Alternate Protocol for Maximizing Transfection Efficiency Using ViaFect™ Reagent
Leta Steffen and Amy Landreman
Promega Corporation
Publication Date: January 2017
Abstract
This article describes an alternate protocol for creating transfection mixtures using ViaFect™ Transfection Reagent in which the reagent:DNA complexes are formed in a lower volume compared to the standard protocol. Two reporters were used to monitor transfection efficiency, luciferase and GFP, and the health of the cells was also measured for each transfection condition. This method offers improved transfection efficiency in some cell lines, including HEK293 cells, and enables reduction in the amount of reagent required for others, while still maintaining the health of the cells during the process.
Introduction
When introduction of DNA into cells is needed, a reproducible, efficient transfection process with minimal impact on cell health is required for optimal results. Maximizing the efficiency of this process can help reduce the amount of DNA and reagent consumed, further improve data and enable assay expansion into additional cell models. Lipid-based transfection is a commonly used method of introducing DNA into the cell. In this method, the reagent associates with the DNA to create a reagent:DNA complex which can then pass through the cell membrane delivering the DNA to your target cell. The method by which these complexes are formed is a critical part of any lipid-based transfection protocol. This article describes an alternate protocol for creating transfection mixtures using ViaFect™ Transfection Reagent in which the reagent:DNA complexes are formed in a lower volume compared to the standard protocol. Two reporters, luciferase and GFP, were used to monitor transfection efficiency. Cell health was also measured for each transfection condition. This method offers improved transfection efficiency in some cell lines, including HEK293 cells, and enables reduction in the amount of reagent required for others, while maintaining cell health during the process.
Materials
Cell Lines and Culture Media
• HEK293 (DMEM + 10% FBS)
• C2C12 (DMEM + 10% FBS)
• A549 (Ham’s F12 + 10% FBS)
• K562 (RPMI 1640 + 10% FBS)
Transfection Reagents
• Opti-MEM™ or other serum-free media (Gibco Cat.# 31985)
• ViaFect™ Transfection Reagent (Cat.# E4981)
• pGL4.13[luc2/SV40] Vector (Cat.# E6681)
• pCAG-GFP plasmid (Addgene Cat.# 11150)
Analysis Reagents
• ONE-Glo™ Luciferase Assay System (Cat.# E6120)
• CellTiter-Fluor™ Cell Viability Assay System (Cat.# G6080)
Instruments
• GloMax® Discover System (Cat.# GM3000)
Alternate Protocol
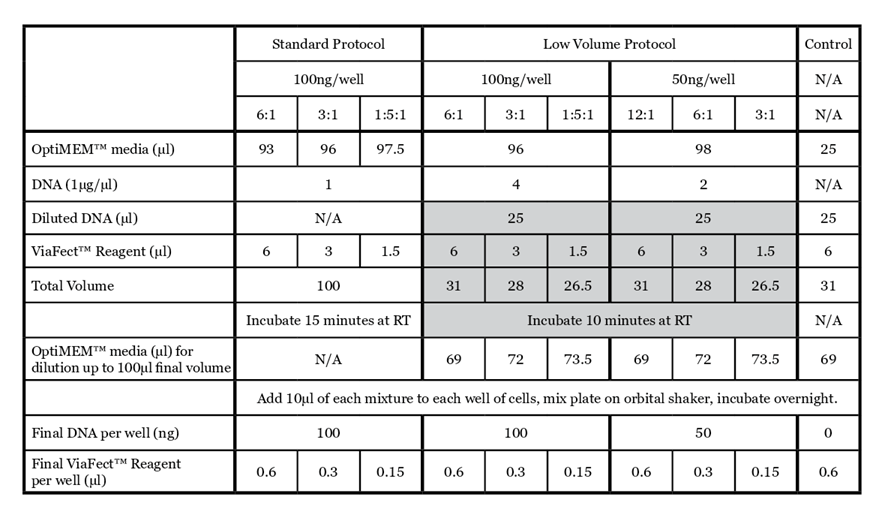
For details on the standard ViaFect™ Transfection Reagent protocol, refer to the ViaFect™ Transfection Reagent Technical Manual #TM409. The protocol below outlines the details of this study; some steps may be optimized for your cell culture system. The alternate protocol is summarized in Table 1 for comparison.
