High-Concentration (>100ng/µl) Genomic DNA From Whole Blood Using Maxwell® 16
Promega Corporation
Publication Date: 2010
Abstract
We developed a novel DNA isolation chemistry with an automated format that allows genomic DNA isolation from whole blood and elution in 50 microliter volumes. This DNA isolation method does not require laborious extraction from buffy coat samples that characterizes most approaches to high-concentration (>100ng/µl) genomic DNA.
Introduction
Molecular techniques are increasingly informative, frequently allowing multiple determinations from an individual genomic DNA (gDNA) isolate. Many methods, such as microarray detection or real-time or endpoint PCR, rely upon high-concentration genomic DNA isolates (>100ng/µl) to ensure success. However, current DNA extraction methods require large input volumes of biological samples and lengthy enrichment processes, such as preparing buffy coats from whole blood, to achieve the concentrations necessary for these applications. At the same time, there is increasing pressure to automate DNA purification methods as laboratories grapple with larger sets of samples. To meet these needs, we recently developed the Maxwell® 16 LEV Blood DNA Kit (Cat.# AS1290), an automated gDNA purification system that yields high-concentration DNA from whole blood without the additional manual preprocessing required to isolate gDNA from buffy coat samples. This purification method relies upon two developments. First, we developed an efficient DNA-isolation chemistry using a novel paramagnetic particle with a binding capacity of at least 20 micrograms of genomic DNA from liquid blood samples. Second, we adapted this paramagnetic particle chemistry to the Low Elution Volume (LEV) configuration of the Maxwell® 16 series of instruments, which allows isolated gDNA to be eluted in volumes as low as 50µl.
"The Maxwell® 16 LEV Blood DNA Kit allows automated genomic DNA isolation from fresh and frozen whole blood samples, resulting in high-concentration, high-molecular-weight DNA samples that are compatible with PCR amplification."
Protocol Overview
First, 300µl of whole blood is combined with 30µl of Proteinase K (PK) Solution (provided in the Maxwell® 16 LEV Blood DNA Kit) and 300µl of Lysis Buffer (also provided in the kit). This combination is vortexed briefly and incubated for 20 minutes at 56°C. After this preprocessing step, lysates are transferred to the first well of a multiwell cartridge prefilled with reagents. Plasticware and 50µl of the provided Elution Buffer are loaded in the appropriate locations on the instrument deck, and the appropriate automated method is run. The method run time is approximately 40 minutes. The resulting eluates in elution tubes can be stored as per laboratory guidelines.
Efficient gDNA Isolation in a 50µl Elution Volume
Crucial to isolation of high-concentration gDNA from blood samples is the development of a high-efficiency gDNA isolation system. In the presence of polyethylene glycol (PEG), nucleic acids bind to a cellulose matrix (1) . For this system, a new cellulose-based paramagnetic particle with a high binding capacity was developed. We have obtained as much as 20µg of DNA from a particle volume that can be automated using the Maxwell® 16 series of instrument. The Maxwell® 16 Instrument is a paramagnetic particle-handling instrument that transfers paramagnetic particles through a series of binding and washing steps in prefilled reagent cartridges. The LEV configuration of the Maxwell® 16 Instrument was developed specifically to isolate DNA from biological samples in a 50µl elution volume (Figure 1). To achieve the desired high-concentration isolates, we developed a gDNA isolation method that rivals precipitation-based purification protocols but can be automated on the Maxwell® 16 Instrument.
 Figure 1. The Maxwell® 16 Low Elution Volume (LEV) configuration.
Figure 1. The Maxwell® 16 Low Elution Volume (LEV) configuration.
Disposable cartridges are placed in an LEV rack. Separate tubes are used to elute DNA in a 50µl volume.
To demonstrate the efficiency of this gDNA isolation chemistry and compare it to commercially available precipitation-based chemistries, we tested a precipitation-based DNA purification method and the Qiagen QIAamp® DNA Blood Mini Kit, both of which isolate gDNA from whole blood. Whole blood from a single donor was collected into Vacutainer® tubes (BD Vacutainer® K2 EDTA Plus collection tubes) and used for each purification chemistry. Genomic DNA was isolated from 300µl of fresh whole blood using the Maxwell® 16 LEV Blood DNA Kit and the precipitation-based DNA purification method, and 200µl of the same blood was processed using the QIAamp® kit. We followed the manufacturer's recommended protocol for each kit and used the recommended elution volumes. The volumes of the resulting gDNA isolates were measured, and DNA concentrations were estimated using a NanoDrop® (ND-1000) spectrophotometer. Yield was estimated by multiplying gDNA concentration by elution volume. These yield estimates were compared to theoretical gDNA yields, which were based on the white blood cell count of the blood samples. In this comparison, the Maxwell® 16 LEV Blood DNA Kit resulted in the highest recovery as a percentage of the available gDNA (Figure 2).
