Mix and Match Primary and Control Reporters using the NanoDLR™ Dual-Luciferase Assay System
Why Choose a Dual-Reporter Assay?
Dual-Reporter Assays give scientists the ability to simultaneously measure two reporter enzymes within a single sample. In dual assays, the activity of a primary reporter is correlated with the effect of specific experimental conditions, while the activity of a control reporter relays the baseline response, providing an essential internal control that reduces variability caused by differences in cell number, lysis efficiency, cell viability, transfection efficiency, temperature, and measurement time.
Normalizing data by calculating the ratio of the experimental reporter over the control reporter can help minimize some types of experimental variability and permit better comparisons between measurements on different luminometers or at different times.
The Nano-Glo® Dual-Luciferase® Reporter (NanoDLR™) Assay provides a choice of two sensitive reporters (firefly and NanoLuc® luciferases) for use in dual-assay format. Both reporters give state-of-the-art functionality, raising the question “Which luciferase should be used as the primary reporter and which should be the control?
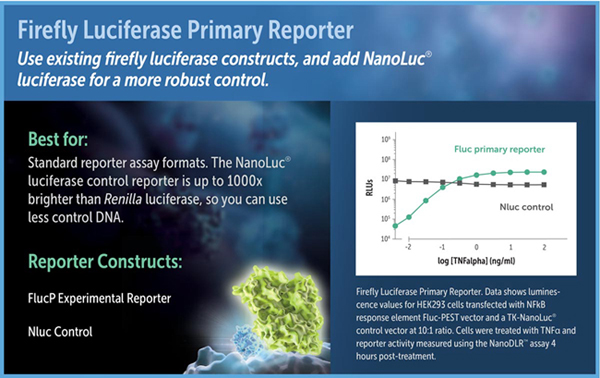
When to Choose Firefly Luciferase as a Primary Reporter
Just as in the older DLR™ and Dual-Glo® Dual-Luciferase Assays, in the NanoDLR™ assay firefly luciferase can be used as the primary reporter to measure a biological response, with NanoLuc® luciferase replacing Renilla luciferase as the constitutive control. In this assay format, firefly luciferase or firefly luciferase-PEST (luc2P) can be used as the experimental reporter. The high-intensity luminescence and stable nature of the NanoLuc® protein distinguish NanoLuc® from Renilla luciferase. When expressed in mammalian cells under equivalent conditions, NanoLuc® luminescence using the NanoDLR™ Assay is often about 1,000-fold brighter than Renilla luminescence using the Dual-Glo® Assay. As a result, lower amounts of control plasmid can be transfected for NanoLuc® luciferase, potentially minimizing transcriptional squelching of the experimental reporter. If sensitivity is an issue for the control, NanoLuc® and the NanoDLR™ Assay will provide the brightest possible signal.
Luciferase reporter vectors suitable for use in assay formats with a firefly luciferase primary reporter include pGL4 Vectors containing the firefly luciferase-PEST sequence, and the pNL1.1.PGK[Nluc/PGK] (Cat.# N1441), pNL1.1.TK[Nluc/TK] Vector (Cat.# N1501), 6r pNL1.1.CMV[Nluc/CMV] (Cat.# N1091) NanoLuc® luciferase control vectors.
When to Choose NanoLuc® Luciferase as a Primary Reporter
For the brightest possible signal and maximal response, use NanoLuc®-PEST luciferase as the experimental reporter and firefly luciferase as the constitutive control. When expressed in mammalian cells under equivalent conditions, the signal from NanoLuc®-PEST using the NanoDLR™ Assay is often more than tenfold brighter than the signal from luc2-PEST using the Dual-Glo® Assay. Because of its short intracellular protein half-life, the NanoLuc®-PEST fusion will also respond more quickly and typically with a greater signal-to-background ratio compared to firefly luciferase or luc2-PEST. Unfused NanoLuc® luciferase also can be used as the experimental reporter to maximize luminescence, but the longer protein half-life can provide a slower response to changes in gene transcription and lower signal-to-background ratios compared to firefly luciferase, luc2-PEST or NanoLuc®- PEST.
