Doing Good Science: Authenticating Cell Line Identity
Jill Harley Dunham1 and Pam Guthmiller2
1Emory University, Department of Pharmacology, and 2Promega Corporation
Publication Date: 2012
Abstract
Much of biomedical research—medicine, genetics, drug discovery, vaccine development, reconstructive medicine, basic science, HIV testing/treatment, and cell biology—is performed with cultured cells obtained from major repositories or fellow researchers. An estimated 15–20% of the time, cells used in experiments have been misidentified or cross-contaminated with another cell line. Although major repositories now authenticate cell line identity, many are calling for all researchers to test and authenticate cell line identity using standard genotyping techniques like STR analysis. Now there is a consensus standard from an ATCC working group that details how to authenticate cell lines for research use.
Introduction
The ability to repeat previously published observations in scientific research is important for confirming new discoveries. If previous data are not repeatable, it is doubtful that anyone will continue to pursue that work. Much of biomedical research—medicine, genetics, drug discovery, vaccine development, reconstructive medicine, basic science, HIV testing/treatment, and cell biology—is done with cultured cells obtained from major repositories such as American Type Culture Collection (ATCC) or from fellow researchers. An estimated 15–20% of the time, cells used in experiments have been misidentified or cross-contaminated with another cell line (1) (2) (3). ATCC, along with the Coriell Institute for Medical Research, European Collection of Cell Cultures (ECACC), Deutsche Sammlung von Mikrorganismen and Zellkulturen (DSMZ), and the Japanese Collection of Research Resources, all have received cell line submissions that, upon authentication, were determined to have been misidentified by the depositor (4) (5). This poses a huge threat to the quality of publications and legitimacy of research findings produced from any of these cell cultures. For this reason, these repositories now authenticate cell line submissions and monitor cross-contamination. Recently, an ATCC working group has developed a consensus standard for identifying and authenticating human cell lines using STR profiling.
Historical Perspective on the Problem
The first human cancer cell line, HeLa, was developed in 1952, and for the next 15 years, additional human cell lines from different tissues continued to be developed (6). In 1968, researchers discovered that many cultured cells exhibited characteristics that did not match the characteristics of the original source. This was the first evidence that particular methods of culturing cells could produce unpredictable changes to the cells or misidentification. While the development of better sterile techniques helped decrease cross-contamination, few tests were available to determine which cells were already affected. Since then, however, standardized methods have been developed that are both quick and inexpensive to perform. While these improvements should have eliminated much cellular cross-contamination, it remains a prominent issue. ATCC and other similar repositories now monitor cross-contamination and authenticate all cell lines that they distribute, but most individual investigators do not adhere to the same meticulous authentication processes. In fact, a 2004 survey of approximately 500 biologists by Gertrude Buehring of the University of California—Berkeley, and her colleagues showed that less than half of all researchers regularly verify the identities of their cell lines using any standard techniques, such as DNA fingerprinting by short tandem repeat (STR) analysis (6). Without requiring that all cell lines be authenticated, misidentification will remain a significant problem.
Authentication Saves Time and Money
Aside from the issue of inconsistent or questionable data, cross-contamination also wastes time and money. For instance, Mordechai Liscovitch, a cancer researcher in Israel, says that he and researchers in his lab spent three years working on two breast cancer cell lines (MCF-7 and MCF-7/AdrR, now renamed NCI/ADR-RES) that they believed were related, only to discover later that the cell lines were actually unrelated. Although some researchers suspected the misidentification of these cell lines as early as 1998, the actual identity of these two cell lines was not confirmed until 2007 (5) (7). NCI/ADR-RES is not derived from the breast cancer cell line MCF-7 as originally thought but rather from the ovarian cancer cell line OVCAR-8 (7). All three of these cell lines are part of the NCI-60 panel of cell lines that are routinely used in drug-screening applications (5).
The Liscovitch lab cancelled the publication of a manuscript that contained erroneous conclusions based on the mistaken cell line identity; however, an unknown number of studies have been published that contain conclusions based on misidentified cell lines. Charles Patrick Reynolds of the University of Southern California and the Children’s Hospital Los Angeles’ Institute for Pediatric Clinical Research establishes new pediatric cancer cell lines and tests potential cancer drugs on existing lines. According to his estimations, up to 35–40% of previously published cell biology papers would need to be retracted due to invalid data. These estimates have caused Roland Nardone at The Catholic University of America to call for authentication both as a condition for receipt of grant funds from major agencies such as the National Institutes of Health and American Cancer Society as well as for publication of cell culture-based research in leading journals. He also is requesting education for technicians and scientists regarding prevention and detection of cross-contamination, including relevant professional societies’ endorsements of these proposed policies and sponsored conferences and workshops to facilitate adoption of the standards. Dr. Nardone’s belief that these changes are vital to scientific research has even led him to cocreate the Cell Line Authentication Global Awareness Month [May 2008], started by “an ad hoc group of scientists because of the chaos and waste caused by rampant misidentification and cross-contamination of cell lines.”
