10 Things You May Not Know About OSIRIS
National Center for Biotechnology Information, National Library of Medicine and National Institutes of Health, Bethesda, Maryland, USA.
Publication Date: 2016

Introduction
The Open Source Independent Review and Analysis System (OSIRIS) was developed in response to recommendations of the Kinship and DNA Advisory Panel (KADAP) brought together by the US Department of Justice to develop guidelines and recommendations during the World Trade Center victim identification effort. OSIRIS was tested in the Hurricane Katrina victim identification, and further developed in collaboration with State and Federal laboratories. OSIRIS is fast, powerful short tandem repeat (STR) analysis software designed for forensic and clinical use and is approved by the National DNA Index System (NDIS) for use as an expert system.
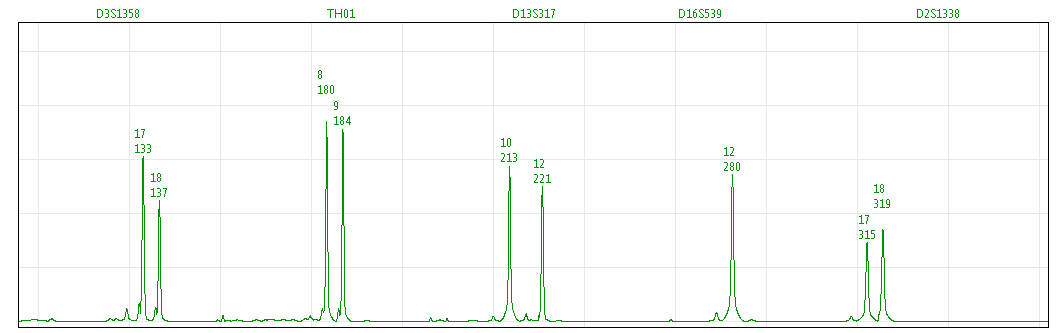 Figure 1. A NIST Sample (MIX05 Case 3 victim) amplified with Identifiler® and analyzed with OSIRIS software.
Figure 1. A NIST Sample (MIX05 Case 3 victim) amplified with Identifiler® and analyzed with OSIRIS software. 1. OSIRIS is powerful, open source software for STR analysis.
OSIRIS is grounded in fundamental mathematical and physical principles. It uses sophisticated algorithms to match mathematically defined curves to peaks and the baseline in the raw data produced by genetic analyzers (1) . This allows OSIRIS to calculate unique quality metrics such as peak fit (shape), displacement (shifting), residual displacement (displaced shifting) and sample-to-ladder fit for alleles, artifacts, noise and the baseline that allow improved allele calling and artifact identification. This makes it possible for OSIRIS to accurately identify low-level peaks and artifacts at very low analytical calling thresholds, which reduces the amount of editing that analysts must do, and the information provided to the analyst aids in making editing decisions.
2. OSIRIS can speed up STR analysis and increase reproducibility.
OSIRIS can increase the efficiency and accuracy of STR profile analysis. A plate of 96 samples can process in under 30 seconds. OSIRIS can analyze multiple runs at a time. A lab can analyze 100 plates of samples in a single OSIRIS batch in under an hour.
OSRIS identifies over 100 artifacts, which reduces the amount of manual editing. One lab’s evaluation found a tenfold reduction of editing in an analysis of heavily loaded reference samples as compared with GeneMapper® ID-X (Personal communication, Chris Hefford, Forensic Science SA, Australia). OSIRIS displays artifact information to the analyst, which can aid in complex calls and in training new analysts. Automated and accurate artifact identification increases analyst-to-analyst reproducibility, which is one of the current challenges in mixture analysis.
OSIRIS checks the accuracy of controls—including ladders, negatives, blanks, positives and even lab-custom positive controls—and also for missing positives and negatives. OSIRIS is Y- and X-STR aware and will accurately call missing loci in marker sets that include both Amelogenin and sex chromosome-specific markers.
OSIRIS predicts what sort of reanalysis is required when a sample fails based on software algorithms and user settings. This information can be exported to a LIMS system, and in a high-throughput system such as CODIS (Combined DNA Index System) laboratory, it can be used to automatically re-queue samples for reanalysis.
3. OSIRIS is used globally.
OSIRIS is used not only in the United States but also around the globe. Users include laboratories that perform forensic analysis, CODIS indexing, cell line authentication, paternity testing and other identity testing. OSIRIS supports the health-care mission of the National Institutes of Health (NIH), with users in clinical testing and research, including the NIH Clinical Center Molecular Hematology Laboratory, the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) and the National Institute of Allergy and Infectious Diseases (NIAID).
4. OSIRIS works with most marker systems.
OSIRIS comes with all of the common commercial marker sets defined. These include: Promega PowerPlex® Fusion, Fusion 6C, Y23, ESI 16/17 and ESX 16/17 Systems; Applied Biosystems GlobalFiler®, Yfiler® Plus PCR Amplification Kits, NGM™ and NGM SElect kits; and the Qiagen Investigator® 24plex QS kit. OSIRIS also includes legacy marker set definitions to allow reanalysis of legacy STR data. OSIRIS has been used with custom marker sets including ones for cannabis (2) and Fragile X syndrome.
