Measuring cAMP Levels and Cytotoxicity in a Single Plate Well
Mark Bratz, Brad Hook and Trista Schagat
Promega Corporation
Publication Date: January 2013
Abstract
Multiplexing is a technique that allows the researcher to gain more information from a single sample well. Here we describe the multiplexing of a fluorescent real-time cytotoxicity assay with a bioluminescent cAMP assay to study changes in cAMP levels in the context of cytotoxic effects.
Introduction
cAMP is important as a second messenger in a variety of signaling pathways. The cAMP-Glo™ Max Assay was developed to test compounds for their ability to modulate cAMP levels in cells. This assay allows researchers to examine G-protein-coupled receptors (GPCRs) coupled with adenylate cyclase. A compound that activates or blocks GPCRs will alter intracellular cAMP levels, thus resulting in a signal change in the assay.
The cAMP-Glo™ Max Assay measures cAMP levels through protein kinase A (PKA), which is activated upon release of its regulatory subunits after binding to cAMP (Figure 1). PKA is a component of the cAMP Detection Solution, and once activated, PKA will use any ATP present to phosphorylate its substrate. Thus, the cAMP-Glo™ Max Assay produces a luminescent signal that is inversely proportional to the amount of cAMP present in the reaction. Increasing concentrations of cAMP leads to higher ATP turnover by PKA, resulting in less ATP to drive the luminescence reaction (using Ultra-Glo™ Recombinant Luciferase). Conversely, a sample with low amounts of cAMP will have high amounts of ATP and thus a high luminescent signal. In cell-based experiments where treatment results in changes to cAMP, it is important to understand that change in relation to potential cytotoxic effects or changes in cell number due to the treatment.
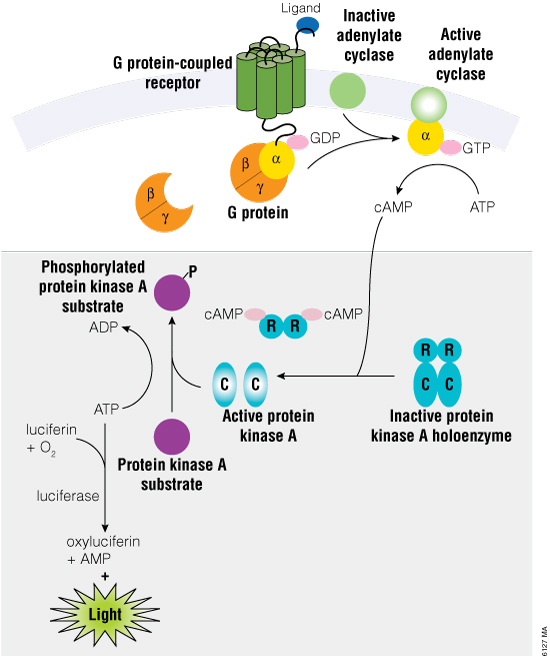
Figure 1. Schematic diagram of cAMP activation during the induction phase of the cAMP-Glo™ Max Assay (upper portion of the diagram). The shaded lower portion of the diagram depicts the assay after lysis with the cAMP Detection Solution plus the addition of the Kinase-Glo® Reagent (as described in Technical Manual #TM347).
The CellTox™ Green Cytotoxicity Assay detects the cytotoxic effects of compounds on cells. The CellTox™ Green Dye is a membrane impermeable, fluorescent, DNA-binding dye. It cannot penetrate viable cell membranes, but it can pass through the compromised membrane of dead cells to stain DNA (Figure 2). An increase in fluorescence occurs when the dye binds to DNA. The increased fluorescence correlates to increased cytotoxicity. Unlike other markers of necrotic cell death that require maintenance of enzymatic activity after release from the cytoplasm, staining of DNA provides a long lasting fluorescent signal.
The CellTox™ Green Dye is also stable to multiple fluorescence readings. This stable signal makes it ideal for extended treatment exposure times, for as long as 72 hours, allowing measurement of cytotoxicity either in real-time or as an endpoint assay. The reagent is nonlytic, allowing downstream assays to be multiplexed to gain more detailed data from each well.
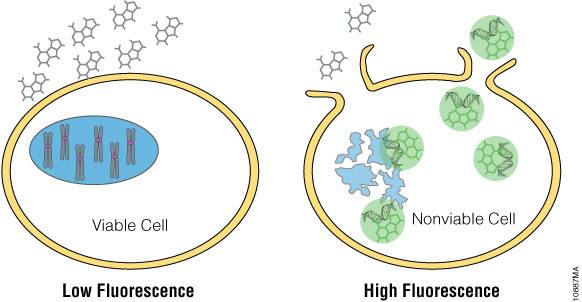
Figure 2. CellTox™ Green Cytotoxicity Assay. CellTox™ Green Dye is excluded from viable cells, but it binds to DNA from dead cells with compromised membrane integrity.