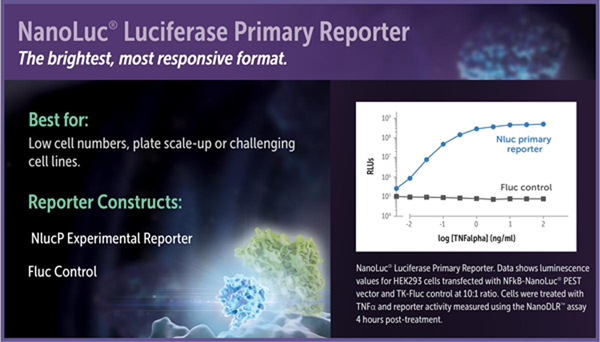

Luciferase reporter vectors suitable for use in assay formats with a NanoLuc® Luciferase primary reporter include the NanoLuc® genetic reporter vectors and the pGL4.53[luc2/PGK] (Cat.# E5011), pGL4.54[luc2/TK] (Cat.# E5061), pGL4.50[luc2/CMV/Hygro] (Cat.# E1310) and pGL4.51[luc2/CMV/Neo] (Cat.# E1320) firefly luciferase control vectors.
Options for Using Two Primary Reporters
The sensitivity of the NanoDLR™ Assay also enables assay formats where both luciferases are used as primary reporters. The potent inhibition of firefly luciferase in the NanoDLR™ Assay assures that basal levels of NanoLuc®-PEST expression can be detected without the risk of signal bleedthrough, even when luc2-PEST expression levels are very high. This assay format can increase the amount of information that is obtained from a given sample. However, without the internal control reporter, it is important that the experiment is designed to minimize variability from cell number, transfection efficiency, pipetting volumes, etc. The most responsive reporters are the PEST-destabilized forms of firefly and NanoLuc® luciferase.
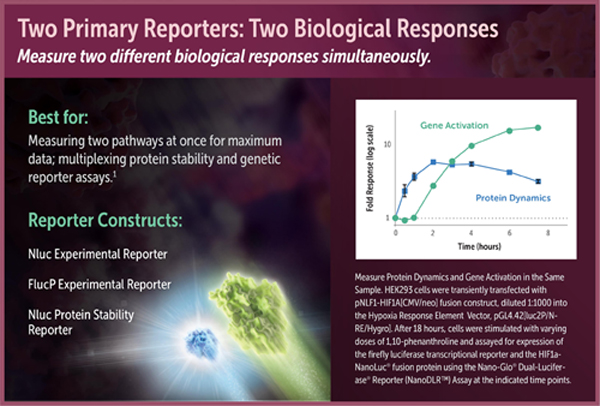
Firefly Transcriptional Reporter Plus Protein Fusion to NanoLuc® Luciferase
One assay format enabled by the NanoDLR™ Assay is coupling transcriptional readout of a firefly reporter gene to measurement of the stability of a protein fusion to NanoLuc® luciferase. For instance, many stress response pathways are regulated by dynamic protein turnover of transcription factors. Monitoring the luminescence of NanoLuc® fusions can be a simple way to follow this regulated event that occurs upstream of transcription and translation. Because of its brightness and small size, NanoLuc® luciferase is an ideal fusion partner to act as a protein stability sensor. Brightness is critical because it allows the NanoLuc® fusion protein to be expressed at much lower concentrations than fusions to other luciferases, i.e., at more physiologically relevant levels. Therefore, the most relevant biological response will generally be obtained by expressing the NanoLuc® fusion protein off a relatively weak constitutive promoter, transfecting low amounts of DNA, or diluting the DNA into promoterless carrier DNA (e.g., Transfection Carrier DNA; Cat.# E4881).
Luciferase reporter vectors suitable for use in assay formats with two primary reporters include pGL4 Vectors containing the firefly luciferase-PEST sequence and the NanoLuc® genetic reporter vectors. For protein stability studies, we offer the protein stability vectors pNLF1-HIF1A [CMV/neo] (Cat.# N1381) and pNLF1-NRF2 [CMV/neo] (Cat.# N1391).
