RNA-Free Plant DNA Using the Maxwell® RSC PureFood GMO and Authentication Kit
László Kupcsik1, Chris Moreland2, and Carlos Rossini3
1Laboratory of Plant Physiology and Molecular Genetics, Université libre de Bruxelles, Campus Plaine CP 242, Bd du Triomphe, 1050 Brussels, Belgium, 2Promega Corporation, 3Promega Benelux, Netherlands
Publication Date: January 2017
Abstract
Here we describe a modification of the Maxwell® RSC PureFood GMO and Authentication Kit Protocol that allows the extraction of RNA-free DNA from Arabidopsis suitable for downstream applications such as next-generation sequencing.
Introduction
Recent years have seen the rapid expansion of large-scale genomics and transcriptomics screening with the advent of next-generation DNA sequencing (NGS) methods. This experimental approach necessitates the extraction of DNA from several specimens in parallel. Automated DNA extraction systems, such as the Maxwell® automated platforms, greatly reduce sample processing overhead and allow researchers to focus on research questions rather than nucleic acid extraction.
Library preparations for sequencing are often carried out by external or core laboratories with strict sample submission guidelines, because the purity of the starting genomic DNA (gDNA) for NGS is important to achieve meaningful data. These sample quality guidelines often define that the gDNA must be of sufficient quantity and free of RNA, proteins, and phenol contamination that can impair downstream reactions and therefore the quality of sequencing results. For example, the ratio of OD260/280 is used as an indicator of sample purity, where values of approximately 1.8 are an indication of a good gDNA sample. However, this measurement can be compromised by the presence of RNA or small nucleic acid fragments such as nucleotides. Thus, care must be taken when choosing a gDNA extraction method so that samples are free of any contaminants.
This paper describes a modification of the Maxwell® RSC PureFood GMO and Authentication Kit protocol for the extraction of RNA-free DNA from plant material. It improves the DNA yield compared to the Plant DNA Kit and eliminates the substantial RNA contamination present in the eluates of the PureFood Kit.
Materials and Methods
The gDNA extraction experiments were performed at the Laboratory of Plant Physiology and Molecular Biology at the Université Libre de Bruxelles. Their research is focused on genome-wide association studies (GWAS) of Arabidopsis thaliana and requires high-quality gDNA. For all isolations, a Maxwell® 16 Research Instrument (Cat.# AS2000) and the FFS custom kit (X9431, currently sold as the Maxwell® RSC PureFood GMO and Authentication Kit, Cat.# AS1600) were used.
Arabidopsis leaves were flash-frozen in liquid nitrogen, and ground into a homogeneous, fine powder using a mortar and pestle. This powder was used in DNA extraction reactions directly or after storage at –80°C for several weeks.
The samples were processed according to the method published by the Bayerisches Landesamt für Gesundheit und Lebensmittelsicherheit (Bavaria State Office for Health and Food Safety) for the isolation of gDNA from feed and seed samples(1). The sample preparation and enzymatic pretreatment was modified in these experiments, as shown in Table 1.
| Table 1. Sample Preparation and Enzymatic Pretreatment Modifications. | |||
|---|---|---|---|
| Maxwell® 16 Application Protocol E35 |
E35 Application Protocol Scaled Up |
RNA-Free Protocol | |
| Starting Material | 200mg/ml CTAB | 400mg/ml CTAB | 200mg/ml CTAB (up to 500mg/ml possible) |
| Sample Pretreatment | 65°C for 90 minutes with Proteinase K and RNase A | 65°C for 90 minutes with Proteinase K and RNase A |
1. 65°C for 60 minutes with Proteinase K 2. 95°C for 15 minutes 3. 10 minutes at room temperature with RNase A |
| Extraction |
1. Centrifuge samples 10 minutes at 16,000 × g. 2. Add 300µl of lysis buffer to well 1 of the Maxwell® cartridge. 3. Proceed with the E35 protocol. |
||
| In all cases, 40µl of Proteinase-K (20mg/ml) and 20µl RNase A (4mg/ml) were added to 1 ml of CTAB solution. | |||
Results
At the time of the experiments, the recommended DNA extraction protocol for the FFS Kit (X9431) was the Maxwell 16® Application Protocol E35.(1) This protocol results in a substantial amount of RNA in the isolates, as shown in Figure 1.
