LCN DNA Analysis: Limitations Prevent 'General Acceptance'
Faculty of Health Science & Medicine, Bond University, Gold Coast, Queensland, Australia
Publication Date: 2010
Introduction
DNA analysis is considered the gold standard amongst the forensic disciplines, as highlighted by the recent report on the forensic sciences from the National Academy of Science(1). DNA typing is an extremely effective tool for determining the contributor of biological material associated with a crime. The exquisite sensitivity of the polymerase chain reaction (PCR) allows minute quantities of DNA to be analyzed. Underpinning the robust nature of forensic DNA analysis are extensive validation studies, which have established the conditions under which the assays produce reliable results(2)(3)(4).
The phenomenal success of DNA profiling has increasingly led to attempts to analyze more difficult and challenging samples such as those containing DNA from only a few cells. Attempts to increase the sensitivity of STR assays by increasing the number of PCR cycles were first described more than a decade ago(5)(6). This approach, which has become known as low copy number (LCN) DNA typing, has led to the analysis of a new category of samples called variously "touch", low-template or high-sensitivity(7). Despite the more than ten years since the Forensic Science Service (FSS) in the UK documented its analysis of extremely low levels of DNA, many forensic laboratories have not formally adopted the technique, although many are analyzing samples containing less than 100–200pg.
The limitations of LCN arise from the exaggerated stochastic effects seen with the very low levels of input template DNA. These effects are exacerbated by the fact that the majority of LCN samples are mixture samples(8)(9).
This paper will address two questions regarding LCN analysis:
1. Has LCN typing been validated and accepted in nonforensic areas of DNA testing?
2. Is it better to consume a sample with a single amplification to increase the possibility of obtaining a few more alleles and the possibility of a more complete profile vs. duplicate/triplicate amplifications with repetition of data?
"In order for the forensic community to generate reliable results from these low-level samples, significant improvements to the current LCN methods must be made."
General Acceptance
The first question addresses the legal standard for general acceptance under Frye and the first prong of acceptance under Daubert for admission of expert evidence. The Daubert standard, as well as requiring widespread acceptance in the relevant scientific community, necessitates assessment of additional criteria such as testability, error rate and peer review publication. It has been argued that LCN is not novel because the same process (PCR) and same kits are used (Linacre, ISFG 2009). This does not translate to reproducibility or reliability, and without these fundamental scientific standards there should not be general acceptance. That a person's life or liberty might hinge on a method that is not reproducible or reliable is anathema to the goals of the forensic science community. The lack of reproducibility of LCN analysis is well documented(5)(10)(11). The same overall result is not expected when the sample is analyzed twice. On this basis alone LCN typing cannot be considered robust or reliable, although it was so described in a recent review of low template DNA analysis, commissioned by the Forensic Regulator in the UK in response to criticisms by the judge in the Omagh bombing case(12). It is a combination of what is used and how it is used that determines the reliability of any method. It is important to recognize that a lack of novelty does not translate to general acceptance. If lack of novelty was the criterion for general acceptance, the courts would hear evidence on polygraphs, intelligent design, astrology and other areas widely recognized as not meeting the necessary scientifically accepted standards.
The use of PCR for analysis of very low levels of DNA in the field of pre-implantation genetic diagnosis (PGD) has been used to justify general acceptance of LCN analysis in the wider scientific community(13). This argument is flawed. PGD analysis is not analogous to LCN analysis for several reasons. PGD uses pristine cellular DNA from a single source, whereas forensic LCN samples are mostly mixtures and are likely in a state of apoptosis. The complexities of profile interpretation issues seen with STR analysis (e.g., stutter) are not an issue for PGD testing(14). However the most significant difference is that the samples derived from the mother, father and embryo are single-source and the parental samples are of known genotype. The PGD analysis is undertaken to determine the genotype of the embryo. In the situation where both parents are carriers for a recessive genetic disease (Dd), embryos are tested in order to select a nondisease embryo for implantation. The genotype of the embryo will be either DD, Dd or dd, and one in four embryos would be expected to result in a disease genotype (dd). If a D allele is detected the embryo can be implanted. The occurrence of allele d drop-out such that a Dd embryo was mistyped as DD would not be of concern. It is of course possible that a Dd embryo is mistyped as dd as a result of D allele drop-out. This may result in choosing to discard a healthy embryo. However, it is possible to use haplotyping of markers surrounding the parental D and d alleles to confirm or reject allele drop-out and therefore substantially reduce erroneous results (unlike in LCN typing).
It is worth noting that it is not possible for the forensic science community to properly assess the general acceptance of LCN since the primary advocates of LCN typing [FSS in the UK, Office of the Chief Medical Examiner (OCME) in New York and Environmental Science & Research (ESR) in New Zealand] have declared that their protocols are proprietary (personal communication). Such nondisclosure means it is not possible for the scientific community to assess the validity of practices when these protocols are concealed. Thus, there can be no peer review of the methods and how they are applied.
