Evaluation of Haplotypes to Enhance the Power of STR Test Kits in Cases of Questioned Family Relationships
Department of Forensic Sciences, Center for Health Sciences Oklahoma State University 1Current address: Texas Department of Public Safety
Publication Date: 2012
Introduction
Multiplex STR typing kits available commercially today are robust and exhibit a very high discriminatory power in cases of questioned family relationships, especially cases in which the questioned relationship is one of alleged parent and child. While existing kits are very powerful, there are cases in which the questioned relationship is one involving individuals more distant within the pedigree. Often, immigration cases submitted to a parentage testing laboratory involve individuals who claim to be an alleged aunt, uncle, niece, nephew or half-sibling. In such cases, standard STR kits may fail to resolve the case. Other examples of cases at risk to remain unresolved include those involving the identification of human remains in which only distant family members of the deceased are available to provide reference samples. In addition, in a portion of those cases, the victim and/or key family reference samples will be female, so Y-STR analysis will be uninformative.
One strategy to increase the discriminatory power of an STR test battery is to add additional loci. Supplemental STR kits have been developed, and some are available commercially such as the PowerPlex® CS7 System (Promega Corporation, Madison, WI). A 26-plex STR typing kit has been developed and may become available in the future (1) . A second approach could be to identify additional genetic markers linked to existing STR loci in a kit such that haplotypes rather than allele frequencies could be used in relatedness calculations. This strategy was applied by Lewis et al. (2) to resolve a case of questioned relatedness in which a World War II victim was identified through the use of haplotype comparisons between the remains and an alleged second cousin. The increase in discriminatory power results from the lower frequency of an obligate haplotype as opposed to that for the individual obligate alleles that compose it. Such has been the case with HLA haplotypes used in parentage testing for years, with haplotypes exhibiting much lower frequencies than the individual HLA antigens that compose them (3) (4) (5) .
The FESFPS and Penta E loci contained within the PowerPlex® 16 and CS7 Systems (Promega Corporation) have been shown to be linked on chromosome 15 (6) . The distance between the loci is estimated to be about 6 centimorgans (1 centimorgan [cM] corresponds to about 1% recombination between the loci) (7) (8) . Even though these loci might not represent an ideal haplotype model for family relationship testing due to the rather moderate linkage exhibited by the loci, this model nonetheless provided an opportunity to investigate whether or not haplotypes used for family relationship testing increased the power of a test battery in any useful way.
Materials and Methods
Source of DNA samples: Archival parentage casework was subjected to DNA typing with the FFFL or Penta E typing kits (Promega Corporation). In some later analyses, the PowerPlex® CS7 multiplex kit, which contains these loci plus five others, was used. One hundred cases were chosen from each of the Black and Caucasian ethnic groups and involved a mother, one or more children and an alleged father who was conclusively shown to be the father of the child(ren). Haplotypes were identified in the child by determining the pair of alleles for the FESFPS and Penta E loci transmitted from each parent. Of the cases chosen, most were found to be informative in both Blacks and Caucasians for the transmitted haplotype. Eighteen families in the Black group included multiple children, which enabled recombinants between FESFPS and Penta E to be identified and the frequency of recombination between the loci to be roughly estimated.
Data analysis: Haplotype frequency databases were created from test results obtained from both Black and Caucasian ethnic groups. Four hundred haplotypes were scored in the Black population and 396 in the Caucasian population. The distribution of haplotypes in the two populations was evaluated using a contingency table producing a chi square statistic (Table 1) and also using Kolmogorov-Smirnov distribution analysis. For the chi square analysis, haplotypes were grouped based on the FESFPS allele in the haplotype, and counts in some categories were pooled based on the FESFPS allele in the haplotype to obtain sufficient numbers for the analysis (see Table 1 for the contingency table groupings).
