DNA Extraction from Fungi Environmental Field Samples
M. Rischer1,2, R. Neumann3 and S. Domey3
1Friedrich-Schiller-University, Institute for Microbiology, Jena
2Currently: Leibniz Institute for Natural Product Research and Infection Biology – Hans Knöll Institute, Jena
3Thüringer Landesanstalt für Landwirtschaft, Jena
Publication Date: February 2016
Abstract
In this article we compare the DNA extraction performance of the Maxwell® 16 Instrument to the InnuSpeed Bacteria/Fungi DNA Kit and the NucleoSpin® Soil Kit.
Introduction
Species of the genera Cladosporium, Acremonium, Verticillium, Aureobasidium, Alternaria, Epicoccum and Fusarium are typical representatives of the flora of cereals (1) (2). Particular attention is paid to the genus Fusarium, because many of the ubiquitous species cause damage and yield reduction in plants by fusariosis, and ingestion of the produced mycotoxins can have toxic effects. Due to the complexity and variability of morphological identification data for Fusarium species, molecular biological methods have become more important in diagnostics. Current efforts focus on the validation and standardization of protocols for DNA isolation, PCR and sequencing as well as data analysis (3) (4). In particular, the thick cell walls of plant pathogenic fungi can complicate cell lysis for DNA extraction (5).
In the present study we investigated DNA extraction with a new automatic DNA extraction instrument, the Maxwell® 16 Instrument (Promega). We used the Maxwell® Instrument to extract DNA from fungal material and compared the results with those from other commercially available extraction kits.
Sample Types and Extraction Methods
Fusarium fungi were extracted from freshly harvested grain from Thuringia. Nine samples of winter barley, two of spring barley and one winter wheat sample were used. In total four different species were chosen: F. avenaceum, F. graminearum, F. poae and F. tricinctum. To standardize molecular biology methods, three different DNA extraction methods were used. The Maxwell® 16 Instrument, and corresponding Maxwell® 16 FFS Kit (Promega) was compared to two commercially available extraction kits, the InnuSpeed Bacteria/Fungi DNA kit (Analytik Jena), and the NucleoSpin® Soil kit (Macherey Nagel).
The fungal species were incubated for eleven days on potato dextrose agar (PDA) media and approximately 18mg of mycelium were transferred to tubes with beads. The samples for the Maxwell® 16 Instrument and the NuceloSpin kit were mixed with 500μl cetyltrimethylammonium bromide buffer (CTAB) and shaken (2 × 3 minutes) using a vortexer. The InnuSpeed samples were shaken with 100ml distilled water. To each Maxwell® sample, 10μl RNase (10mg/ml), 10μl lysozyme (10mg/ml) and 20μl proteinase K were added (20mg/ml) and incubated for 90 minutes at 65°C with gentle shaking. The DNA was extracted according to the protocol for each kit.
Sample Quantitation and Amplification
The amount of extracted DNA extracted with each kit was determined using a spectrophotometer (NanoDrop® 1000). The DNA extracted using the Maxwell® 16 instrument were further used for performing the standard PCR protocol of the ‘Bund/Länder’ working group on Genetic Engineering (B/LAG, 2012) and for DNA sequencing.
Results and Discussion
For most samples tested, we found that the Maxwell® 16 extraction protocol provided higher yields of DNA (Figure 1). In addition, automated extraction with the Maxwell® 16 instrument saved the analyst time (90 minutes). During extraction there was sufficient time to perform other laboratory activities. This is not true for the two manual kits evaluated. Furthermore, the Maxwell® system sped up the extraction process because multiple samples were processed in one run without fluid losses.
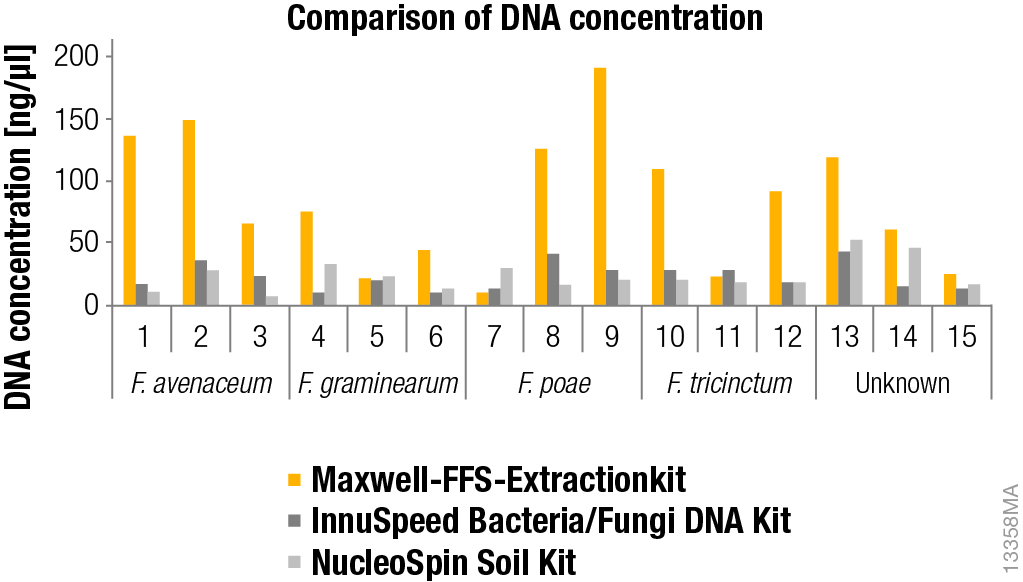
Figure 1. DNA concentration after extraction from mycelia from Fusaria fungi with different methods. DNA was extracted using the Maxwell® 16 Instrument and a custom kit, the NucleoSpin® Soil Kit or the InnuSpeed Bacteria/Fungi DNA Kit as described in the materials and methods section. The amount of DNA was determined using a spectrophotometer (NanoDrop® 1000 PEQLAB GmbH, Erlangen, Germany). Unknown fungi were later observed as Microdochium nivale and Fusarium langsethiae by sequencing.
Conclusion
We found that DNA could be successfully extracted from environmental fungi samples using the automated Maxwell® 16 system. The system yields a high amount of clean DNA. Following DNA purification, we were able to successfully use the DNA in a standard ITS PCR assay and for sequencing analysis.
References
- Sampson, R.A. et al. (2000) Introduction to food- and airborne fungi. CBS–Centralbureau voor Schimmelcultures, Utrecht.
- Neumann, R. (2012) Qualitätsuntersuchungen der Thüringer Getreide- und Rapsernte anhand von repräsentativen Ernteproben und langjährige Gesamtübersicht. Thüringer Landesanstalt für Landwirtschaft, Jena.
- BVL, Methodensammlung der Bund/Länder-AG Gentechnik (LAG) (2012) Molekularbiologische Identifizierung von Pilzen mittels ITS-PCR und nachfolgender Sequenzierung. Journal of Consumer Protection and Food Safety 7, 71–6.
- Gherbawy, Y. et al. (2008) Molecular identification of fungi, quantitative detection of fungi by molecular methods: a case study on Fusarium. Heidelberg: Springer Verlag.
- Gonzalez-Mendoza, D. et al. (2010) A rapid method for isolation of total DNA from pathogenic filamentous plant fungi. Gen. and Mol. Res 9, 162–6.
Learn more about the Maxwell® RSC Instrument