If a decrease in cAMP levels is observed, it could be through inhibition of the adenylate cyclase pathway, an increase in cell death or an error in plating, with uneven cell distribution. Addition of CellTox™ Green Dye to the assay will help to differentiate these possible scenarios.
Here we describe multiplexing the CellTox™ Green Cytotoxicity Assay with the cAMP-Glo™ Max Assay to yield data on cAMP modulation as well as the cytotoxic effects of the compound being tested and the effects of cell plating variations. Furthermore, CellTox™ Green solution can be added directly to the cAMP-Glo™ Max Assay buffer preparation, simplifying experimental setup.
In this report we tested cAMP levels in cells in the presence of two compounds, forskolin and cholera toxin. We added CellTox™ Green Dye to these assays and determined if there were any effects on the luminescent signal produced by the cAMP-Glo™ Max Assay.
Materials to Be Supplied by the User
- IBMX (Acros Organics, Cat.# 228420010)
- Forskolin (Acros Organics, Cat.# 328240250)
- cAMP-Glo™ Max Assay (Promega, Cat.# V1681)
- CellTox™ Green Cytotoxicity Assay (Promega, Cat.# G8741)
- Cholera Toxin (from Vibrio cholera; Sigma-Aldrich, Cat.# C8052-1MG)
- Ro 20-1724 (Tocris Bioscience, Cat.# 0415)
Procedure
The cAMP-Glo™ Max Assay was performed as described in the cAMP-Glo™ Max Assay Technical Manual #TM347, with a few exceptions. Here we provide a detailed protocol:
- HEK293 cells were plated (50µl per well; 1 × 105 cells/ml) into two 96-well plates for each compound tested for a total of four plates. Note: Cells were plated in rows B–G and columns 1–11. DMEM (100µl) was plated in the remaining wells.
- Cells were incubated overnight at 37°C with 5% CO2.
- An Induction Buffer stock was made by diluting IBMX to 500µM, Ro 20-1724 to 100µM and MgCl2 to 25mM in 1X PBS.
- CellTox™ Green (40µl) was added to a 10ml aliquot of Induction Buffer.
- For each treatment compound, a fivefold series of dilutions was prepared in both of the Induction Buffers (with and without CellTox™ Green Dye). Forskolin dilutions were made from 100µM–51pM. Cholera toxin dilutions were made from 1µM–0.51pM.
- Medium was aspirated from all plate wells.
- Forty microliters (40µl) of compound was added to the wells in rows B–G and columns 1–11 of the appropriate plate. Column 12 was a no-cell control for all plates. For each compound one plate was treated with compound + Induction Buffer containing CellTox™ Green and the other plate was treated with the compound + Induction Buffer only (no CellTox™ Green).
- The plates were shaken for 1–2 minutes.
- The cholera toxin plates were incubated for 45 minutes at 37°C, then equilibrated to room temperature for 15 minutes. The forskolin plates were incubated for 15 minutes at room temperature.
- Fluorescence was measured (excitation 490nm, emission 510–550nm) for the plates containing Induction Buffer + CellTox™ Green.
- cAMP Detection Solution + PKA (10µl/well) was added to all wells.
- The plates were incubated for 20 minutes at room temperature.
- Kinase-Glo® Solution (50µl/well) was added to all wells.
- The plates were incubated at room temperature for 10 minutes.
- Luminescence was measured for all four plates.
- A second fluorescence measurement was made for the Induction Buffer + CellTox™ Green-containing plates only.
Results
The CellTox™ Green cytotoxicity measurement was taken before addition of lysis solution (Step 10). Any signal at this point would be due to normal background cell death and death caused by the test compound treatment. The measurements in our experiments indicate that the compounds used did not kill the cells.
The total cell measurement was taken after the addition of both the cAMP Detection Solution and Kinase-Glo® Reagent (Step 15). All cells should be lysed at this point and the DNA released. Thus, signal was proportional to the total number of cells in each well. Measuring this signal ensures that the plates were seeded correctly and that there were approximately the same number of cells in each well. The total cell measurement can be used to normalize the cAMP-Glo™ Max data relative to number of cells in the well if large discrepancies are found.
Forskolin was the first compound tested. Forskolin binds directly to adenylate cyclase and activates cAMP production. The cells were then incubated for 15 minutes at room temperature before the cytotoxicity fluorescent read. The EC50 values determined using the cAMP-Glo™ Max Assay showed no significant difference between cells with cAMP-Glo™ Max and cells with cAMP-Glo™ Max + CellTox™ Green Assay. The overall luminescence with CellTox™ Green present was slightly reduced, with an average loss of 16%, but the large dynamic range of the luminescence assay prevents any change to the data interpretation, as seen with the EC50 values (Figure 3, Panel A). The CellTox™ Green cytotoxicity and total cell reads (Figure 3, Panel B) show that forskolin did not cause cell death within the concentrations and time assayed and that there was minimal cell number variation between wells, indicating consistent cell number in all wells.