The problems of cross-contamination have been recognized already by a few organizations, such as the ATCC (7) and FDA, which requires that in-process materials, such as cell lines, that are used to produce pharmaceuticals be tested for identity and purity (8). Similarly, Nature recently mandated STR fingerprint data for papers reporting new human embryonic stem cell lines but not other lines (5). Although journals and funding agencies recognize the problems of cell line misidentification, they are uncertain how best to address them. When would researchers be asked to confirm cell line identity, before or after peer review? Where would the resources come from to confirm authors’ assertions of cell line identity?
In addition to wasted time and money, inconsistent or nonreproducible findings, and retraction of publications, a potential for health consequences also exists in misidentified cell lines. Drugs, vaccines and other biomedicines all are created based on findings in the lab, often times via cell culture. Products made using misleading or false data can cause major delays in the production and availability of treatments for a variety of diseases. The longer it takes for a treatment to be developed, the more people these diseases affect.
To address the issue of cell line authentication, the American Type Culture Collection (ATCC) Standards Development Organization (SDO) assembled a working group in January 2009. They were tasked with developing guidelines by which human cell lines will be authenticated and identified using short-tandem repeat (STR) profiling. A universal database that includes STR information, electropherograms, and other relevant information about a cell line would be created and maintained for researchers to identify and verify cell lines and assess cell line relationships.
The ATCC SDO presented a new consensus standard ASN-0002 “Authentication of Human Cell Lines: Standardization of STR Profiling” in 2010 and submitted to the American National Standards Institute regarding human cell line authentication using profiles generated from STRs. This standard was officially published in February 2012. Human cell line identification and authentication can affect scientists from grant submission to article publication. In fact, journals like In Vitro Cellular and Developmental Biology, International Journal of Cancer, Cell Biochemistry and Biophysics and more require cell line authentication prior to publication (9). Resources for checking if a human cell line is misidentified or cross-contaminated (e.g., Chang liver cells are actually HeLa cells) include the European Collection of Cell Cultures, ATCC and Wikipedia.
STR-Based Methods Provide Easy, Quick Cell ID
All of these misidentification issues can be prevented with inexpensive and now standard procedures used to authenticate cell lines, but the procedures must be performed. Roderick MacLeod and his colleagues at DSMZ, German Collection of Microorganisms and Cell Cultures, have found that about 90% of scientists ignore or refuse a cell bank’s request to send in new lines, preventing the establishment of cell line DNA fingerprints for future attempts at verification (7). Researchers need to be educated early on in their research to learn how to detect intra- and inter-species cross-contamination, as well as why it is so important to do so.
Many methods, such as isoenzyme analysis, karyotyping, human lymphocyte antigen (HLA) typing, and characterization of amplified fragment length polymorphisms (AFLP), have been used to identify cross-contamination in cell culture. However, a superior method is STR-profiling, well established in the field of DNA-based forensic identification. At the ATCC, STR analysis is performed using multiplex PCR (Promega PowerPlex® 1.2 System) to simultaneously amplify eight STR loci plus Amelogenin for gender determination (10) (11). A unique pattern of repeating DNA is generated for each human cell line analyzed, so the DNA profile of each new stock is verified by comparing to the baseline profile. STR profiling already has prevented the further distribution of six different cell lines at ATCC after it revealed the presence of Y chromosome-specific amplification products in cell lines that are derived from females. The research community hopes that STR profiling will provide a global reference technique to detect and eliminate cell line contamination.
Cell ID System Allows STR-Based Authentication of Your Cell Line
Promega is a leader in providing STR-profiling systems for forensic and paternity applications, and the PowerPlex® 1.2 STR analysis system has become the “gold standard” tool used by cell culture facilities to authenticate cell lines. To support the need for simpler methods to authenticate cell lines, the Cell ID™ System was developed and offers an improved system that includes the reagents required to successfully and simply identify and authenticate human cell lines as well as detect intra-species cell line cross-contamination.
