Measuring Intracellular Protein Lifetime Dynamics Using NanoLuc® Luciferase
Matthew Robers, Brock Binkowski, Jim Hartnett, Jennifer Wilkinson, Chad Zimprich, Pete Stecha and Mei Cong
Promega Corporation, 2800 Woods Hollow Road, Madison WI 53711
Publication Date: 2/2014; tpub 139
Abstract
Adaptive stress response pathways are tightly regulated by the localization and concentration of key intracellular transcription factors. The rapid changes in protein lifetime that occur following environmental or chemical stress can be exploited using a new genetic reporter strategy that includes NanoLuc® luciferase. NanoLuc® luciferase is ideally suited as a protein function reporter for measuring changes in transcription factor levels following induction of stress response signaling. Via genetic attachment of NanoLuc® luciferase to various target proteins, induction of these pathways can be measured using a simple, luminescent assay readout. The physical properties of NanoLuc® luciferase (e.g., small size, ATP-independence, extreme brightness, etc.) make an excellent reporter for quantifying endogenous levels of these naturally low-abundance proteins. NanoLuc® luciferase-enabled assays for monitoring the intracellular levels of these highly dynamic proteins provides an excellent complement to existing reporter gene assays for toxicological screening and general pathway analysis.
Introduction
Traditional reporter gene assays have proven to be powerful tools to interrogate adaptive stress-response signaling mechanisms. Transcriptional responses occur at distal signaling endpoints, and reporter gene assays successfully integrate numerous signaling mechanisms into a single cellular output. This provides a convenient means for measuring global cellular responses to small-molecule modulators. However, drug actions typically occur at signaling events far upstream of gene activation. For this reason, the use of reporter genes assays to elucidate molecular targets of inhibitors during drug discovery can be challenging, especially within highly complex pathways. Furthermore, transcriptional readouts have limited utility for real-time pathway analysis and often require extended drug treatment times for adequate performance. Consequently, alternate methods that measure signaling responses at upstream pathway nodes are desirable as complements to transcriptional reporters.
Stress response pathways use a common architecture involving dynamic regulation of protein turnover that allows for such alternate methods to be considered. For example, hypoxia-inducible factor-1A (HIF1A) pathway is acutely regulated by the ubiquitin-proteosome system (Figure 1). Under normoxic conditions, the transcription factor HIF1A is repressed by prolyl hydroxylation and VHL-directed ubiquitination. Ubiquitinated HIF1A becomes destined for proteosomal degradation, resulting in low levels of HIF1A protein and low basal levels of transcription of HIF1A-responsive genes. Upon transition to hypoxic conditions or chemical inhibition of HIF1A prolyl hydroxylation, HIF1A proteins decouple from the ubiquitin/proteasome pathway and accumulate in the nucleus. In turn, transactivation of various HIF1A-responsive genes occurs, resulting in a cellular response to hypoxic stress. Small molecule modulation of HIF1A signaling may prove promising in treating cancer or ischemic diseases, underscoring the value of quantitative methods for measuring HIF1A stabilization during drug discovery.
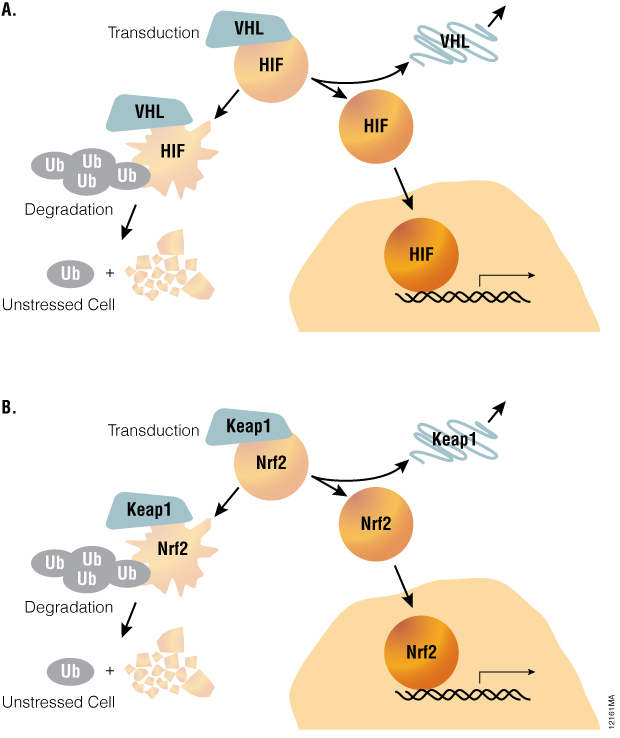
Figure 1. A common pathway architecture governs adaptive stress response signaling. The stability of key transcription factors changes in response to hypoxia or oxidative stress. When induced by hypoxia or chemical hypoxia mimetics, the HIF1A transcription factor decouples from the ubiquitin/proteosome pathway and accumulates in the nucleus to activate target genes (Panel A). Similarly, the NRF2 transcription factor accumulates in response to inducers of oxidative stress (Panel B).
