Functional Proteomics Techniques to Isolate and Characterize the Human Proteasome
Brad Hook and Trista Schagat, Promega Corporation
Publication Date: 2011
Abstract
Identifying protein subunits is a routine application of mass spectrometry. With advances in this technology, characterization of complexes has become accessible to more researchers. Using the human proteasome as our model complex, we demonstrate how new commercial technologies and services allow researchers to quickly characterize a complex with little upfront effort. We combined HaloTag® technology with the services from the Kazusa Institute and MS Bioworks to rapidly identify and characterize subunits in the human proteasome. Using a luciferase-based assay, we were able to assay our complexes for protease activity before identifying the proteins using mass spectrometry. Finally, we used cell-free protein expression to quickly map protein interactions in the human proteasome. This integrated project design required no traditional cloning, little optimization and minimal hands-on time.
Introduction
The proteasome is a multi-catalytic complex in the nucleus and cytosol of all eukaryotic cells that is responsible for proteolysis of ubiquitin-tagged proteins. The 26S proteasome is a large multi-subunit complex containing a 20S proteolytic core particle and a pair of 19S regulatory particles (Figure 1; 1,2). The 20S core is a barrel-shaped assembly of protein subunits that possesses three different proteolytic activities (3,4). The 19S regulatory particles assemble at both ends of the 20S core. The particles recognize ubiquitinized proteins, deubiquitinate them and transfer them into the 20S core for degradation. The 26S proteasome degrades polyubiquitinated proteins in an ATP-dependent manner. This complex processes aberrant and misfolded proteins as well as proteins regulating cell cycle, growth and apoptosis and is essential for cellular function. One of the key aspects of research attained by studying the structure and function of the proteasome is the understanding of how this complex regulates biological processes.
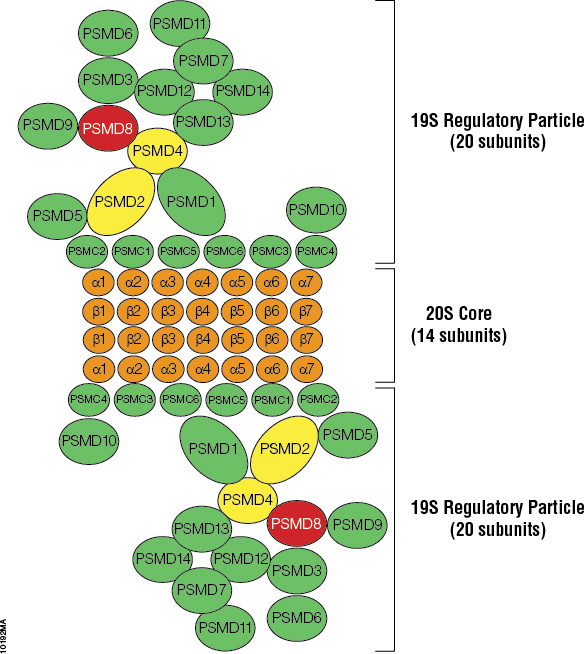
Figure 1. Model of the human proteasome. Red subunits were used in pull-down assays and analyzed with mass spectrometry. Yellow and red subunits were used in protein:protein mapping experiments. Green subunits are found in the 19S regulatory particle. Orange subunits are found in the 20S core. Models were based on yeast proteasome and human mapping data.
We sought to identify subunits of the human proteasome using an assay design that minimized time by integrating commercial products and services. Critical steps in the project used products and services from Kazusa DNA Research Institute, Promega Corporation and MS Bioworks (Figure 2).