5. OSIRIS is compatible with mixture analysis software.
OSIRIS data can be exported in a wide variety of formats for import into continuous and semi-continuous mixture analysis software and LIMS systems. OSIRIS uses an advanced mathematical algorithm for baselining and noise analysis. This gives very accurate peak heights and peaks areas, which increases the accuracy of continuous method mixture analysis and is critical in calculating accurate donor/recipient proportions in stem cell engraftment (chimerism) testing.
6. OSIRIS has the flexibility to be run on most computers.
OSIRIS can analyze both FSA and HID format files produced by the commonly used Applied Biosystems 310, 3100 series, 3500 and 3700 series Genetic Analyzers and data produced by IntegenX and NetBio Rapid DNA analyzers. This provides users with flexibility regarding their analysis platform and the ability to review Rapid DNA profiles.
OSIRIS is designed to be used with a wide range of hardware and software. OSIRIS can be used on Windows and Macintosh computers and will work on Windows XP, 7, 8 and 10, and Macintosh 10.9, 10.10, and 10.11 operating systems. This avoids the need for a new version of analysis software when the lab transitions to a new operating system.
OSIRIS has minimal hardware requirements. The program will occupy about 55Mb on the hard drive. The program does not require any additional RAM beyond what is necessary for the computer’s operating system to run efficiently. OSIRIS can be set up without administrator privileges for evaluation purposes.
7. OSIRIS is network-ready.
OSIRIS can be installed and run on a network, allowing an unlimited number of simultaneous users access to the laboratory’s analysis parameters and analysis data stored on the network. This means that maintenance of analysis settings is easily accomplished in a single location, the analysis parameters can be protected against unauthorized changes, and users can analyze and save data to the network. LIMS systems can take advantage of this to manage data for analysis and storage. Because of the way OSIRIS is designed, multiple users do not impact performance; OSIRIS runs as fast with 100 simultaneous users on the network as it does with a single user.
8. OSIRIS catches problems before they become critical.
OSIRIS can be used for process quality control. It has an automated reporting function that can be used to export processed data without analyst intervention. Information regarding matrix, capillary array, injection and control sample quality can be automatically exported for tracking by the lab. This allows problems—such as an invalid control—to be caught before they become critical and adversely affect the data produced by the laboratory. An analyst or a computerized laboratory process could run OSIRIS on multiple analyzer batches with an automated OSIRIS export that generates a report indicating whether the positive controls, the negative controls and the ladders have passed or failed QC, as determined by OSIRIS algorithms and user-defined settings. While automated control sample checking would be particularly advantageous to a high-throughput laboratory, knowing when to replace a capillary array or remake a color separation matrix could save headaches for any laboratory.
9. OSIRIS works seamlessly in software workflows.
OSIRIS can be closely integrated with other software. It can be run from the command line, which means that it can be called from within other software packages or by a laboratory LIMS. This makes it possible to integrate OSIRIS with other software in the laboratory or for OSIRIS to be run automatically as a part of a laboratory analysis pipeline workflow.
10. OSIRIS was developed by the community, for the community.
OSIRIS is an open source project of the National Center for Biotechnology Information (NCBI) in collaboration with NIST, the NIH Clinical Center, the Defense Forensic Science Center (DFSC, previously USACIL), the Illinois State Police Indexing Laboratory and OSIRIS users. The OSIRIS source code is available on the GitHub NCBI page, allowing the STR community to research, reuse, repurpose and improve the software. Additional collaborators are welcome. The improvements that have been and continue to be made are in response to feedback and requests for features from forensic, clinical and research users. OSIRIS is also updated if required by changes in operating systems.
Summary
OSIRIS is an advanced and powerful STR profile analysis program. It is designed to accurately call alleles and identify artifacts in a variety of forensic and clinical casework to increase analyst efficiency, accuracy and reproducibility. OSIRIS is a free download available on the OSIRIS homepage. Also available are the User Guide, a Webinar and conference presentations. Feedback and suggestions are welcomed at the “forensics” e-mail on the OSIRIS home page.
Funding/Acknowledgements
This research was supported by the Intramural Research Program of the NIH, National Library of Medicine.
Article References
- Goor, R. et al. (2011) A Mathematical Approach to the Analysis of Multiplex DNA Profiles. Bulletin of Mathematical Biology 73, 1909–1931.
- Clarke, R. and Coyle, H. (2016) Re-evaluation of the Current NMI01 STR Sizing System of Cannabis DNA Henry C. Lee Graduate Student Showcase, University of New Haven
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Riley, G., Goor, R., Hoffman, D. and Sherry, S. 10 Things You May Not Know About OSIRIS. [Internet] 2016. [cited: year, month, date]. Available from: https://www.promega.com/resources/profiles-in-dna/2016/10-things-you-may-not-know-about-osiris/
American Medical Association, Manual of Style, 10th edition, 2007
Riley, G., Goor, R., Hoffman, D. and Sherry, S. 10 Things You May Not Know About OSIRIS. Promega Corporation Web site. https://www.promega.com/resources/profiles-in-dna/2016/10-things-you-may-not-know-about-osiris/ Updated 2016. Accessed Month Day, Year.
PowerPlex is a registered trademark of Promega Corporation. Identifiler, GeneMapper, Globalflier and Yfiler are registered trademarks of Thermo Fisher Scientific, and NGM is a trademark of Thermo Fisher Scientific. Investigator is a registered trademark of Qiagen.