Sample "Replicates"
The nonreproducible nature of the LCN process is well established(5)(10)(11). In an attempt to address this inherent lack of reproducibility laboratories split the sample into two or three aliquots and report only the alleles that are common in at least two of the replicates(5)(10)(15). This would result in, for example, a 60pg sample (~10 cells) being analyzed in two or three separate reactions, each with five or three cells, respectively. While there is some appeal to this redundancy approach, common sense would intimate that this further reduction in template quantity will exacerbate the vagaries of LCN typing. Table 1 provides examples of cell numbers and corresponding DNA amounts where the sample is or is not split into three aliquots. It is apparent from this table that many LCN samples would have contributor DNA present at the equivalent of less than one cell.
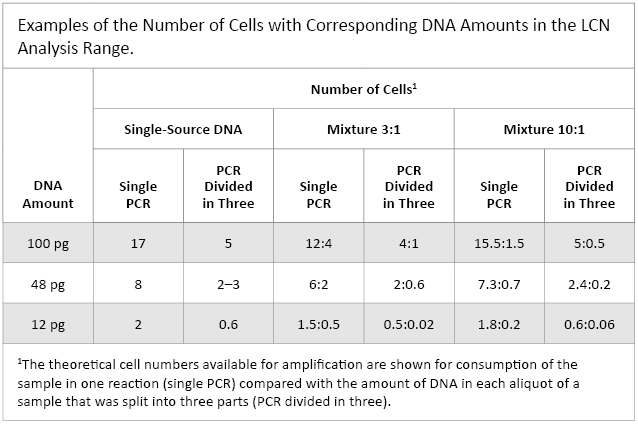 Table 1. Examples of the Number of Cells with Corresponding DNA Amounts in the LCN Analysis Range.
Table 1. Examples of the Number of Cells with Corresponding DNA Amounts in the LCN Analysis Range. Since there are inherent reproducibility problems with LCN typing and replicate analyses are invoked, the interpretation guidelines and statistical assessment of the weight of the evidence must be robust and reliable. However a decade later, the recommended and often cited statistical methods of Gill et al. have not been implemented, not even by the laboratory that developed the approach(5). In fact Gill has stated that the consensus method was never intended to be a permanent solution (ISFG 2009).
Conclusion
In summary, the lack of reproducibility and other issues surrounding current LCN typing demonstrate a lack of scientific reliability and thus acceptability. The practice of further splitting already low amounts of DNA template is questionable. In order for the forensic community to generate reliable results from these low-level samples, significant improvements to the current LCN methods must be made.
Article References
- Mulero, J.J. et al. (2008) Development and validation of the AmpFlSTR® MiniFiler™ PCR Amplification Kit: A miniSTR multiplex for the analysis of degraded and/or PCR inhibited DNA. J. Forensic Sci. 53, 838–52.
- Cotton, E.A. et al. (2000) Validation of the AMPFlSTR® SGM Plus™ system for use in forensic casework. Forensic Sci. Int. 112, 151–61.
- Krenke, B.E. et al. (2002) Validation of a 16-locus fluorescent multiplex system. J. Forensic Sci. 47, 773–85.
- Gill, P. et al. (2000) An investigation of the rigor of interpretation rules for STRs derived from less than 100 pg of DNA. Forensic Sci. Int. 112, 17–40.
- Findlay, I. et al. (1997) DNA fingerprinting from single cells. Nature 389, 555–6.
- van Oorschot, R.A. and Jones, M.K. (1997) DNA fingerprints from fingerprints. Nature 387, 767.
- Budowle, B. et al. (2001) Low copy number — consideration and caution. Proceedings From the Twelfth International Symposium on Human Identification 2001
- Budowle, B., Eisenberg, A.J. and van Daal, A. (2009) Validity of low copy number typing and applications to forensic science. Croat. Med. J. 50, 207–17.
- Caragine, T. et al. (2009) Validation of testing and interpretation protocols for low template DNA samples using AmpFlSTR® Identifiler®. Croat. Med. J. 50, 250–67.
- Gill, P. (2001) Application of low copy number DNA profiling. Croat. Med. J. 42, 229–32.
- Caddy, B., Taylor, G.R. and Linacre, A. (2008) A review of the science of low template DNA analysis.
- People v. Hemant Megnath, 2010 N.Y. Slip Op 20037; 2010 N.Y. Misc. Lexis 223.
- Moretti, T.R. et al. (2001) Validation of short tandem repeats (STRs) for forensic usage: Performance testing of fluorescent multiplex STR systems and analysis of authentic and simulated forensic samples. J. Forensic Sci. 46, 647–60.
- Petricevic, S. et al. (2010) Validation and development of interpretation guidelines for low copy number (LCN) DNA profiling in New Zealand using the AmpFlSTR® SGM Plus™ multiplex. Forensic Sci. Int. Genet. 4, 305–10.
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
van Daal, A. LCN DNA Analysis: Limitations Prevent 'General Acceptance'. [Internet] 2010. [cited: year, month, date]. Available from: https://www.promega.com/resources/profiles-in-dna/lcn-dna-analysis-limitations-prevent-general-acceptance/
American Medical Association, Manual of Style, 10th edition, 2007
van Daal, A. LCN DNA Analysis: Limitations Prevent 'General Acceptance'. Promega Corporation Web site. https://www.promega.com/resources/profiles-in-dna/lcn-dna-analysis-limitations-prevent-general-acceptance/ Updated 2010. Accessed Month Day, Year.
Contribution of an article to Profiles in DNA does not constitute an endorsement of Promega products.