Efficient High-Throughput Protein Purification Using the Magne™ HaloTag® Beads
Promega Corporation
Publication Date: May 2012
Abstract
The growing demand to rapidly purify vast numbers of proteins for structural and functional studies is driving development of high-throughput protein purification methods. The Magne™ HaloTag® Beads allow protein purification through covalent capture of HaloTag® fusion proteins and subsequent release of these target proteins by TEV protease cleavage without multiple centrifugation steps, providing a simple purification that can be performed on robotic platforms. We developed an automated high-throughput pipeline for protein expression and purification on a Tecan Freedom® robotic workstation using the Magne™ HaloTag® Beads. The high binding capacity of the beads allowed efficient purification of proteins over a wide range of molecular weights and expression levels.
Introduction
The emerging field of functional proteomics is driving the need to develop high-throughput approaches to express and isolate a large number of gene products quickly for comprehensive analyses. Successful high-throughput protein purification generally requires efficient expression of these proteins in a soluble form followed by generic purification methods that are applicable to all proteins. The most frequently used host for high-throughput protein expression is E. coli because of its ease of use, rapid cell growth and low cost of culturing (1) . In addition the availability of auto-induction protocols minimizes manipulation and enables fully automated growth of cultures without monitoring growth or adding an inducer at the appropriate time. Once expressed, the simultaneous isolation of many proteins creates a technical challenge because proteins express at different levels and display different folding and biophysical characteristics (2) . To this end, affinity tags are commonly used to provide a generic, single-step chromatography using the affinity property of the tag. Affinity tags allow purification of multiple proteins on a single platform (3) (2) . In addition to facilitating purification, some of these tags also enhance protein expression and solubility in E. coli (3) . Although hundreds of proteins can be purified manually in a 96-well format, the tedious process would benefit from automation. Automation would increase throughput at each stage, enable hands-off operation and reduce human error, resulting in higher reproducibility and less variation among replicates. Magnetic affinity resins are suitable because they eliminate the need for multiple centrifugation steps and are amenable to automation.
We have developed a pipeline for high-throughput protein expression and purification using the HaloTag® technology (Figure 1) (4) . The HaloTag® protein is a fusion tag engineered to enhance expression and solubility of recombinant proteins in E. coli and provides efficient protein purification through covalent capture of HaloTag® fusion proteins onto the Magne™ HaloTag® Beads (Cat. # G7471, G7472, G7473) (5) . The highly specific and irreversible binding to the beads enables efficient protein capture regardless of the expression level and eliminates protein loss during wash steps, resulting in high protein recovery. Since covalently bound fusion protein can’t be eluted by conventional methods, the protein of interest is released from the beads by specific cleavage at an optimized TEV protease recognition site in the linker sequence separating the target protein from the tag.
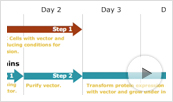
 Figure 1. Schematic overview of the protein expression and purification pipeline.
Figure 1. Schematic overview of the protein expression and purification pipeline. Method Overview
Expression: Recombinant protein clones generated by subcloning into pFN18A/K HaloTag® T7 Flexi® Vector (Cat.# G2751 and G2681) were used to transform Single Step (KRX) cells. E. coli KRX cells contain a chromosomal copy of the T7 RNA polymerase gene tightly regulated by a rhamnose promoter, which is subjected to catabolite repression by glucose. The presence of rhamnose activates the promoter, but only when the glucose is completely consumed, providing control of expression driven by the T7 promoter. Thus auto-induction is possible through titration of glucose and rhamnose concentrations (6) .
Lysis: Because sonication and centrifugation are difficult to incorporate into automated platforms, we took advantage of the FastBreak™ Cell Lysis Reagent (Cat.# V8571), which offers a convenient method for lysis of E. coli cells.
Purification: Following cell lysis, the HaloTag® fusion protein is covalently captured using the Magne™ HaloTag® Beads, and nonspecifically interacting proteins are washed away. The protein of interest is released by specific proteolytic cleavage at an optimized TEV protease recognition site contained within the amino acid linker sequence connecting HaloTag® protein and the protein of interest. The use of a TEV protease fused to HaloTag® (HaloTEV protease, Cat.# G6601, G6602), which is covalently captured on the Magne™ HaloTag® Beads, eliminates the need for post-cleavage removal of the protease, resulting in a streamlined purification process. This straightforward purification uses a single, mild physiological buffer throughout the entire purification process with no need for buffer exchange (Figure 2).