The oxidative stress response pathway is similarly regulated by protein lifetime (Table 1). In unperturbed cells, nuclear factor (erythroid-derived 2)-like 2 (NRF2) is subject to Keap1-directed ubiquitination and proteosomal degradation. Reactive oxygen species disrupt the interaction between Keap1 and NRF2, resulting in NRF2 accumulation and transactivation. NRF2 target genes are then responsible for scavenging reactive oxygen species and a return to cellular homeostasis. While HIF1A and NRF2 mechanisms are key examples of the relevance of protein lifetime in regulation of gene activation, this pathway architecture is conserved within many signaling pathways, as described in Table 1( (1) ). Thus, protein lifetime has the potential to be broadly exploited as a readout for critical upstream signaling nodes in adaptive stress response pathways.
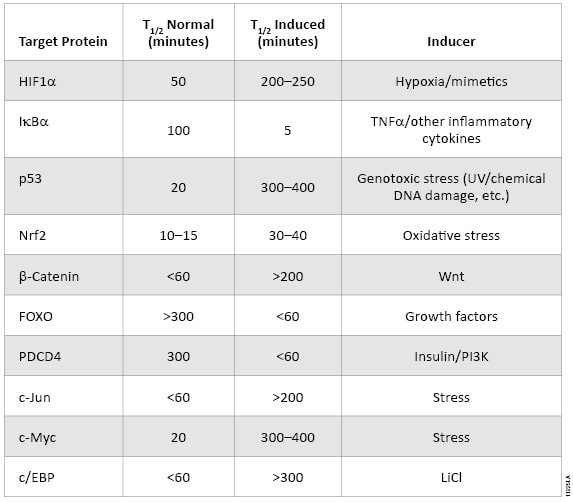
Table 1. The intracellular lifetime of regulatory proteins from key signaling pathways changes in response to pathway activation.
Methods
Due to its small size (19kDa) and intense bioluminescence output (Figure 2), NanoLuc® luciferase is ideally suited as a protein function reporter for quantifying protein lifetime dynamics. Changes in intracellular protein levels can be quantified using Nano-Glo® reagent in a simplified assay format, wherein the luminescence output of NanoLuc® luciferase is used as a surrogate for the intracellular levels of the fusion protein. Via direct genetic fusions of NanoLuc® luciferase with HIF1A or NRF2 proteins, it is possible to measure hypoxia or oxidative stress-response signaling (respectively) in HCT116 colorectal cancer cells. Using a constitutive (e.g., CMV) promoter driving HIF1A-NanoLuc® or NRF2-NanoLuc®, changes in light output correlate to dynamic changes in protein levels. However, use of robust promoters such as CMV may result in unnaturally high intracellular levels of the protein of interest, creating potential assay artifacts.
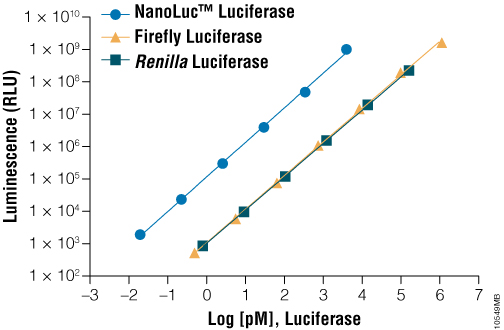
Figure 2. A comparison of luminescence of various purified luciferases in the presence of their optimal substrates. On a molar basis, NanoLuc® luciferase is much brighter than commonly used luciferases such as firefly luciferase and Renilla luciferase.
To address this potential liability, plasmid DNA encoding genetic fusions of NanoLuc® luciferase are serially diluted into promoterless Transfection Carrier DNA to better approximate physiologically relevant expression levels of the protein of interest. To address this need, Promega offers Transfection Carrier DNA (Cat.# E4881), qualified for use in mammalian transfection experiments. In addition, a control vector, pNL3.2 [CMV], encoding NanoLuc® luciferase fused to a PEST destabilization domain (NlucP) is available as a negative control for nonspecific impacts on NanoLuc® signals. These nonspecific impacts may result from toxic chemical insults or nonspecific proteasome inhibitors introduced to cells.