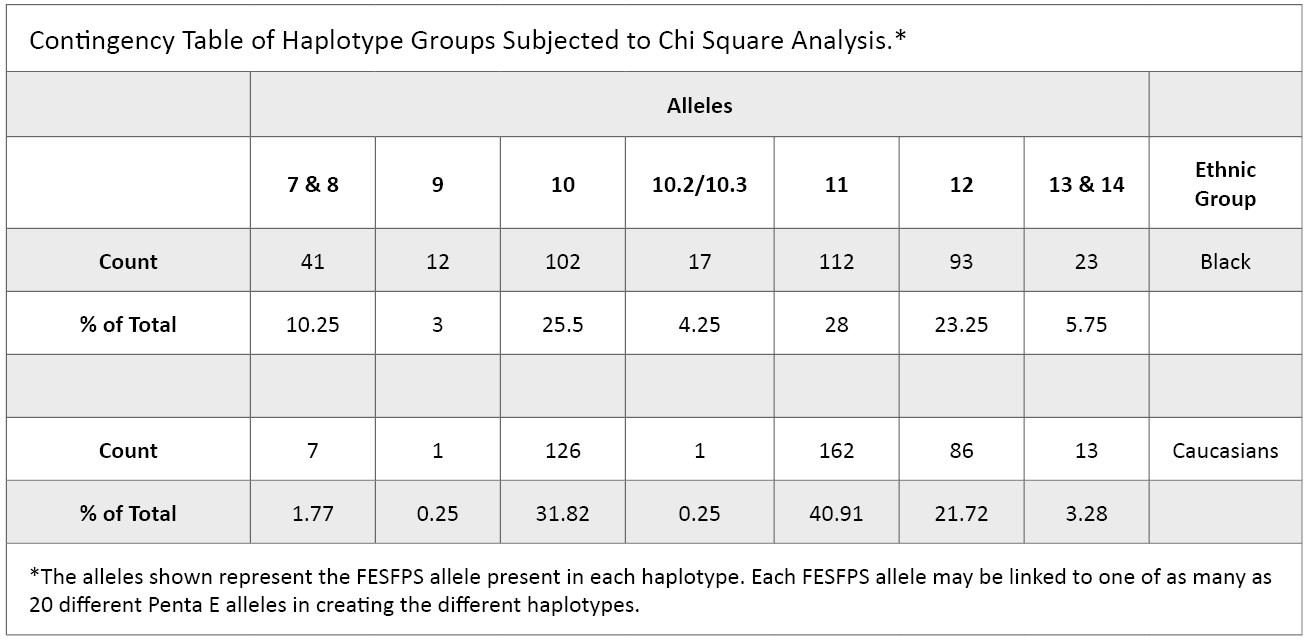 Table 1. Contingency table of haplotype groups subjected to chi square analysis.
Table 1. Contingency table of haplotype groups subjected to chi square analysis. Frequency estimations for all haplotypes were set to the 95% confidence level using the following methods: For haplotypes not seen in one ethnic group the frequency was estimated using the formula 1 – α1/N, where N equals the total number of haplotypes in the database and alpha equals 0.95. For all other haplotypes, the formula p + 1.96√[p(1–p)/N] was applied, where p is the observed frequency of a haplotype and N is the total number of haplotypes in the database.
Haplotype frequencies were used to calculate likelihood ratios (LR) in each of the 200 cases, which were then compared with the FESFPS or Penta E LR values for those cases. The higher LR value for FESFPS or Penta E in each calculation was compared against the LR value produced when using haplotype frequencies instead of allele frequencies.
Results
STR analysis of the family trios for the FESFPS and Penta E loci identified 396 haplotypes in Caucasians, creating 57 different haplotype groups, while in Blacks, 400 haplotypes, which fell into 96 different groups, could be identified (Table 2). In all, 102 different haplotypes were scored in both populations (Table 2). Multiplying the number of FESFPS alleles known to exist by the number of Penta E alleles produces a theoretical number of FESFPS-Penta E haplotypes of 234. Thus, in our population sampling, less than 50% of possible haplotypes were detected.