Coincidence Reporter System
High-throughput screens using reporter gene assays can have a significant number of non-relevant hits generated due to direct interaction of compounds with the reporter enzyme. Compounds truly modulating the biological pathway of interest can be differentiated from those affecting the stability or activity of the reporter enzyme by using a coincidence reporter system in which transcriptional activation leads to stoichiometric expression of two orthologous reporters with dissimilar profiles of compound interference (1). Firefly and NanoLuc® luciferases make a particularly attractive pair of orthologous reporters because of the differences in their respective substrates and their low overlap for interfering compounds (2).
In the coincidence reporter system, firefly luciferase and NanoLuc®-PEST luciferase are expressed from the same promoter, and stoichiometric amounts of the non-fused reporter enzymes are generated using ribosome skipping mediated by the viral P2A peptide sequence (3). Following insertion of the promoter of interest into the luc2-2A-NlucP construct, transcriptional activation is measured using the NanoDLR™ Assay. Compounds increasing transcription will show a coincident increase in the activity of both reporters, while compounds affecting the stability or activity of a reporter will generally show an effect on only one reporter.
Obtaining an independent readout from two distinct reporters can improve the signal-to-noise ratio and increase confidence in hits that cause only small effects on transcription. Also, because of the two bright reporters in the NanoDLR Assay™, the coincidence system is compatible with gene editing techniques, allowing detection of endogenous expression levels for more relevant screening assays.
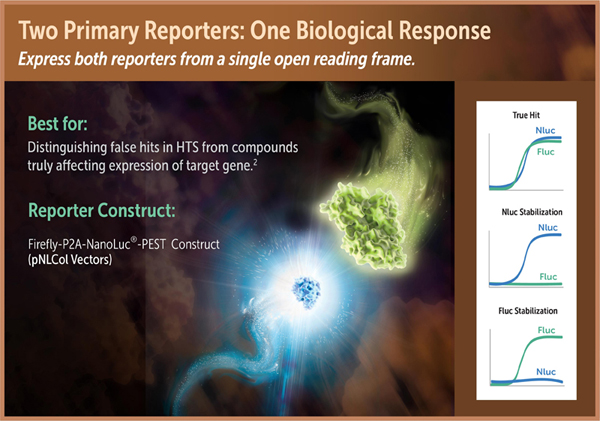

Reporter constructs for use in a coincidence reporter system contain a firefly-P2A-NanoLuc-PEST coincidence circuit from the same promoter, allowing expression of two unfused forms of luciferase with distinct compound interaction profiles. The pNLCol Vectors pNLCoI1[luc2-P2A-NlucP/Hygro] (Cat.# N1461), pNLCoI2[luc2-P2A-NlucP/minP/Hygro] Cat.# N1471), pNLCoI3[luc2-P2A-NlucP/CMV/Hygro] (Cat.# N1481) and pNLCoI4[luc2-P2A-NlucP/PGK/Hygro] (Cat.# N1491) are available for this application.
NanoDLR™ Dual Reporter Assay
The Nano-Glo® Dual-Luciferase® Reporter (NanoDLR™) Assay detects the activities of firefly and NanoLuc® luciferases in a single sample. Improved firefly chemistry and the bright, sensitive NanoLuc® luciferase make this a highly sensitive dual-luciferase assay that is simple to use, provides superior data quality, and allows greater flexibility in assay design.
The information in this article, outlining how to choose primary and control reporters for use with the NanoDLR Assay is available as a downloadable infographic.
References
- Cheng, K.C. and Inglese, J. (2012) A coincidence reporter-gene system for high-throughput screening. Nature Methods 9, 937.
- Ho, P. et al. (2013) Reporter enzyme inhibitor study to aid assembly of orthogonal reporter gene assays. ACS Chem. Biol. 8, 1009–17.
- Hasson, S.A. et al. (2015) Chemogenomic profiling of endogenous PARK2 expression using a genome-edited coincidence reporter. ACS Chem. Biol. 10(5), 1188-97.