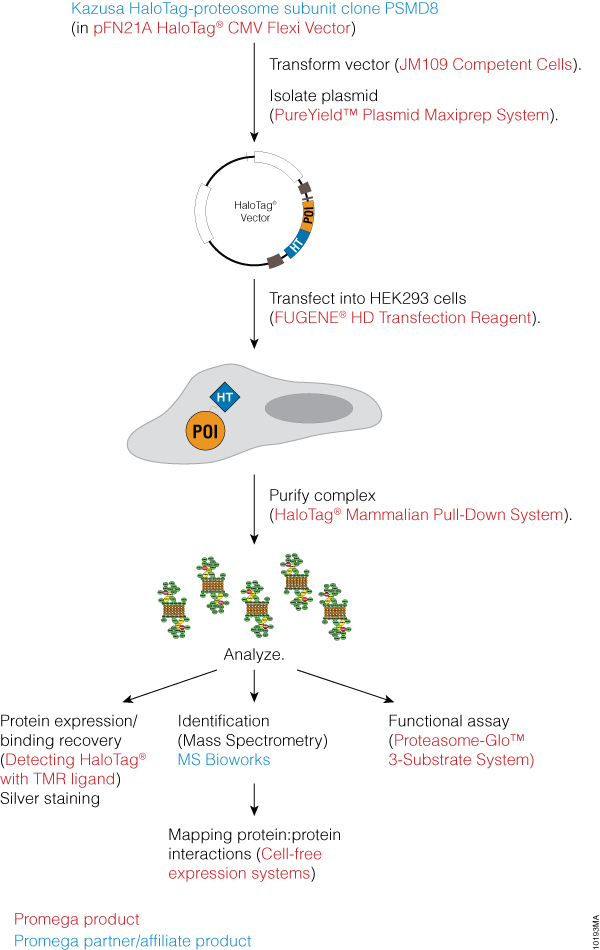
Figure 2. Project design to characterize the human proteasome. Red lettering indicates Promega produces and blue lettering indicates Promega partners.
The initial construct was purchased from Kazusa Institute, which allowed a rapid start to the project with no need for cloning. The construct contained a proteasome subunit fused to the HaloTag® protein, a novel protein fusion tag that allows highly specific, oriented and covalent immobilization of proteins on surfaces. The proteasome subunit was transfected into HEK293 cells using FuGENE® HD Transfection Reagent (Cat.# E2311) and isolated using the HaloTag® Mammalian Pull-Down System (Cat.# G6504). Analysis was performed by MS Bioworks using mass spectrometry. The proteasome was assayed for function using the luciferase-based Proteasome-Glo™ 3-Substrate System Assay (Cat.# G8531). Protein:protein interactions were mapped using TnT® SP6 High-Yield Wheat Germ Protein Expression System (Cat.# L3260). Using commercially available products and services allowed us to rapidly analyze this multi-protein complex.
Methods
Templates
Three constructs were purchased from the Kazusa Institute, each containing one of the human proteasome subunits. PSMD2 (Kazusa Institute Cat.# FHC01526), PSMD4 (Kazusa Institute Cat.# FHC03468) and PSMD8 (Kazusa Institute Cat.# FHC02693) are in the pFN21A HaloTag® CMV Flexi® Vector (Cat.# G2821), which contains an amino-terminal HaloTag® protein, a CMV promoter and a T7 promoter. The HaloTag® Control Vector (Cat.# G6591) was used as a pull-down control. Vectors were transformed into JM109 Competent Cells (Cat.# L1001, L2001), grown overnight in Terrific Broth at 37°C and purified using the PureYield™ Plasmid Maxiprep System (Cat.# A2392). For cell-free protein expression in the TnT® SP6 High Yield Expression system, each proteasome subunit was transferred to the pFN19K HaloTag® T7 SP6 Flexi® Vector (Cat.# G1841) using the Flexi® System, Entry/Transfer (Cat.# C8640).
Collecting Mammalian Cells Expressing HaloTag® Fusion Proteins
For processing, 1 × 107 HEK293 cells were plated into T150 flasks (30ml) for each sample. The cells were grown overnight in DMEM medium (containing 10% FBS) at 37°C and 5% CO2. HaloTag® Control Vector (Cat.# G6591) and the vectors containing the three proteasome subunits were transfected into the HEK293 cells using FuGENE® HD Transfection Reagent (Cat.# E2311, see #TM328 for details). The cells were harvested 48 hours after transfection. The media was removed, and the cells were washed with 25ml of cold PBS. After removing PBS, 30ml of fresh, cold PBS was added to the flasks. The cells were scraped from the surface and collected into conical tubes. The cells were centrifuged at 4°C for 5 minutes at 2,000 × g, and the PBS was discarded. The cell pellets were stored overnight at –80°C.