As shown in Figure 3, when HCT116 cells are transiently transfected with a 1:1,000 dilution of HIF1A-NanoLuc® plasmid DNA, 1, 10-phenanthroline (a known hypoxia mimetic) induces a dose- and time-dependent accumulation of the reporter fusion. Similarly, an NRF2-NanoLuc® fusion protein accumulates in HCT116 cells upon treatment with D,L-sulforaphane (a known inducer of reactive oxygen species). To ensure proper regulation of NRF2-NanoLuc® levels in unperturbed conditions, cells are cotransfected with KEAP1 expression DNA. These results serve to demonstrate the use of NanoLuc® luciferase as a protein function reporter for monitoring regulated changes in protein stability within stress response pathways. Furthermore, these responses occurred within 2–3 hours, compared to traditional reporter gene assays that require up to 16 hours to achieve optimal responses. Protein stability reporters of HIF1A and NRF2 provide a proximal readout of the targets of these chemical stressors, enabling a highly precise mode of compound action for toxicological screens during drug discovery.
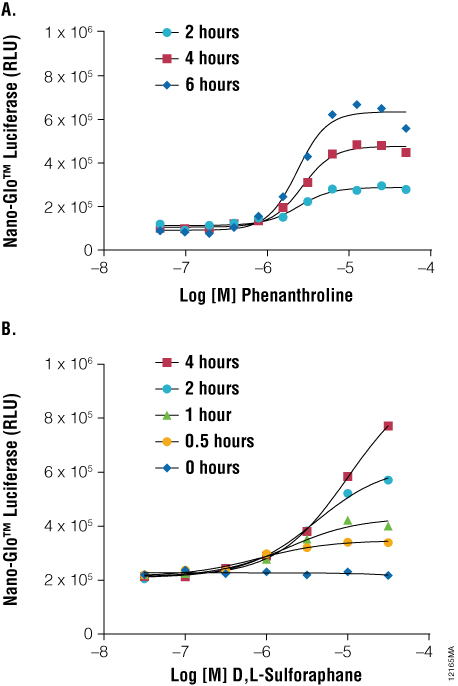
Figure 3. As a protein function reporter, NanoLuc® Luciferase enables quantitation of changes in protein stability following induction of stress response signaling. Panel A. HCT-116 cells were transiently transfected with the pNLF1-HIF1A[CMV/neo] Vector and Transfection Carrier DNA in hypoxia mimetic. Panel B. HCT-116 cells were transiently transfected with the pNLF1-NRF2[CMV/neo] Vector and pKEAP1 DNA and exposed to a stimulant. Treatments occurred as indicated and NanoLuc® Luciferase detected using Nano-Glo™ Luciferase Assay at the indicated time points using the GloMax®-Multi+ Plate Reader.
NanoLuc® luciferase also enables applications of protein stability sensors in advanced cell models. Induced pluripotent stem cell (iPSC)-derived cardiomyocytes are an attractive cell model for studying HIF1A signaling in ischemic disease ( (2) ). As shown in Figure 4, Panel A, induction of HIF1A levels can be monitored in iCell® Cardiomyocytes transiently transfected with HIF1A-NanoLuc® fusion proteins. Prolyl hydroxylase inhibitors such as IOX2, CoCl2, ML-228, and 1, 10-phenanthroline each induced dose-dependent increases in HIF1A-NanoLuc® proteins after only 3 hours of stimulation. These results are consistent with results generated using a hypoxia response element (HRE-luc2P) firefly reporter gene assay, following 6-hour stimulation (Figure 4, Panel B). The application of the HIF1A-NanoLuc® stability sensor within iCell® Cardiomyocytes provides an elegant assay readout in a physiologically-relevant cellular context for studying HIF1A signaling.
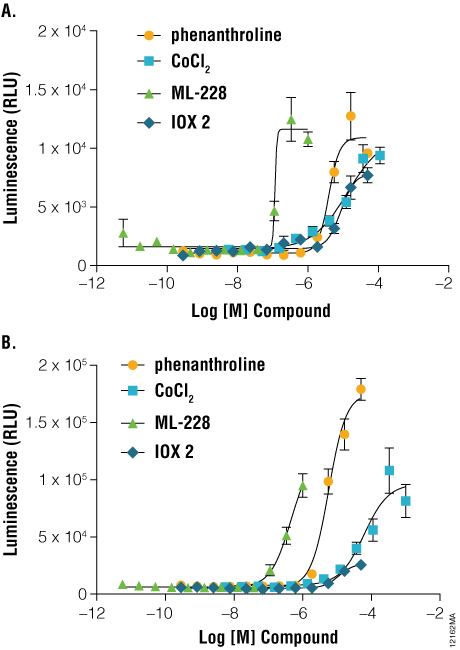
Figure 4. A comparison of hypoxia response reporters in iPS-derived cardiomyocytes. Three to five days post seeding, iCell® Cardiomyocytes were transiently transfected using either the pNLF1-HIF1A[CMV/neo] fusion construct, diluted 1:100 into Transfection Carrier DNA (Panel A) or pGL4.42[luc2P/HRE/Hygro] HRE response element Vector (Panel B). Twenty-four hours post-transfection, cells were stimulated for either 3 hours (HIF1A-NanoLuc® fusion) or 6 hours (HRE reporter) with the indicated compounds and assayed using either ONE-Glo™ Luciferase Assay System or Nano-Glo™ Luciferase Assay Reagents, respectively. Luminescence was quantified on a GloMax®-Multi+ Plate Reader.