Day 1 – Cell Plating
Plate cells at 1 × 104 cells in 100µl and incubate overnight at 37°C with 5% CO2 so that they will be approximately 75% confluent on the day of transfection. Due to differences in growth rates, confluence is variable between cell lines. Plate K562 cells at 1 × 104 in 100µl at the time of use (no overnight incubation needed).
Day 2 – Transfections
- Allow transfection reagents to warm to room temperature before use. Mix by vortexing.
- Prepare transfection complexes with a given plasmid according to Table 1.
- a. Dilute DNA by adding OptiMEM™ media and DNA to a microfuge tube. Vortex to mix.
Note: Adjust total volume of diluted DNA for the appropriate number of wells being transfected. - b. To a microfuge tube or a V-bottom plate, add 10µl of diluted DNA.
- c. Add ViaFect™ Transfection Reagent and mix by pipetting.
- d. Incubate for 10 minutes at room temperature.
- e. Dilute complexes with OptiMEM™ media to reach a final volume of 100µl and pipette gently to mix.
- a. Dilute DNA by adding OptiMEM™ media and DNA to a microfuge tube. Vortex to mix.
- Add 10µl of transfection complex or control treatment to each well of cells in culture. Additional controls include no ViaFect™ Transfection Reagent/no DNA treatment and a no-cells background control for viability measurements.
- Mix plate on an orbital plate shaker for 10–30 seconds.
- Incubate cells overnight up to 28 hours at 37°C with 5% CO2.
Day 3 – Measurements & Data Analysis
-
Measure cell viability of luciferase transfected cells using CellTiter-Fluor™ Cell Viability Assay. See the CellTiter-Fluor™ Cell Viability Assay Technical Bulletin #TB371 for a multiplex assay protocol.
- a. For background correction, subtract the average fluorescence of the no-cells control wells (with reagent) from all other fluorescence measurements.
- b. For percent viability, divide the background corrected values by the average fluorescence for the untreated cell controls.
-
Measure luminescence of luciferase transfected cells using ONE-Glo™ Luciferase Assay System. See the ONE-Glo™ Luciferase Assay System Technical Manual #TM292 for protocol.
- a. For background correction, subtract the average luminescence of the no-cell control wells (with reagent) from all other luminescence measurements.
- Measure fluorescence of GFP-transfected cells using the GloMax® Discover instrument (Ex 475nm, Em 500–550nm).
- Subtract the average of the non-transfected control wells from the GFP measurements.
Results and Conclusions
Compared to the standard ViaFect™ transfection protocol, low-volume complex formation improved the transfection efficiency for some cell lines tested and yielded equivalent results for others. For both luciferase and GFP reporters, HEK293 cells showed markedly improved efficiency when the alternate protocol was used for complex formation (Figure 1). When the same amount of ViaFect™ Transfection Reagent and DNA were used, the alternate protocol significantly increased reporter expression. The improved efficiency of the alternate protocol also enabled equivalent results when either less DNA or ViaFect™ Transfection Reagent was used.
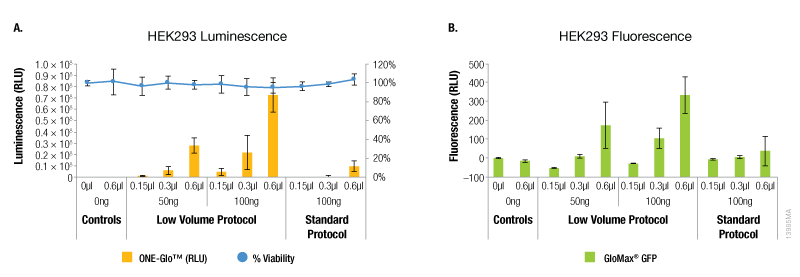
Figure 1. Transfection efficiency for HEK293 cell line. ViaFect™ (μl) and plasmid DNA (ng) used per well are indicated on the x-axes. Panel A. Luminescence and viability after transfection of HEK293 with pGL4.13[luc2/SV40] DNA. Panel B. Fluorescence after transfection of HEK293 with pCAG-GFP DNA.
In contrast, A549 cells showed equivalent levels of reporter expression when the alternate protocol was used (Figure 2). This can be achieved with the alternate protocol by using either less ViaFect™ Transfection Reagent or DNA, providing an option to decrease the amount of reagents consumed. Similarly, results in C2C12 cells demonstrated modest improvements with the low volume protocol. These results were also observed when less ViaFect™ Transfection Reagent was used for the transfection (Figure 3).