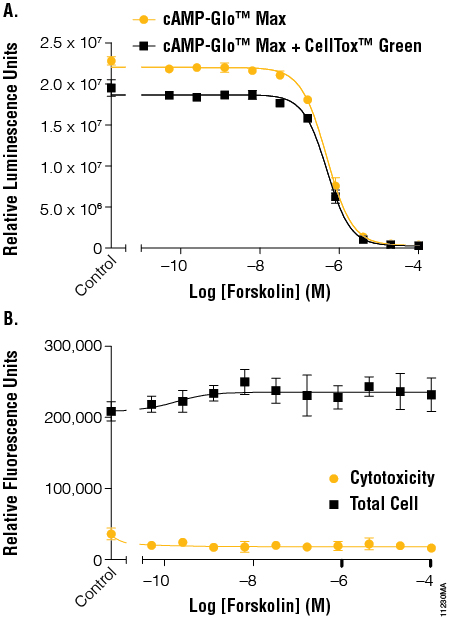
Figure 3. The effects of forskolin on cAMP levels in the context of cytotoxicity and cell number normalization. Panel A. A series of forskolin dilutions was added to cells and tested for cAMP activation using the cAMP-Glo™ Max Assay in the absence and presence of CellTox™ Green Dye. EC50 values were calculated using the average of 6 replicates using GraphPad Prism® software. The resulting EC50 values were 4.20–5.45 × 107. Panel B. Fluorescent signal generated by the CellTox™ Green Dye binding to DNA from dead cells was determined before addition of lytic cAMP-Glo™ Max reagent to determine cytotoxicity and after addition of the lytic reagents to determine total cell DNA levels. EC50 values were 4.13–5.76 × 107.
Next, we tested cholera toxin using this multiplexing protocol. Cholera toxin is known to increase intracellular cAMP levels. The A subunit of the toxin binds with the Gαs subunit of GPCRs, maintaining adenylate cyclase activation.
We incubated the cholera toxin dilutions for 45 minutes at 37°C with 5% CO2 and then equilibrated to room temperature for 15 minutes before beginning the CellTox™ Green cytotoxicity read. The EC50 values determined using the cAMP-Glo™ Max Assay with and without CellTox™ Green Dye were again very similar, with a small loss (<16%) of luminescent signal when the CellTox™ Green Dye was present (Figure 4, Panel A). The CellTox™ Green Assay showed that the cholera toxin was not toxic to the cells under the conditions tested as there was no significant increase in fluorescence signal (Figure 4, Panel B).
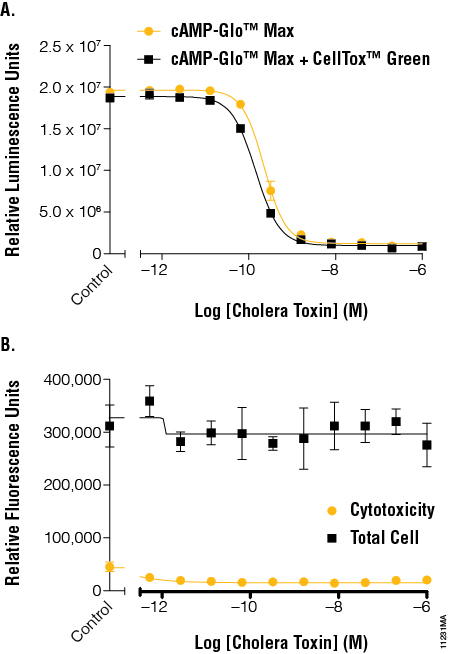
Figure 4. The effects of cholera toxin on cAMP levels in the context of cytotoxicity and cell number normalization. Panel A. A cholera toxin dilution series was added to cells and tested for cAMP activation using the cAMP-Glo™ Max Assay in the absence and presence of CellTox™ Green Dye. EC50 values were calculated for an average of 6 replicates using GraphPad Prism® software. The resulting EC50 range was 2.03–2.49 × 1010. Panel B. Fluorescent signal generated by the CellTox™ Green Dye binding DNA of dead cells was determined before addition of lytic cAMP-Glo™ Max reagent to determine cytotoxicity and after addition of the lytic reagents to determine total cell DNA levels. EC50 values were calculated for an average of 6 replicates using GraphPad Prism® software. The resulting EC50 range was 1.31–1.57 × 1010.
Conclusions
assay since the cAMP-Glo™ Max Assay has a large luminescence dynamic range. The EC50 values determined using forskolin and cholera toxin were not significantly changed by the presence of CellTox™ Green Dye.
CellTox™ Green Cytotoxicity Assay is a good choice for multiplexing with the cAMP-Glo™ Max Assay. The simplicity of a no-step addition for the CellTox™ Green Cytotoxicity Assay Dye at the time of cell dosing makes it quick and easy to detect cytotoxicity during the experiment. The average of 16% loss in luminescent signal due to addition of CellTox™ Green Dye had little impact on the