Analyses of gene disruption by whole-cell PCR using the GoTaq® Green Master Mix
Howard Hughes Medical Institute and Department of Cell and Developmental Biology, Vanderbilt University Medical Center, Nashville, TN 37232
Publication Date: 2005
Abstract
We used GoTaq® Green Master Mix for PCR directly from yeast colonies to analyze targeted gene disruptions in S. pombe.
Introduction
We used Promega GoTaq® Green Master Mix to check the targeted deletion of genes from the genome of fission yeast, S. pombe. We tested the targeted disruption of ase1+, encoding a conserved microtubule-bundling protein(1)(2). GoTaq® Green Master Mix is a premixed, ready-to-use solution containing GoTaq® DNA Polymerase, dNTPs, MgCl2, reaction buffers and gel loading solution.
Materials and Methods
An S. pombe strain was transformed with a DNA fragment carrying the kanMX6 cassette, conferring G-418 resistance, flanked by ase1 target sequences(1). Colonies resistant to G-418 were screened by PCR for correct disruption of the ase1+ gene.
S. pombe strains were maintained in YE medium(3). The oligonucleotide primers designed to check the disruption of ase1+ were obtained from IDT DNA technology (Coralville, IA) and are as follows:
Ase1 5´FL: 5´ ttactttcgcaggaaaaggc 3´
KanMXRev: 5´ ccggcgcaggaacactgcc 3´
We tested reactions in a final volume of either 10µl or 20µl. In our experience, the 20µl reactions worked best. For each primer, 0.5µl of a 20µM stock was added to each reaction and was diluted to 10µl with water. To this primer mix, 10µl of GoTaq® Green Master Mix was added to have a final volume of 20µl per reaction. A toothpick of cells was added to the reactions, and PCR was performed under the following conditions: 95°C for 5 minutes; 12 cycles of 95°C for 45 seconds, 60°C for 1 minute and 72°C for 2.5 minutes; 25 cycles of 95°C for 45 seconds, 52°C for 45 seconds and 72°C for 2.5 minutes. The reaction products were separated on a 1% agarose gel and visualized.
Results
The results of the PCR are shown in Figure 1. Lanes 1, 2 and 6 contain a PCR product of ~800bp representing the disruption of ase1+ with kanMX6. Lanes 9 and 10 represent positive and negative control for the disruption, respectively. The low molecular weight bands presumably represent excess primers and/or denatured DNA from the reactions. As shown in Figure 1, the GoTaq® Green Master Mix is very efficient in the amplification of DNA fragments from yeast colonies. In summary, GoTaq® Green Master Mix is easy to use, reliable, versatile and efficient in the generation of PCR products from yeast colonies.
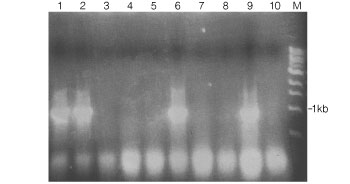 Figure 1. Agarose gel stained with ethidium bromide to detect PCR products generated from S. pombe colonies.
Figure 1. Agarose gel stained with ethidium bromide to detect PCR products generated from S. pombe colonies.
Lanes 1–8, colonies that were G-418 resistant; lane 9, an ase1::kanMX6 strain (a positive control for disruption); lane 10, a wildtype strain that is G-418 sensitive (a negative control for disruption); and lane M, a 1kb DNA ladder.
Related Products
Related Resources
Article References
- Loiodice, I. et al. (2005) Ase1p organizes antiparallel microtubule arrays during interphase and mitosis in fission yeast. Mol. Biol. Cell 16, 1756–68.
- Yamashita, A. et al. (2005) The roles of fission yeast ase1 in mitotic cell division, meiotic nuclear oscillation, and cytokinesis checkpoint signaling. Mol. Biol. Cell 16, 1378–95.
- Moreno, S., Klar, A. and Nurse, P. (1991) Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 194, 795–823.
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Venkatram, S. Analyses of gene disruption by whole-cell PCR using the GoTaq® Green Master Mix. [Internet] 2005. [cited: year, month, date]. Available from: https://www.promega.com/resources/pubhub/enotes/analyses-of-gene-disruption-by-whole-cell-pcr-using-the-gotaq-green-master-mix/
American Medical Association, Manual of Style, 10th edition, 2007
Venkatram, S. Analyses of gene disruption by whole-cell PCR using the GoTaq® Green Master Mix. Promega Corporation Web site. https://www.promega.com/resources/pubhub/enotes/analyses-of-gene-disruption-by-whole-cell-pcr-using-the-gotaq-green-master-mix/ Updated 2005. Accessed Month Day, Year.
Products may be covered by pending or issued patents or may have certain limitations on use.
 Watch this animated, step-by-step introduction to PCR--a landmark molecular biology technique.
Watch this animated, step-by-step introduction to PCR--a landmark molecular biology technique.