 Figure 2. Schematic diagram of protein purification using the Magne™ HaloTag® Beads.
Figure 2. Schematic diagram of protein purification using the Magne™ HaloTag® Beads. Automated Protein Purification Workstation
The entire protein purification process was performed on the deck of a Tecan Freedom® robotic workstation (Figure 3). The LiHa 8-tip liquid handling arm was used to transfer the lysates from the 96-well 2ml block to the 96-well 500µl block. The TeMO 96-tip liquid-handling system was used in all the other steps. The MagnaBot® II Magnetic Separation Device (Cat.# V8151), a 24-pin magnet designed to process 96-well plates, was positioned on the deck and was used for all magnetic separations. Processing one 96-well plate from cells to purified proteins requires 4.5 hours; five plates can be processed in parallel within seven hours.
 Figure 3. Tecan Freedom® robotic workstation.
Figure 3. Tecan Freedom® robotic workstation. The entire purification is performed on the deck of the Tecan Freedom® robotic workstation. The flowthrough and washes can be collected for analysis or discarded.
High Recovery and Reproducibility
To demonstrate the efficiency and reproducibility of the high-throughput purification using the Magne™ HaloTag® Beads, we monitored the binding, recovery and well-to-well variability using two reporter genes: Firefly (FFLY) and Renilla (hRL) luciferase. We inoculated a 96-well plate with KRX cells containing HaloTag®:FFLY and HaloTag®:hRL fusions in an alternate-column pattern and followed the protocols for expression and purification outlined in Figures 1 and 2. Since the crude lysate often contains soluble as well as insoluble protein, we took advantage of the reporter genes and monitored only the binding and purification of soluble functional protein. Our results indicate that for both reporters, 65% of the functional HaloTag® fusion was bound to beads, and more than 65% of the functional protein of interest was purified (Figure 4). This highly efficient purification is probably the result of covalent protein binding, which results in highly efficient capture of HaloTag® fusion proteins directly from the crude lysate and minimizes protein loss during wash steps. The low well-to-well variability with relatively low standard deviations of 5–7% indicates a robust and highly reproducible method.
 Figure 4. Efficient protein purification with reproducible yield and low variability.
Figure 4. Efficient protein purification with reproducible yield and low variability. Normalized volumes of crude lysate, flowthrough and eluted protein were analyzed for luciferase activity. Firefly luciferase activity (Panel A) was determined using the Bright-Glo™ Luciferase Assay (Cat.# E2610), and Renilla luciferase activity (Panel B) was determined using the Renilla-Glo™ Luciferase Assay (Cat.# E2710). Luminescence was measured using a microplate luminometer. Binding efficiency (%) was determined by comparing the activities in the crude lysate and the flowthrough. The recovery (%) was calculated as the fraction of the active protein that was purified. The average recovery for n = 40 was calculated to be 68% for Renilla luciferase and 69% for Firefly luciferase with standard deviations of 7% and 5%, respectively.
Minimal Cross-Contamination
To demonstrate the low cross-contamination of this method, we used the same reporter genes: firefly (FFLY) and Renilla (hRL) luciferase. Proteins were purified from a 96-well plate containing E. coli KRX cells expressing HaloTag®:FFLY and HaloTag®:hRL fusions plated in an alternate-column pattern. Purified proteins from each well were analyzed by SDS-PAGE, and no significant contamination was observed (Figure 5, Panel A). To increase the sensitivity of the cross-contamination test, each well was assayed for FFLY as well as hRL activities (Figure 5, Panel B). Our results show no significant activity cross-contamination between wells (<0.12%).
 Figure 5. Analysis of cross-contamination between wells.
Figure 5. Analysis of cross-contamination between wells. Panel A. SDS-PAGE analysis of purified proteins. Purified HaloTag®-firefly and Renilla proteins were resolved by SDS-PAGE and visualized by SimplyBlue™ staining. M = molecular weight markers. Panel B. Luciferase activity of purified proteins in each well. Purified protein in each well was tested for firefly luciferase activity using the Bright-Glo™ Luciferase Assay and Renilla luciferase activity using the Renilla-Glo™ Luciferase Assay. Luminescence was measured on a microplate luminometer.
Parallel Protein Purification from Multiple Clones with Different Expression Levels
To demonstrate further the utility of automated protein purification using the Magne™ HaloTag® Beads, we purified 12 different proteins with a broad range of molecular weights and expression levels. KRX cells harboring 12 different clones were inoculated into a 96-well plate (each column with a different clone) and processed following the protocols outlined in Figures 1 and 2. Our results indicated that all 12 proteins, regardless of the expression level, could be purified (Figure 6). Furthermore, all of the protein preparations were highly pure, as determined by SDS-PAGE analysis. These parallel purifications also can be used as a screening tool for expression levels and purification yields of different variants or mutants prior to scaling up.