Pull-Down of the Proteasome
We used the HaloTag® Mammalian Pull-Down and Labeling Systems (Cat.# G6500), following the protocol given in the HaloTag® Mammalian Pull-Down and Labeling System Technical Manual #TM342, to isolate the proteasome. Proteins were eluted from the HaloLink™ Resin (Cat.# G1912) in 50ul of buffer using ProTEV Plus protease (30 units; Cat.# V6101). ProTEV Plus protease cleaves an optimized TEV cleavage site that was engineered into the pFN21A HaloTag® CMV Flexi® Vector. The protocol for cleavage is found in the HaloTag® Mammalian Pull-Down and Labeling System Technical Manual #TM342. The ProTEV Plus protease was not removed from eluates.
Detecting HaloTag® Fusion Proteins
Detailed instructions for detecting HaloTag® fusion proteins is available in the HaloTag® Mammalian Pull-Down and Labeling System Technical Manual #TM342. A 10μl aliquot of diluted lysate was added to 19μl of water and 1μl of 50μM HaloTag® TMR Ligand (Cat.# G8251, G8252). Following a 15-minute, room-temperature incubation, 10μl of 4X SDS gel loading buffer was added. The samples were heated to 70°C for 5 minutes, and 10μl was loaded onto a 4–20% SDS-polyacrylamide gel. The gel was run at 150V for 1.5 hours, and the bands were detected using a BioRad XR imager.
Proteasome Activity Assay
The Proteasome-Glo™ 3-Substrate System (Cat.# G8531) was used to detect three types of protease activities found in the proteasome, following the protocol given in the Proteasome-Glo™ 3-Substrate System Technical Bulletin, #TB349. A 10μl aliquot of each of the eluates was added to 40μl of 1mM HEPES (pH 7.5) in a white 96-well plate. Next, 50μl of the reconstituted Proteasome-Glo™ Reagent (see #TB349, Reagent Preparation) was added to each well. After mixing at 500rpm for 30 seconds on a plate shaker, the samples were incubated at room temperature for 30 minutes. Luminescence was recorded using a GloMax® Multi Luminometer (Cat.# E7031). A background control consisting of 40μl of 1mM HEPES buffer (pH 7.5) and 10µl of 1X ProTEV Plus Buffer was included. To ensure that the ProTEV Plus in the eluates was not exhibiting the protease activities found in the Proteasome-Glo™ 3-Substrate System, 30 units of ProTEV Plus was added to 1mM HEPES buffer (pH 7.5; final volume 50μl) and tested in the assay.
Gel Analysis of Eluates
A 10μl aliquot of each sample was combined with 10μl of 2X SDS loading dye, heated to 95°C for 5 minutes, and 20μl was loaded onto a 4–20% SDS-polyacrylamide gel. The gel was run at 150V for 1.5 hours. The gel was placed in water and then stained using Pierce silver stain kit, following the manufacture’s instructions.
Mass Spectrometry
Mass spectrometry was performed by MS Bioworks (www.msbioworks.com). Samples were separated using SDS-PAGE, stained using Coomassie®, and each lane was excised into ten equally sized segments. Gel pieces were processed using a robot (ProGest, DigiLab). Each gel digest was analyzed by nano LC/MS/MS with a Waters NanoAcquity HPLC system interfaced to a ThermoFisher LTQ Orbitrap Velos. Peptides were loaded on a trapping column and eluted over a 75μm analytical column at 300nL/minute; both columns were packed with Jupiter Proteo resin (Phenomenex). The mass spectrometer was operated in data-dependent mode, with MS performed in the Orbitrap at 60,000FWHM resolution and MS/MS performed in the LTQ. The fifteen most abundant ions were selected for MS/MS. Data was processed by MS Bioworks using Mascot. Mascot DAT files were parsed into the Scaffold software for validation, filtering and to create a non-redundant list for each sample. Data were filtered using a minimum protein value of 90%, a minimum peptide value of 50% (Prophet scores) and requiring at least two unique peptides per protein.
