Protease for Fusion Protein Processing. Promega eNotes
Featured Product
This article highlights use of ProTEV Protease, an engineered protease with greater stability and prolonged activity over native TEV protease.