 Figure 2. DNA recovery from whole blood samples.
Figure 2. DNA recovery from whole blood samples.
Genomic DNA was isolated from 200µl of liquid blood using the QIAamp® Blood Kit and from 300µl of liquid blood using the Maxwell® 16 LEV Blood DNA Kit and a precipitation-based DNA purification method. All three sample sets were derived from the same volume of liquid blood on the same day. One standard deviation is indicated for each set.
One research goal was to develop a gDNA isolation system that is as efficient as precipitation-based DNA purification, and another was to develop an automated method that isolates DNA in as little as 50µl to yield a high-concentration gDNA eluate. Using the manufacturer's elution volume recommendation, the Maxwell® 16 LEV Blood DNA Kit provided a higher DNA concentration than the other kits tested (Figure 3).
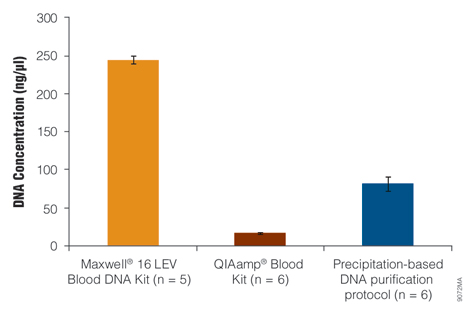 Figure 3. Concentration of DNA isolated from whole blood.
Figure 3. Concentration of DNA isolated from whole blood.
Genomic DNA was isolated from 200µl of liquid blood using the QIAamp® Blood Kit and from 300µl of liquid blood using the Maxwell® 16 LEV Blood DNA Kit and a precipitation-based DNA purification method. All three sample sets were derived from the same volume of liquid blood on the same day. One standard deviation of values is indicated for each treatment set.
DNA Purification Method is Compatible with Common Anticoagulants and Storage Conditions
Whole blood was collected from two individuals into EDTA, sodium citrate and sodium heparin blood collection tubes and stored at 4°C. Genomic DNA was isolated from 300µl of blood from each individual in each collection tube type using the Maxwell® 16 LEV Blood DNA Kit and eluted in a final elution volume of 50µl. Each DNA sample was quantified using the NanoDrop® ND-1000 spectrophotometer. The resulting concentrations and purity (A260/A280 and A260/A230 values) demonstrate that neither yield nor sample purity were affected by the blood collection tube type (Figure 4).
 Figure 4. Results of the Maxwell® 16 LEV Blood DNA Kit with EDTA, sodium citrate and sodium heparin collection tubes.
Figure 4. Results of the Maxwell® 16 LEV Blood DNA Kit with EDTA, sodium citrate and sodium heparin collection tubes.
Blood from two individuals was collected into EDTA, sodium citrate and sodium heparin collection tubes, then processed using the Maxwell® 16 LEV Blood DNA Kit. The resulting gDNA eluates were quantified using the NanoDrop® ND-1000 spectrophotometer to estimate DNA concentrations (Panel A) and A260/A280 and A260/A230 ratios (Panel B). n = 4
To address potential effects of anticoagulant type on downstream PCR results, we performed the following experiment: We incubated sterile phosphobuffered saline overnight in each tube type, processed these samples in parallel with the other samples and used the resulting eluates as reagent blanks when co-amplifying known amounts of human DNA in a sensitive, PCR-based STR multiplex (PowerPlex® 16 System, Cat.# DC6530). Amplified products were detected using capillary electrophoresis, and the resulting amplification signal was analyzed. No PCR inhibition was detected (data not shown).
Fresh versus Frozen Blood Samples
Blood samples from four individuals were obtained in EDTA collection tubes. Each sample was stored fresh (4°C) or frozen (–80°) overnight. Frozen samples were equilibrated to room temperature, and 300μl of fresh or frozen whole blood was processed using the Maxwell® 16 LEV Blood DNA Kit. DNA was eluted in 50µl of Elution Buffer. Each sample was processed in duplicate. The resulting eluates were quantified using the NanoDrop® ND-1000 spectrophotometer. The average DNA concentrations demonstrate comparable gDNA isolation results from fresh and frozen blood samples (Figure 5).
 Figure 5. Results from fresh and frozen blood samples from four donors.
Figure 5. Results from fresh and frozen blood samples from four donors.
