T7 RiboMAX™ Express: Generation of 27kb In Vitro Transcripts in Minutes
Barbara Schelle and Volker Thiel
Institute of Virology and Immunology, University of Würzburg, 97078 Würzburg, Germany
Publication Date: 2002
Abstract
In vitro transcription using bacteriophage DNA-dependent RNA polymerases has been optimized to mediate the synthesis of large amounts of RNA. Here we show that in vitro transcripts of extraordinary size (27.3kb) can be synthesized in quantitative amounts using the T7 RiboMAX™ Express Large Scale RNA Production System (Cat.# P1320). Furthermore, full-length transcripts are produced in as little as 10 minutes. Best results with respect to integrity and yield of the RNA were obtained at incubation times of 30 minutes to 2 hours.
Introduction
In vitro transcription using bacteriophage DNA-dependent RNA polymerases has become an almost indispensable tool in molecular biological research. Enzymes like bacteriophage SP6, T3 and T7 RNA polymerases have been used to produce synthetic RNA for a wide variety of applications (1) (2) (3) (4) (5). In the field of RNA viruses, particularly for positive-stranded RNA viruses, the generation of full-length in vitro transcripts from cloned cDNA represents one of the most common strategies to produce synthetic RNA virus genomes (6). Reverse genetics systems have been established based upon the synthesis of full-length “infectious” RNA in vitro. However, for RNA viruses with extremely large genomes, the generation of synthetic genome-length RNA is still a challenging task. We have shown for coronaviruses, which have the largest genome of all RNA viruses (27–32kb), that infectious in vitro transcripts can be obtained using the T7 RiboMAX™ Large Scale RNA Production System (Cat.# P1300) (7). Here we report the synthesis of a 27.3kb RNA representing the human coronavirus (HCoV) genomic RNA in as little as 10 minutes using the T7 RiboMAX™ Express Large Scale RNA Production System (Cat.# P1320).
The DNA Template
We studied the ability of the T7 RiboMAX™ Express Large Scale RNA Production System to generate long RNA transcripts. We then used our recently described reverse genetics system for the generation of recombinant HCoV (7), based upon the synthesis of in vitro transcribed RNA from a full-length HCoV cDNA cloned in a vaccinia virus vector. The template for in vitro transcription is derived from genomic DNA of a recombinant vaccinia virus, vHCoV-inf-1. This virus contains the full-length HCoV cDNA of 27.3kb cloned downstream of a bacteriophage T7 RNA polymerase promoter and one additional G nucleotide to facilitate initiation of transcription. The HCoV cDNA is terminated by a synthetic polyA sequence of 42 nucleotides and a ClaI restriction site (Figure 1). Template DNA for in vitro transcription was prepared from purified vaccinia virus vHCoV-inf-1 stocks by proteinase K treatment, phenol extraction and ethanol precipitation. In order to produce run-off transcripts, genomic vaccinia virus vHCoV-inf-1 DNA was cleaved with ClaI, followed by phenol extraction and ethanol precipitation. The integrity of the ClaI-digested template DNA was analyzed by agarose gel electrophoresis (Figure 2).
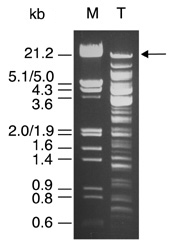
analyzed by agarose gel electrophoresis (lane T). The fragment containing the 27.3kb HCoV cDNA template is indicated. Lane M shows 500ng of EcoRI and HindIII-digested bacteriophage lambda DNA.
In Vitro Transcription
In vitro transcription reactions were performed according to the manufacturer's instructions in the T7 RiboMAX™ Express Large Scale RNA Production System Technical Bulletin (#TB298) in a 20µl reaction containing the following: 10µl of RiboMAX™ Express T7 2X Buffer, 6µl of nuclease-free water, 2µl of ClaI-digested template DNA (250ng/µl) and 2µl of Enzyme Mix. Note that the genomic DNA of vHCoV-inf-1 has a size of about 210kb compared to the HCoV-inf-1 cDNA insert of 27.3kb, which is the actual transcribed target sequence. Thus, only about 13% of the vHCoV-inf-1 genome represents the functional template DNA. In vitro transcription reactions were incubated at 37°C for time periods of 10 minutes to 2 hours.