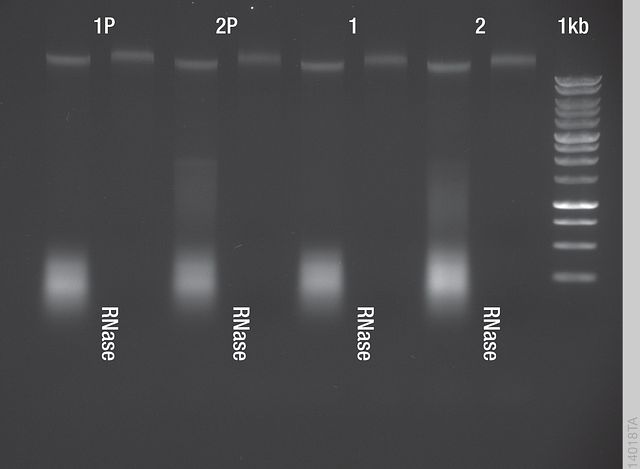
Figure 1. PureFood Kit Eluates. Four isolates (1P, 1P, 1, 2 [left lanes]) were run on a 1% agarose gel with 0.5 × TAE buffer next to a Promega 1kb DNA ladder. Half of the samples were RNase-treated after the elution (right lanes). Arabidopsis genomic DNA runs slightly above 10kb. The smear observed between 100–4,000bp is RNA.
Three protocols were tested to improve DNA purity: the original one, a modification using more starting material (Upscaled), and a sequential enzymatic treatment (RNA-free; Figure 2). The upscaled protocol yielded more DNA than the original method, but the RNA contamination also increased proportionally. The sequential enzymatic pretreatment almost completely eliminated the RNA from the eluates. During routine application of the RNA-free protocol, we found that up to 500mg leaf tissue can be used, which improves DNA yield without considerable RNA contamination.
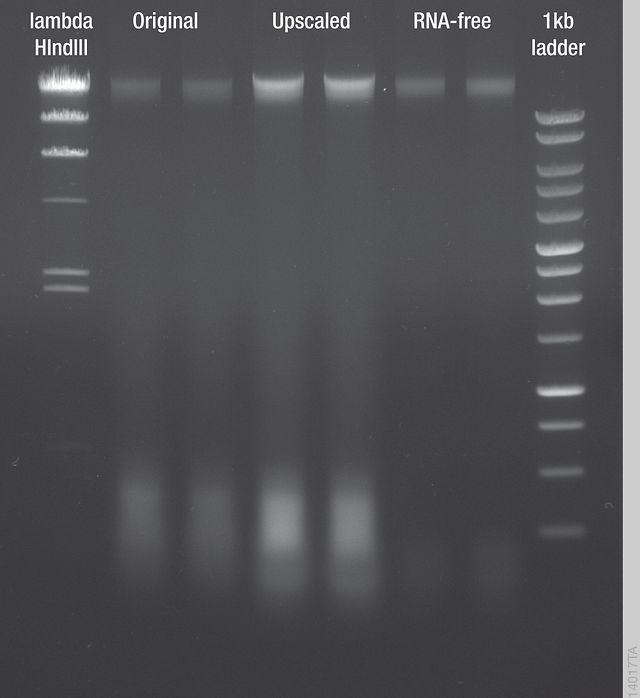
Figure 2. Comparison of DNA Extraction Methods. DNA samples extracted from leaves of two Arabidopsis thaliana plants using the three different protocols described in Table 1.
Conclusions
The Maxwell PureFood kit was designed to isolate nucleic acids from hard-to-process materials, such as seeds and processed food products. In principle, it can also be used to isolate DNA from any plant tissue. The kit protocol includes a Proteinase K treatment to inactivate DNA-modifying enzymes and an optional RNase treatment to remove RNA. We found that the proposed protocol is inefficient in eliminating RNA from the isolates, probably because the Proteinase K rapidly inactivates the RNase.
These results show that the optimal procedure begins with a Proteinase K digestion(2), followed by heat-inactivation of the enzyme, which allows for an effective RNase treatment thereafter. This improved protocol both protects the DNA and effectively eliminates RNA from the sample.
The PureFood kit may also be preferred over the standard Plant DNA kit because it allows for processing more starting material, which significantly increases the DNA yield per reaction. All in all, the presented protocol is ideal for the automated extraction of high-quality plant DNA suitable for NGS sequencing.
References
- Guertler, P. et al. (2013) Development of a CTAB buffer-based automated gDNA extraction method for the surveillance of GMO in seed. European Food Research and Technology 236, 599–606.
- Wiegers, U. and Hilz, H. (1971) A new method using proteinase K to prevent mRNA degradation during isolation from HeLa cells. Biochem. Biophys. Res. Comm. 44, 513–19.
Learn more about the Maxwell® RSC PureFood GMO and Authentication Kit