Protein Mapping using Cell-Free Expressed Proteins
Cell-free expression reactions: HaloTag®-PSMD2, 4 and 8 and HaloTag®-GFP were expressed from the pFN19K HaloTag® T7 SP6 Flexi® Vector (Cat.# G1841) using the TnT® SP6 High-Yield Wheat Germ Protein Expression System following the instructions in the TnT® SP6 High-Yield Wheat Germ System Technical Manual #TM282. Ten micrograms of plasmid was used for each 50µl reaction. Reactions were incubated at 25°C for 2 hours. Prey protein: 1µl of HaloTag® TMR Ligand (0.5mM; Cat.# G8251, G8252) was added to each 50μl reaction. Labeling was performed at room temperature for 15 minutes. The reactions were stored on ice. Bait protein: For each interaction assay a 50μl cell-free reaction was prepared. For each reaction, a 10μl aliquot of settled HaloLink™ magnetic beads was equilibrated with TMNEN150 buffer (50mM Tris [pH8.0], 0.5% IGEPAL®, 0.5mM EDTA, 2mM MgCl2, and 150mM NaCl). Each 50μl cell-free reaction was added to the 10μl of settled beads and incubated with mixing at room temperature for 15 minutes. Following HaloTag binding, the beads were washed three times with 500μl TMNEN150. Binding reaction: 10μl of the labeled prey reaction was added to the 10μl of HaloTag®-fusion bound beads. Reactions were incubated at 4°C for 2 hours. After binding, the beads were washed 4 times with 500μl of TMNEN150. Analysis: 20μl of 2X SDS was added to the settled beads and incubated at 70°C for 10 minutes, and 15μl of each reaction was added to a 4–20% SDS-PAGE gel. After separation using electrophoresis, a BioRad XR imager was used to detect the HaloTag® TMR Ligand. In these experiments HaloTag®-GFP was used as a negative control.
E. coli were transformed with the HaloTag®-PSMD2, PSMD4 and PSMD8 plasmids purchased from the Kazusa Institute. The plasmid DNA was purified using the PureYield™ Plasmid Maxiprep System (Cat.# A2392). The HaloTag®-PSMD2-, PSMD4- and PSMD8-containing plasmids as well as the HaloTag® Control Vector were then transfected into HEK293 cells. Cells were harvested after 48 hours and lysed using the lysis buffer provided in the HaloTag® Mammalian Pull-Down System. The HaloTag® fusion proteins were labeled with TMR Ligand and analyzed using SDS-PAGE (Figure 3). All the HaloTag®-proteasome subunit fusion proteins and the 35kDa HaloTag® control protein were expressed and detected in the cell lysates. The lysates were incubated with HaloLink™ Resin and then washed using buffer provided in the HaloTag® Mammalian Pull-Down System. The proteins that did not bind the HaloLink™ Resin (flowthrough) also were labeled with TMR Ligand and analyzed using SDS-PAGE (Figure 3).
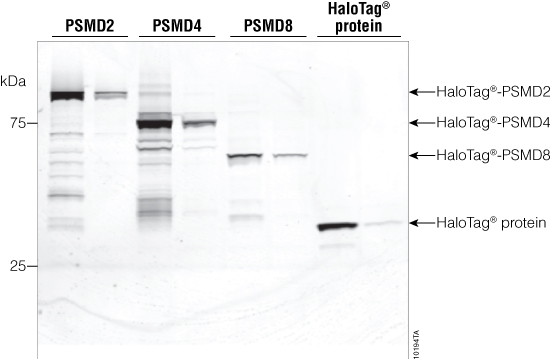
Figure 3. SDS-PAGE analysis of proteasome subunits. Samples were labeled using the HaloTag® TMR Ligand, and the PSMD2 (135kDa), PSMD4 (76kDa) and PSMD8 (65kDa) proteins were separated using SDS-PAGE. Lanes 1, 3, 5 and 7: lysates containing TMR-labeled, expressed HaloTag® proteins. Lanes 2, 4, 6 and 8: flowthrough off the HaloLink™ Resin. All the proteasome subunits are HaloTag® fusion proteins. Lanes 7 and 8 show expression and depletion of the HaloTag® protein (35kDa) alone.