As a protein function reporter, NanoLuc® luciferase also provides a quantitative measure of protein levels from endogenously expressed genetic loci. As overexpression of the protein of interest may lead to assay artifacts, the ability to express the target protein at endogenous levels may be desirable. However, use of reporters with limited sensitivity provides a major challenge when the target protein is produced at endogenous levels, especially when the target protein is naturally unstable. To explore the utility of NanoLuc® luciferase as a reporter of endogenous protein stability, Horizon’s proprietary GENESIS™ platform was used to introduce the NanoLuc® gene into specific chromosomal loci as in‐frame protein fusions to HIF1A or NRF2.
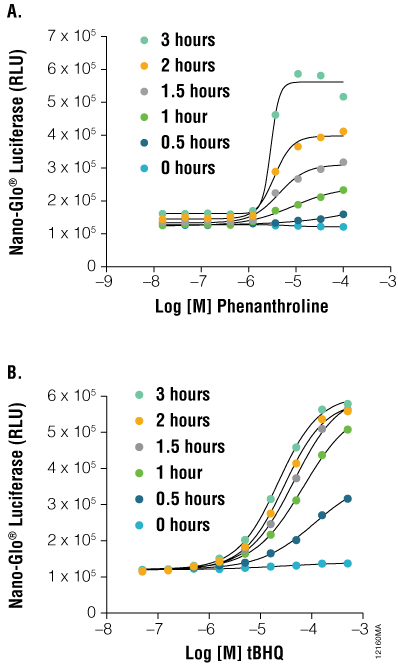
Figure 5. As a protein function reporter, NanoLuc® Luciferase enables detection of proteins expressed from endogenous gene loci. Engineered HCT116 cells expressing NanoLuc® Luciferase fused to endogenous HIF1A (Panel A) or NRF2 (Panel B) were stimulated with either tBHQ or 1,10-phenanthroline, respectively, with detection performed using the Nano-Glo™ Luciferase Assay, at the indicated time points. Both fusion constructs showed a dose- and time-dependent response.
As shown in Figure 5, HCT116 cells engineered with NanoLuc® as an in-frame fusion to a single allele of HIF1A can be used to monitor inducible changes in HIF1A levels upon treatment with 1, 10 phenanthroline. HCT116 cells similarly engineered with a NRF2-NanoLuc® fusion responded in a dose- and time-dependent manner to tertbutylhydroquinone (tBHQ). These results validate the use of NanoLuc® luciferase as an ultrasensitive reporter of protein stability dynamics without the need for overexpression.
Promega has provided a suite of NanoLuc® fusion vectors to enable intracellular protein stability assays. These vectors facilitate subcloning of N-terminal and C-terminal fusions with NanoLuc® luciferase using standard restriction enzyme cloning or Promega’s convenient Flexi® cloning system (Table 2). Flexi® System-compatible vectors facilitate transfer of the gene of interest from ORF clone gene panels, as provided within the Promega “Find My Gene” offering.
For more information about NanoLuc® fusion vectors, including HIF1A-NanoLuc® and NRF2-NanoLuc® expression vectors, visit: www.promega.com/products/pm/nanoluc-redefining-reporter-assays/
References
- Simmons, S.O., Fan, C.Y. and Ramabhadran, R. (2009) Cellular stress response pathway system as a sentinel ensemble in toxicological screening. Toxicol. Sci. 111, 202–25.
- Zhi D, Irvin MR, Gu CC, Stoddard AJ, Lorier R, Matter A, Rao DC, Srinivasasainagendra V, Tiwari HK, Turner A, Broeckel U, Arnett DK. (May 2012) Whole-exome sequencing and an iPSC-derived cardiomyocyte model provides a powerful platform for gene discovery in left ventricular hypertrophy. Front. Genet. 3:92.