After adding 1U of RQ1 RNase-free DNase to the reaction and incubating at 37°C for 15 minutes, the RNA was precipitated by adding 30µl of nuclease-free water and 25µl of LiCl solution (7.5M lithium chloride, 50µM EDTA). The reaction was chilled for at least 30 minutes at –20°C and centrifuged (maximum speed in a microcentrifuge at 4°C for 15 minutes). The resulting RNA pellet was washed with 70% ethanol, dried and resuspended in 1mM sodium citrate (pH 6.4).
Results
For each reaction we determined the total yield of in vitro transcripts and analyzed the RNA by agarose gel electrophoresis (Figures 3 and 4). Surprisingly, full-length transcripts became apparent after only 10 minutes incubation (Figure 4, lane 2). Under these conditions, the total yield was 8µg. Using longer incubation times the total yield of synthesized RNA increased up to 78µg. Full-length transcripts are easily detectable as the predominant RNA when reactions were incubated for 30 minutes, 1 hour and 2 hours (Figure 4, lanes 4–6). The RNA yields were 35µg, 50µg and 78µg, respectively. Overall, our data demonstrate the ability of the T7 RiboMAX™ Express Large Scale RNA Production System to mediate the synthesis of 27.3kb in vitro transcripts in quantitative amounts within short periods of time.
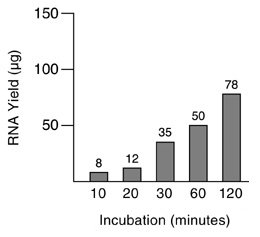
Figure 3. RNA quantification. The RNA yield of in vitro transcription reactions incubated for the indicated time periods were determined by ultraviolet light absorbance at a wavelength of 260nm. The numbers on top of each bar represent the RNA yield in micrograms resulting from in vitro transcription reactions with a volume of 20µl.
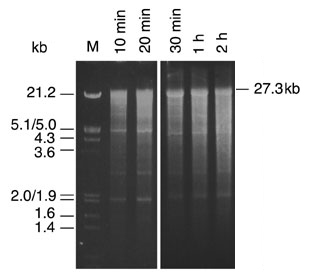
Figure 4. Analysis of in vitro transcripts by agarose gel electrophoresis. In vitro transcribed RNAs were visualized by agarose (1%) gel electrophoresis in 1X TBE/0.1% SDS buffer and subsequent ethidium bromide staining. Each lane contains 1µg of RNA resulting from in vitro transcription reactions incubated for the indicated time periods. Full-length RNA of 27.3kb is indicated. DNA fragment sizes of 500ng of EcoRI- and HindIII-digested bacteriophage lambda DNA (lane M) are shown.
References
- Chamberlin, M. et al. (1983) Isolation of bacterial and bacteriophage RNA polymerases and their use in synthesis of RNA in vitro. Methods Enzymol. 101, 540–68.
- Melton, D.A. et al. (1984) Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 12, 7035–56.
- Krieg, P.A. and Melton, D.A. (1984) Functional messenger RNAs are produced by SP6 in vitro transcription of cloned cDNAs. Nucleic Acids Res. 12, 7057–70.
- Nielsen, D.A. and Shapiro, D.J. (1986) Preparation of capped RNA transcripts using T7 RNA polymerase. Nucleic Acids Res. 14, 5936.
- Leary, S.L. et al. (1991) DNA-dependent RNA polymerase from bacteriophage T3 transcribes and amplifies an RNA template in vitro. Gene 106, 93–6.
- Bredenbeek, P.J. and Rice, C.M. (1992) Animal RNA virus expression systems. Sem. Virol. 3, 297.
- Thiel, V. et al. (2001) Infectious RNA transcribed in vitro from a cDNA copy of the human coronavirus genome cloned in vaccinia virus. J. Gen. Virol. 82, 1273–81.
Learn more about the T7 RiboMAX™ Express Large Scale RNA Production System