Results
A comparison of the lysate and flowthrough lanes indicates that most of the protein was depleted or bound to the HaloLink™ Resin. The bound complexes were eluted with ProTEV Plus protease, and the eluates were analyzed by SDS-PAGE and silver stain (Figure 4). Since the ProTEV Plus protease (48kDa) was not removed from the eluates, some of the bands in each of the lanes are from ProTEV Plus. The ProTEV Plus control lane shows the bands that are associated with ProTEV Plus. All HaloTag®-PSMD pull-down eluates had many unique proteins compared to the HaloTag® control.
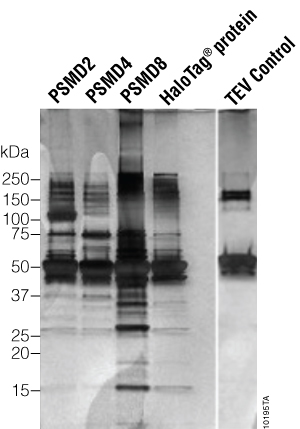
Figure 4. Silver-stained SDS-PAGE analysis of eluates from the HaloTag® Pull-Down. Labels under the figure specify the protein used for the pull-down. “TEV con” is a ProTEV Plus protease control in which an equivalent amount of protease as in each pull-down experiment is added to the gel.
Before sending samples for mass spectrometry analysis, eluates were assayed for proteolytic activity. The Proteasome-Glo™ 3-Substrate System consists of three homogeneous bioluminescent assays that measure the three proteolytic activities associated with the proteasome: trypsin-like, chymotrypsin-like and caspase-like activities (Figure 5, Panel A). Each substrate detects a different protease activity. Each substrate is added to a buffer system optimized for proteasome and luciferase activity to make a Proteasome-Glo™ Reagent. The substrate-specific Proteasome-Glo™ Reagent is added to test samples in an “add-mix-measure” format. Substrate cleavage generates a luminescent signal produced by the luciferase reaction, with the signal proportional to the amount of proteasome activity. Eluates from the PSMD2, PSMD4, PSMD8 and HaloTag® control pull-down reactions were assayed for activity using the Proteasome-Glo™ 3-Substrate System (Figure 5, Panel B). All three proteasome subunit pull-down samples contained all three protease activities. Activity measurements ranged from 7- to 17-fold above background. The HaloTag® control sample had little protease activity. To determine if the ProTEV Plus in the samples contributes to the protease signal observed, ProTEV Plus was independently tested for activity in the Proteasome-Glo™ System. These samples resulted in no significant activity, indicating the activity from the proteasome pull-down eluates are from isolated proteasome and not background proteins or ProTEV Plus (data not shown).
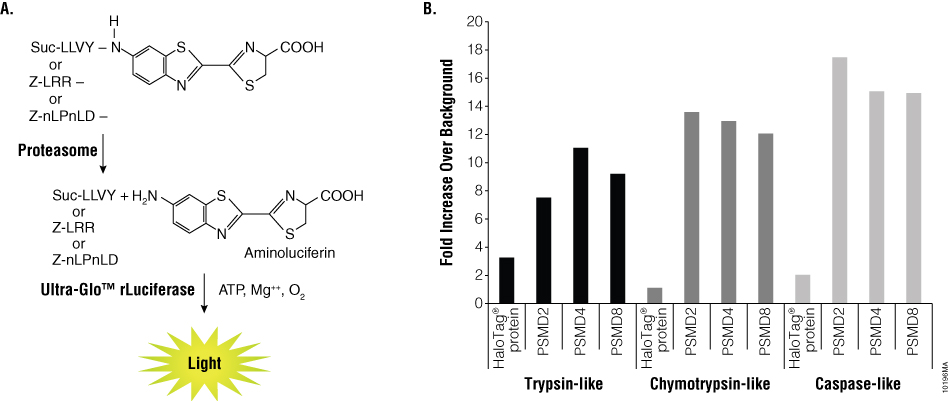
Figure 5. Analysis of sample activity using the Proteasome-Glo™ 3-Substrate System. Panel A. The luminogenic substrates containing the Suc-LLVY, Z-LRR or Z-nLPnLD sequence are recognized by the 20S proteasome. Following cleavage by the 20S proteasome, the substrate for luciferase (aminoluciferin) is released, resulting in light production by the luciferase reaction. Panel B. Graph showing sample activity in each of the substrates in the Proteasome-Glo™ System (trypsin, chymotrypsin and caspase).