The Cell ID™ System uses STR analysis of specific, highly polymorphic loci in the human genome through simultaneous amplification and three-color detection of ten loci (nine STR loci and Amelogenin for gender identification; including D21S11, THO1, TPOX, vWA, Amelogenin, CSF1PO, D16S539, D7S820, D13S317 and D5S818). These loci collectively provide a genetic profile with a random match probability of 1 in 2.92 × 109 (Figure 1). The system includes a hot-start Taq DNA polymerase for convenient room temperature reaction assembly. Following amplification, samples are analyzed by capillary electrophoresis in a single injection in conjunction with provided standards to assist in determining allele sizes for the different loci (Figure 2). The genetic profile is determined using allele-calling software. Most research scientists have access to institute core facilities and service companies that have the instrumentation (capillary electrophoresis), software and experience to perform cell line profiling, even if they do not have the same capability within their own labs. General recommendations in the literature suggest that investigators authenticate an early passage (first week of culture) of their cells to establish the identity of the cell line. Cells should be authenticated again before freezing, once every two months that the culture is actively growing, and before publication. If a lab is using more than one cell line, all lines should be tested initially to rule out cross-contamination (8).
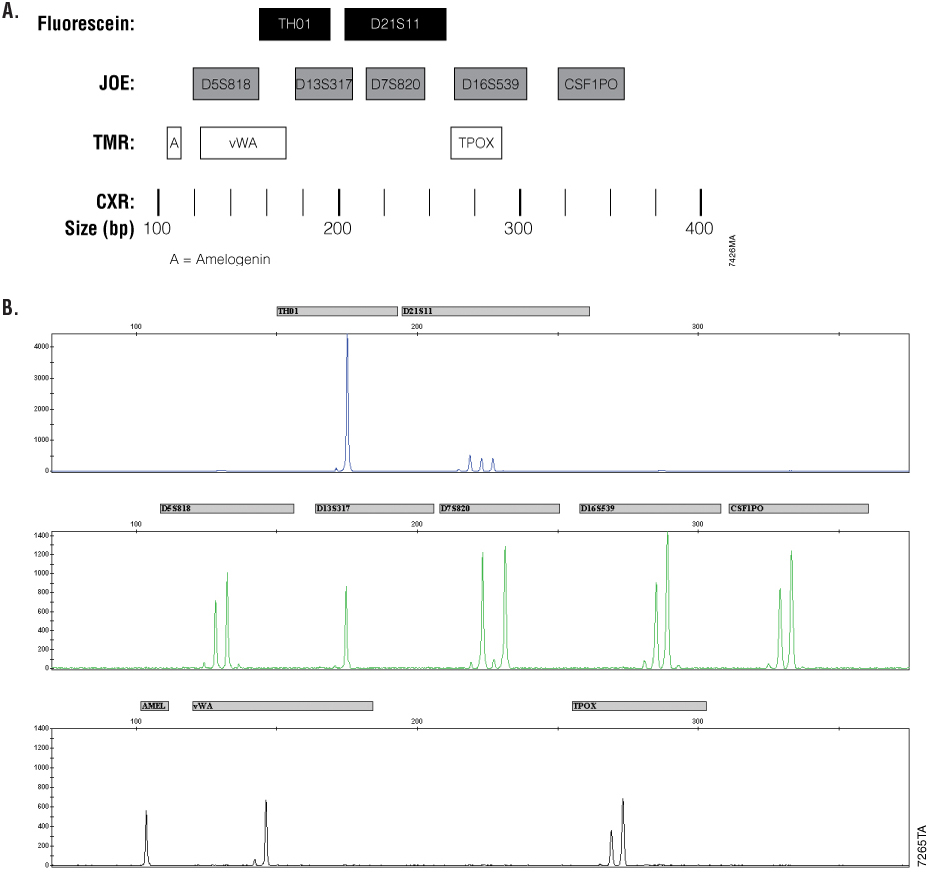
Figure 1. Panel A. Allele ranges for the Cell ID™ System. STR fragments amplified by the Cell ID™ System are labeled with different dyes and are separated by capillary electrophoresis based on size. A size standard is included in each sample to determine the size of the fragments. JOE-labeled loci are shown in gray. Fluorescein-labeled loci are shown in white. The CXR-labeled Internal Lane Standard 600 fragments are represented by black bars. Panel B. K562 cell line DNA profile. After amplification with the Cell ID™ System and subsequent detection on a capillary electrophoresis instrument, the resulting sample data are displayed as a series of dye-labeled allele peaks.