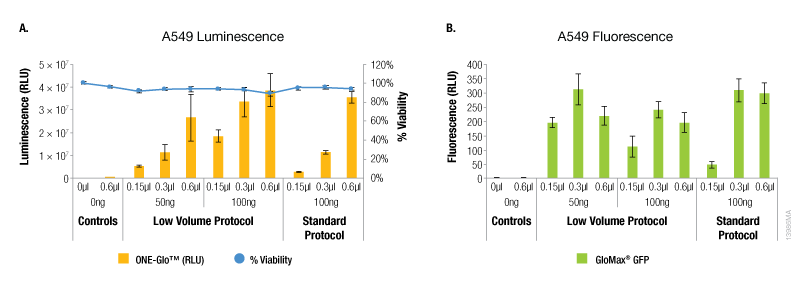
Figure 2. Transfection efficiency for A549 cell line. For this cell line, the transfections indicated in Panels A and B were performed on different days. Panel A. Luminescence and viability after transfection of A549 cell line with pGL4.13[luc2/SV40] DNA. Panel B. Fluorescence after transfection of A549 cell line with pCAG-GFP DNA.
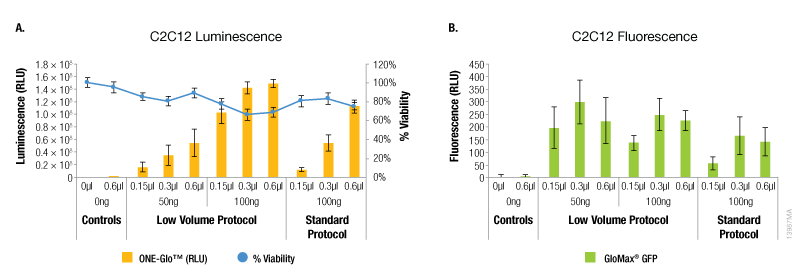
Figure 3. Transfection efficiency for C2C12 cell line. Panel A. Luminescence and viability after transfection of C2C12 cell line with pGL4.13[luc2/SV40] DNA. Panel B. Fluorescence after transfection of C2C12 cell line with pCAG-GFP DNA.
The results with the K562 suspension cells showed a different trend compared to the adherent cell lines tested. For these cells, the maximum expression of both reporters was observed with the lowest amounts of ViaFect™ Transfection Reagent using either protocol (Figure 4). However, the low volume protocol did deliver the highest levels of reporter expression and this was observed with the lowest amounts of DNA and ViaFect™ Transfection Reagent. Minimal impact on cell viability was observed across all experimental conditions using both transfection protocols.
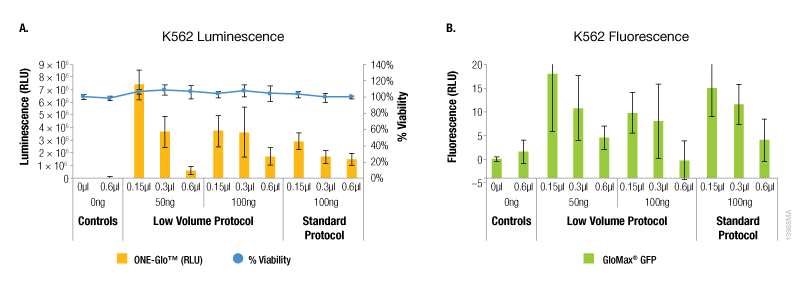
Figure 4. Transfection efficiency for K562 cell line. Panel A. Luminescence and viability after transfection of K562 non-adherent cell line with pGL4.13[luc2/SV40] DNA. Panel B. Fluorescence after transfection of K562 non-adherent cell line with pCAG-GFP DNA.
Generating the ViaFect™ Transfection Reagent:DNA complexes in a lower volume provides an optional protocol for those users looking to optimize their transfection protocols for maximum efficiency. For some cell lines, a substantial increase in transfection may be achieved by using the new protocol with the same amount of DNA and ViaFect™ Transfection Reagent specified in the old protocol. For other cell lines, the new protocol may provide an option to reduce the amount of ViaFect™ Transfection Reagent or DNA used while maintaining similar transfection efficiency.
Learn more about the ViaFect™ Transfection Reagent