Having determined that the proteasome pull-down samples contained active proteasome subunits, we sent only one sample (PSMD8) and one control (HaloTag® control) to MS Bioworks for mass spectrometric analysis. The samples were analyzed as described in the methods section. The mass spectrometric analysis detected 31 of the 34 proteasome subunits (Figure 6). The three missing subunits were small 20S core proteins and might have been missed because of size or inaccessibility. Mass spectrometric analysis of the HaloTag® control pull-down sample did not detect any proteasome subunits. As expected, the Normalized Spectral Abundance Factor numbers show that PSMD8 was the most abundant subunit detected. The results show a high percentage enrichment of proteasome subunits.
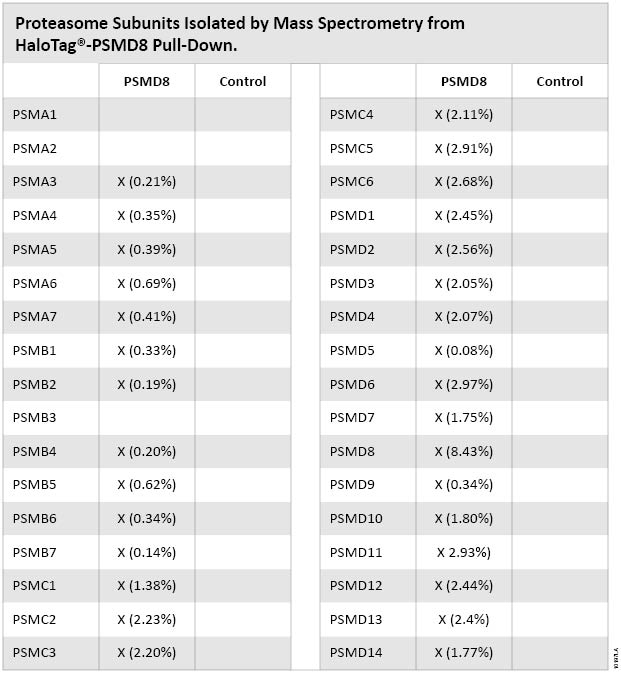
Figure 6. Proteasome subunits isolated by mass spectrometry from HaloTag®-PSMD8 pull-down. “Control” is HaloTag® pull-down. Numbers correspond to NSAF% (Normalized Spectral Abundance Factor).
To demonstrate the advantages of using cell-free translation systems in protein mapping experiments, we mapped the interactions between PSMD2, 4 and 8. Figure 7, Panel A, diagrams the experimental procedure. Each of the subunits were expressed in a cell-free system as HaloTag® fusion proteins and labeled using the fluorescent HaloTag® TMR Ligand (prey). In separate cell-free synthesis reactions, the same PSMD2, 4, 8 HaloTag® fusion proteins were expressed and immobilized on a solid surface by interaction between the HaloTag® protein and HaloLink™ Magnetic Beads (bait). Combinations of the prey and bait were mixed together, washed, eluted and detected using SDS-PAGE and a fluoroimager (Figure 7, Panel B). HaloTag®-GFP was used as a negative control (data not shown). Each subunit was successfully expressed and labeled with the HaloTag® TMR Ligand and visible in the input lanes. A positive interaction was observed between immobilized PSMD2 and prey PSMD4 but not prey PSMD8. The immobilized PSMD4 bound both PSMD2 and PSMD8, and immobilized PSMD8 bound PSMD4 but not PSMD2. These studies map direct interactions between these proteins that is consistent with studies published for the yeast proteasome.