Each sample was processed using the Maxwell® 16 LEV Blood DNA Kit, then characterized using a NanoDrop® (ND-1000) spectrophotometer to estimate DNA concentrations (Panel A) and A260/A280 and A260/A230 ratios (Panel B).
High-Molecular-Weight Genomic DNA is Suitable for Amplification
The Maxwell® 16 LEV Blood DNA Kit is designed primarily to isolate high-concentration gDNA eluates, but it also is important that the gDNA is not fragmented or degraded. To gauge DNA integrity, blood samples were obtained from four individuals and stored overnight at 4°C or –80°C. All samples were brought to room temperature and mixed briefly, then 300µl of blood was processed using the Maxwell® 16 LEV Blood DNA Kit. DNA was eluted in 50µl of Elution Buffer. The purified DNA was separated by gel electrophoresis in a 0.7% agarose gel. High-molecular-weight Human Genomic DNA (Cat.# G3041) was run as a positive control, while NANOpure® water was run as a negative control. Lambda DNA/HindIII Markers (Cat.# G1711) also were run to estimate the size of the resulting gDNA fragments. The gel was stained with ethidium bromide, destained and visualized with a UV transilluminator (Figure 6). The size of the gDNA was comparable to that of the high-molecular-weight control DNA. Eluates processed using the Maxwell® 16 LEV Blood DNA Kit were successfully amplified in a multiplex STR amplification (PowerPlex® 16 System) and in real-time qPCR using the Plexor® HY System (Cat.# DC1001).
 Figure 6. Agarose gel electrophoresis results from fresh and frozen blood samples.
Figure 6. Agarose gel electrophoresis results from fresh and frozen blood samples.
Three hundred microliters of fresh or frozen liquid blood from four donors was processed using the Maxwell® 16 LEV Blood DNA Kit, and the DNA was separated by agarose gel electrophoresis. Lane 1, donor 1, fresh blood; lane 2, donor 2, fresh blood; lane 3, donor 3, fresh blood; lane 4, donor 4, fresh blood; lane 5, donor 1, frozen blood; lane 6, donor 2, frozen blood; lane 7, donor 3, frozen blood; lane 8, donor 4, frozen blood, lane 9, Human Genomic DNA (Cat.# G3041); lane 10, NANOpure® water as the negative control; lane M, Lambda DNA/HindIII Markers.
Summary
We have developed a new automated system for the Maxwell® 16 series of instruments to efficiently isolate high-concentration, high-purity genomic DNA from whole blood. This system uses a new, cellulose-based paramagnetic particle that allows microgram amounts of gDNA to be isolated and eluted in 50μl volumes. This DNA isolation protocol is compatible with commonly used blood storage and collection methods. Downstream analysis indicates that gDNA isolated using the Maxwell® 16 LEV Blood DNA Kit is of high molecular weight and allows robust PCR amplification.
Acknowledgment
The authors acknowledge the assistance of Steve Ekenberg and Brian Andersen, both of Promega Corporation, in the design and development of the MagnaCel™ paramagnetic particle.
Related Products
Related Protocols
Related Resources
Article References
- Su, X. and Comeau, A.M. (1999) Cellulose as a matrix for nucleic acid purification. Anal. Biochem. 267, 415–8.
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Mandrekar, P. V., Ma, Z., Krueger, S. and Cowan, C. High-Concentration (>100ng/µl) Genomic DNA From Whole Blood Using the Maxwell® 16 Low Elution Volume Instrument. [Internet] 2010. [cited: year, month, date]. Available from: https://www.promega.com/resources/pubhub/high-concentration-genomic-dna-from-whole-blood-using-the-maxwell-16-low-elution-volume-instrument/
American Medical Association, Manual of Style, 10th edition, 2007
Mandrekar, P. V., Ma, Z., Krueger, S. and Cowan, C. High-Concentration (>100ng/µl) Genomic DNA From Whole Blood Using the Maxwell® 16 Low Elution Volume Instrument. Promega Corporation Web site. https://www.promega.com/resources/pubhub/high-concentration-genomic-dna-from-whole-blood-using-the-maxwell-16-low-elution-volume-instrument/ Updated 2010. Accessed Month Day, Year.
Maxwell, Plexor and PowerPlex are registered trademarks of Promega Corporation. MagnaCel is a trademark of Promega Corporation.
NanoDrop is a registered trademark of Thermo Fisher Scientific. NANOpure is a registered trademark of Barnstead/Thermolyne Corporation. QIAamp is a registered trademark of Qiagen GmbH Corp. Vacutainer is a registered trademark of Becton, Dickinson and Company.
Products may be covered by pending or issued patents or may have certain limitations. Please visit our web site for more information.