 Figure 6. SDS-PAGE Analysis of purified proteins.
Figure 6. SDS-PAGE Analysis of purified proteins. Purified proteins from two wells for each of the 12 clones is shown. Proteins were resolved by SDS-PAGE and visualized by SimplyBlue™ staining. *Apparent MW of Protein G is 30kDa.
Experimental Considerations
We found that several parameters can greatly affect reproducibility and efficiency of protein purification. 1) Induction at a late growth stage increased the fraction of soluble protein, especially for hard-to-express proteins. The auto-induction protocol described here induced expression at a late stage in the growth phase. 2) The use of a 500µl block for purification minimized splashing and enabled robust purification with no significant loss of beads. 3) In the wash and cleavage steps, pipetting after each addition was necessary for resuspension of the beads.
Conclusions
An ideal protein affinity purification resin is defined by several criteria: minimal nonspecific protein binding, high binding capacity for the fusion protein and efficient recovery of the desired protein. The Magne™ HaloTag® Beads meet these criteria by combining the benefits of good binding capacity (>20mg/ml), low nonspecific binding (< 0.1% of the crude lysate) and highly efficient protein capture through covalent immobilization. The covalent capture allows efficient immobilization of HaloTag® fusion proteins regardless of expression levels and minimizes loss of bound protein during washes, resulting in high recovery of the expressed protein. The paramagnetic property of the beads makes them amenable to automated high-throughput protein purification. Here we combined a Tecan Freedom® robotic platform with the Magne™ HaloTag® Beads to provide an excellent method for high-throughput parallel protein production. This combination provides highly reproducible yields, minimal cross-contamination and minimal hands-on work. The methodology provides an efficient and reliable means to increase the number of protein constructs that can be purified and tested and enables screening of conditions for protein expression and purification. These beads can be used on other robotic platforms compatible with magnetic separation such as the Maxwell® 16 Instrument (7) .
Related Products
Related Protocols
Related Resources
Article References
- Baneyx, F. (1999) Recombinant protein expression in Escherichia coli. Curr. Opin. Biotechnol. 10, 411–21.
- Lesley, S.A. (2001) High-throughput proteomics: Protein expression and purification in the post-genomic world. Protein Expr. Purif. 22, 159–64.
- Nilsson, J. et al. (1997) Affinity fusion strategies for detection, purification, and immobilization of recombinant proteins. Protein Expr. Purif. 11, 1–16.
- Encell, L.P. et al. (in press) Development of a dehalogenase-based protein fusion tag capable of rapid, selective and covalent attachment to customizable ligands. Curr. Chem. Genomics
- Ohana, R.F. et al. (2009) HaloTag7: A genetically engineered tag that enhances bacterial expression of soluble proteins and improves protein purification. Protein Expr. Purif. 68, 110–20.
- Schagat, T. et al. (2008) KRX auto-induction protocol: A convenient method for protein expression. Promega Notes 98, 16–8.
- Bratz, M. et al. (2012) Using cell-free expression systems to study HDAC protein functions. [Internet; cited: 2012, 04/20] Promega PubHub
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Ohana, RF, Vidugiris, G, Encell, LP and Slater, MR Efficient High-Throughput Protein Purification Using the Magne™ HaloTag® Beads. [Internet] May 2012. [cited: year, month, date]. Available from: https://www.promega.com/resources/pubhub/efficient-high-throughput-protein-purification-using-the-magne-halotag-beads/
American Medical Association, Manual of Style, 10th edition, 2007
Ohana, RF, Vidugiris, G, Encell, LP and Slater, MR Efficient High-Throughput Protein Purification Using the Magne™ HaloTag® Beads. Promega Corporation Web site. https://www.promega.com/resources/pubhub/efficient-high-throughput-protein-purification-using-the-magne-halotag-beads/ Updated May 2012. Accessed Month Day, Year.
Freedom is a registered trademark of Tecan AG Corporation. GenBank is a registered trademark of US Dept of Health and Human Services. IGEPAL is a registered trademark of Rhone-Poulenc AG Co. SimplyBlue is a trademark of Invitrogen Corporation.
 Learn about competent cells designed for tightly controlled protein expression.
Learn about competent cells designed for tightly controlled protein expression.