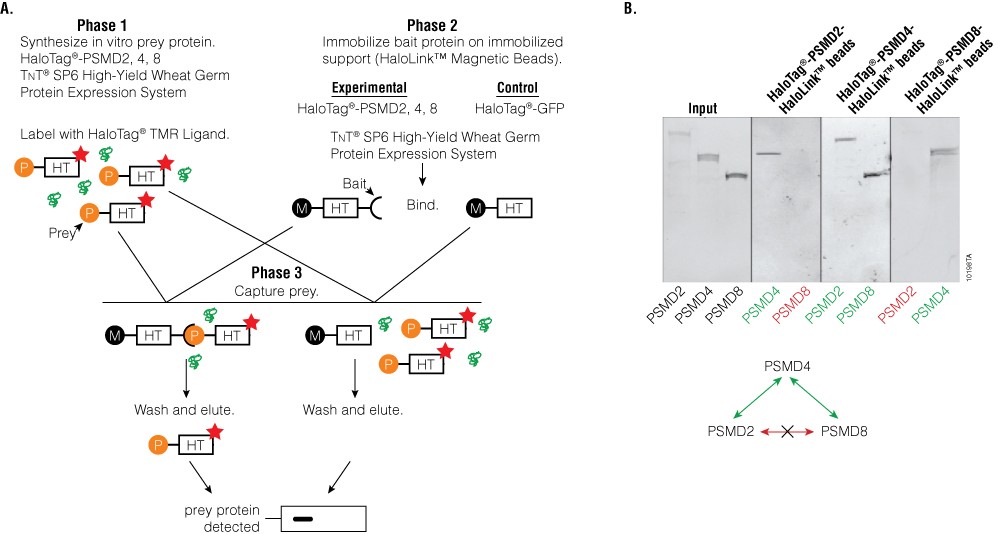
Figure 7. Using cell-free expression to map proteasome interactions. Panel A. Protein:protein interaction scheme. Orange circles represent TMR-labeled HaloTag-fused PSMD2, 4, 8. Black circles represent HaloLink™ Magnetic Beads. HT = HaloTag® protein. Panel B. SDS-PAGE analysis of protein:protein interactions. The labels above the gels indicate the bait proteins, and the labels below the gel indicate prey proteins. Red lettering represents no interaction, and green lettering represents positive interaction. The diagram below the gel represents the interaction map between these proteasome subunits.
Conclusion
We used commercial products and services from Promega Corporation, Kazusa Institute and MS Bioworks to quickly isolate subunits of the human proteasome, test their activity and characterize the subunit interactions. The proteasome constructs contained one of the human proteasome subunits fused to the HaloTag® Protein. Using HaloTag® technologies, we were able to rapidly isolate the functional proteasome from HEK293 cells, label them for fluorescent detection and analysis of complex subunits using mass spectrometry. Finally, cell-free protein expression was used as an easy and fast method for confirming and mapping protein:protein interactions discovered by mass spectrometry.
References
- Chen, C. et al. (2008) Subunit-subunit interactions in the human 26S proteasome. Proteomics 8, 508–20.
- Guerrero, C. et al. (2008) Characterization of the proteasome interaction network using a QTAX-based tag-team strategy and protein interaction network analysis. Proc. Natl. Acad. Sci. USA 105, 13333–8.
- Rechsteiner, M. and Hill, C.P. (2005) Mobilizing the proteolytic machine: Cell biological roles of proteasome activators and inhibitors. Trends Cell. Bio. 15, 27–33.
- Kisselev, A.F. et al. (2002) Binding of hydrophobic peptides to several non-catalytic sites promotes peptide hydrolysis by all active sites of 20S proteasomes. J. Biol. Chem. 277, 22260–70.
Learn more about the Proteasome-Glo™ Assays