Phenotypic Consequence of Target Degradation
Targeted protein degradation is a promising approach for developing new drugs, especially for diseases caused by proteins that are difficult to inhibit using traditional small molecule inhibitors. By understanding the phenotypic consequences of degrading a specific protein, researchers can assess the potential therapeutic benefits and risks of using protein degradation as a treatment strategy.
Here we describe a fast and efficient way to characterize protein degradation phenotype using HaloPROTAC3.
What Is the Phenotypic Consequence of Target Degradation?
Temporal degradation of proteins inside a cell often elicits a much different phenotype than genetic knockouts or protein mutations. HaloPROTAC3, a fusion of a HaloTag® label and a PROTAC, is a rapid and highly effective way to understand and characterize protein degradation phenotype.
HaloPROTAC3 recruits an endogenous VHL E3 ligase component to a HaloTag® fusion protein, resulting in ubiquitination and degradation via the proteasome pathway. HaloPROTAC3 contains a degradation-inducing acylamine moiety, coupled to a chloroalkane moiety by a linker of variable length.
To study endogenous protein loss in relevant cell backgrounds, we recommend incorporating a tag into the target protein loci via CRISPR/Cas9 gene editing. Loss of protein is monitored in cells treated with HaloPROTAC3 using live-cell luminescence. HaloPROTAC3 shows fast burst loss that is sustained over time with endogenously tagged HaloTag® fusion proteins.

Schematic overview of HaloPROTAC function.


The HiBiT-BRD4 degradogram. BRD4, endogenously tagged with HiBiT and HaloTag® label, is degraded using increasing concentrations of HaloPROTAC and quantified using HiBiT luminescence.
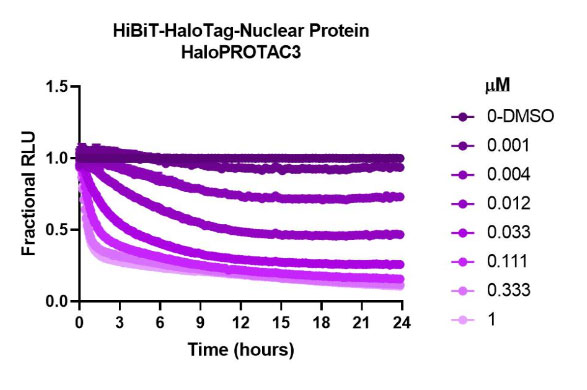
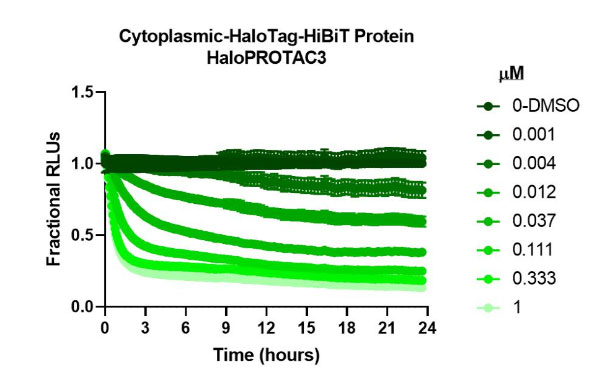
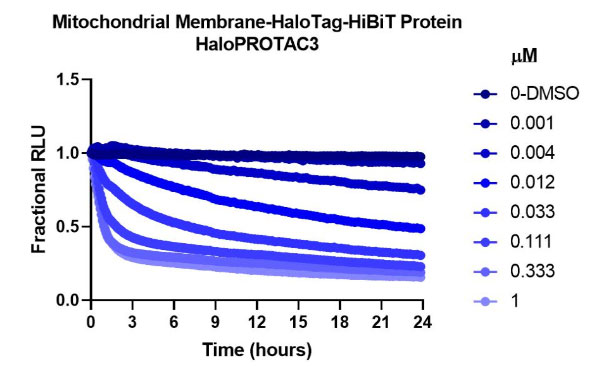
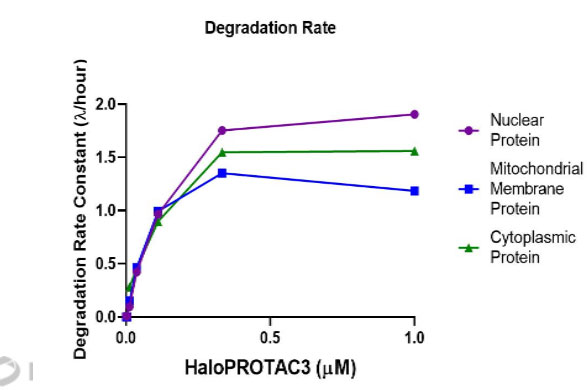
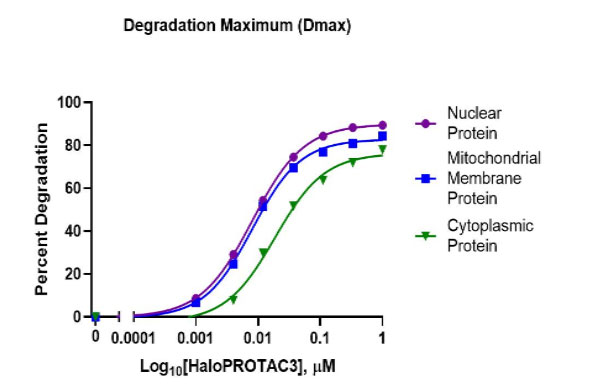
HaloPROTAC3 degradation of HiBiT-HaloTag CRISPR targets in various compartments. Results show rapid, burst degradation of targets in multiple cellular compartments, including single-pass mitochondrial membrane protein, with HaloPROTAC3. This confirms the proteins can be targeted via proteasomal degradation strategies.
Ready to start your degradation assays with HaloPROTAC3?
View HaloPROTAC Ligands
Need help with Assay Development, Screening or Profiling?
Learn more about our targeted protein degradation services.