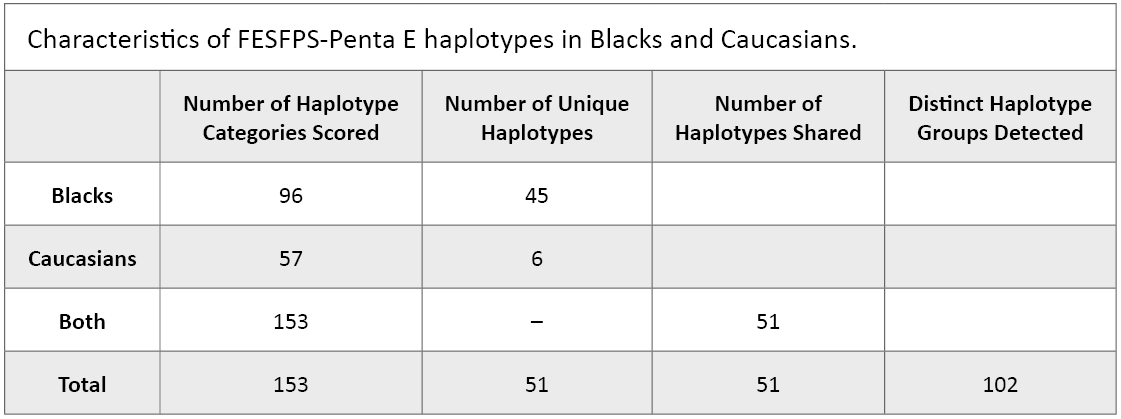 Table 2. Characteristics of FESFPS-Penta E haplotypes in Blacks and Caucasians.
Table 2. Characteristics of FESFPS-Penta E haplotypes in Blacks and Caucasians. In Caucasians, six haplotypes were scored that were not seen in Blacks, whereas 45 haplotypes were scored in Blacks that were not seen in Caucasians (Table 2). A simple scan of the frequencies in the two groups suggested that haplotype variability in Blacks considerably exceeded that of Caucasians, an observation underscored by the number of distinct haplotypes in the two groups (Table 2).
The distributions of haplotypes in the two racial groups were compared in several ways. The Kolmogorov-Smirnov analysis can be used to compare the distribution of two groups of data and provide a sense of how similar or different they are. Kolmogorov-Smirnov analysis of haplotypes demonstrated that they were quite dissimilar (p<0.0001). As a second evaluation of haplotype distributions, contingency table analysis using the chi square approach confirmed that the distributions of haplotypes in the two groups were significantly different (p<0.0001).
It should be noted that in many family trios, the haplotype observed in a child and thus traced back to a parent could represent a recombination event in the gametes produced by either parent that contributed to the conception of that child. Since the child’s haplotype forms the starting point for identifying haplotypes in the parents, the identification and scoring of parental haplotypes in the frequency database may occasionally be in error (perhaps by as much as 16% of the observed frequency for a particular haplotype, see below).
Among the parentage cases from the Black population, there were 18 families with multiple children. Out of 18 multichild families, with a total of 51 children, eight families had a child in which the FESFPS-Penta E haplotype differed from the other children in the family. This is most logically explained through recombination, which based on these data, suggests a recombination rate between FESFPS and Penta E of about 16%. There was an insufficient number of Caucasian families with multiple children to detect any recombination events. In families with only two children, recombination could be detected. However, distinguishing which haplotype was the recombinant and which was the nonrecombinant was not possible when only two children were available for testing.