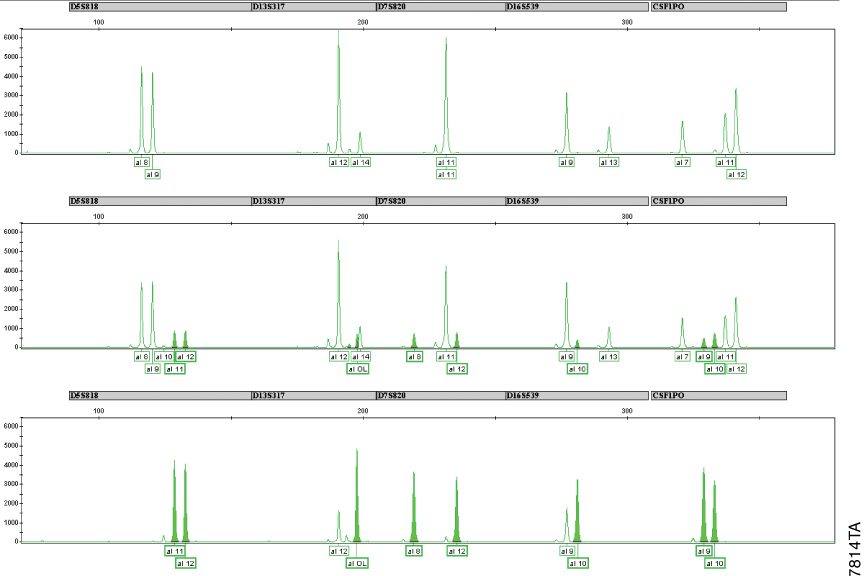
Figure 2. Determination of the cell line contamination using the Cell ID™ System.
Panel A. HEK293 cell line STR profile. Panel B. STR profile of HEK293 Cell Line with 29% HeLa cell line contamination. Panel C. HeLa cell line STR profile. DNA was extracted from 104 cells using the Maxwell® 16 Cell LEV DNA Purification Kit then amplified with the Cell ID™ System. Amplified products were detected on a capillary electrophoresis instrument. For simplicity, only the JOE-labeled allele profiles are shown.
Summary
Because of the importance of cell culture to biomedical research and technology, proper cell line authentication is in everyone’s best interest. However, cross-contamination continues to be a problem. With the increasing number of new cell lines and the high rate of cell culture use in labs worldwide, significant gaps have been created in basic principles of quality control (i.e., cell line authentication). From research articles published with misidentified cell lines and resultant questionable results, to stem cell lines and other lines destined for clinical uses, cross-contamination affects science in all realms—from lab bench to clinic. Without significant change to the handling and treatment of cell cultures, it will become only a larger and more serious issue.
References
- Cabrera, C.M. et al. (2006) Identity tests: Determination of cell line cross-contamination. Cytotechnology 51, 45–50.
- Drexler, H.G., MacLeod, R.A. and Dirks, W.G. (2001) Cross-contamination: HS-Sultan is not a myeloma but a Burkitt lymphoma cell line. Blood 98, 3495–6.
- Drexler, H.G., Dirks, W.G. and MacLeod, R.A. (1999) False human hematopoietic cell lines: Cross-contaminations and misinterpretations. Leukemia 13, 1601–7.
- MacLeod, R.A. et al. (1999) Widespread intraspecies cross-contamination of human tumor cell lines arising at source. Int. J. Cancer 12, 555–63.
- Chatterjee, R. (2007) Cell biology. Cases of mistaken identity. Science 315, 928–31.
- Buehring, G.C., Eby, E.A. and Eby, M.J. (2004) Cell line cross-contamination: How aware are Mammalian cell culturists of the problem and how to monitor it? In Vitro Cell. Dev. Biol. Anim. 40, 211–5.
- Liscovitch, M. and Ravid, D. (2007) A case study in misidentification of cancer cell lines: MCF-7/AdrR cells (re-designated NCI/ADR-RES) are derived from OVCAR-8 human ovarian carcinoma cells. Cancer Lett. 245, 350–2.
- FDA. General Requirements for Laboratory Controls. 21 CFR 211.160 and 21 CFR 610.18.
- American Type Culture Collection Standards Development Organization Workgroup ASN-0002. (2010) Cell line misidentification: The beginning of the end. Nat. Rev. Cancer 10, 441–8.
- ATCC Connection Newsletter (2000) 21, 1–2.
- Masters, J.R. et al. (2001) Short tandem repeat profiling provides an international reference standard for human cell lines. Proc. Natl. Acad. Sci. U.S.A. 98, 8012–7.
Learn more about cell line authentication.