The principal goal of this study was to investigate the enhanced discriminatory power of an STR test battery afforded through the use of linked STR loci. The LRs from the initial testing in the 200 family trios were compared with LRs calculated with haplotype frequency. A comparison is shown graphically in Figure 1. For the Black racial group, the use of haplotypes enhanced the LR value for the parent:child comparison in about 85% of calculations (i.e., the haplotype LR/allele LR ratio exceeded 1.0). For Caucasians, about 95% of the haplotype LR/allele LR ratios exceeded 1.0 (Figure 1). The small number of calculations in which the use of allele frequencies produced LR values greater than that produced with haplotype frequencies is likely to stem from the use of frequencies corrected to the 95% confidence interval, which creates a minimal haplotype frequency that is greater than the minimal allele frequency due to the size of the respective databases. In general, use of haplotype frequencies rather than allele frequencies raised the combined LR for the case. In some cases, the LR was increased by five- to tenfold (especially in the Black group). In other cases, use of haplotype frequencies had only a marginal effect on the combined LR. The average increase in LR for cases in the Black racial group was 2.4-fold, whereas in Caucasians it was 1.8-fold.
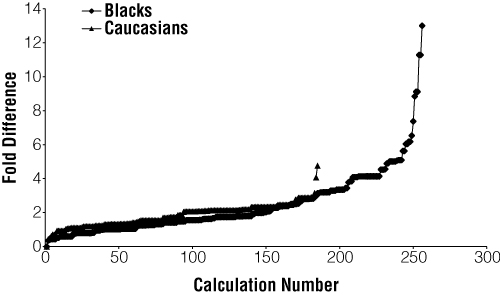 Figure 1. Comparison of fold increase (or decrease) in the likelihood ratio produced using allele frequencies for either the FESFPS or Penta E locus (whichever was higher) or the FESFPS-Penta E haplotype.
Figure 1. Comparison of fold increase (or decrease) in the likelihood ratio produced using allele frequencies for either the FESFPS or Penta E locus (whichever was higher) or the FESFPS-Penta E haplotype. Likelihood ratios (LRs) were calculated using FESFPS or Penta E allele frequencies (whichever produced the higher value), and FESFPS-Penta E haplotypes for both paternity and maternity for each child in the 100 archived cases for each racial group. The LR values produced with each method were compared as a ratio of values calculated using haplotype frequencies versus allele frequency. The ratios were then sorted and plotted from smallest to largest for each racial group. Abscissa: The LR values produced in the various parentage tests (sorted according to the ratio produced). Ordinate: The magnitude of the ratio comparing LR produced with haplotype frequency versus that produced using the FESFPS or Penta E allele frequency(ies), depending on which value produced the higher value.
Discussion
Relationship testing laboratories are increasingly facing challenges in resolving questioned pedigrees in which individuals being tested are presumably related in some way other than parent and child, or perhaps sibling. Such situations are common in immigration cases and cases involving identification of human remains. While one possible solution in such cases is to apply additional STR markers, currently available STR kits may still fail to provide sufficient power. We therefore investigated one potential approach to increasing the discriminatory power of our test battery using readily available STR loci (FESFPS and Penta E) that were recently reported to be statistically linked (8) . By collecting haplotypes of FESFPS-Penta E alleles, it was possible to use both loci in LR calculations (rather than using only one, normally the highest in LR). The average increase in combined LR when using haplotypes was about twofold in both Blacks and Caucasians (2.43 versus 1.84, respectively). Undoubtedly, this average would change, perhaps significantly, if more haplotypes were in the database.
When ratios of LRhaplotype/LRallele were plotted in a sorted fashion, 6% or 15% (depending on race) of ratios fell below 1.0. Of course this would be unexpected unless one considers the minimal frequency for an allele from a database of 400–500 individuals versus the minimal frequency that would be assigned to rare haplotypes in a database less than half that size. In this scenario, some haplotypes might actually have calculated frequencies higher than those of the alleles that compose them. It is likely that with more haplotypes in the database, there would be few if any ratios at or below 1.0. It should be noted that even though the average fold increase in LR associated with the use of haplotypes was about 2.0, in some cases a twofold increase in cumulative LR for a case could resolve it.
There was almost a twofold difference in the number of distinct haplotypes observed in Blacks and Caucasians (96 versus 57), suggesting that haplotype diversity is much higher in Blacks than Caucasians. This observation is important for the application of haplotype frequency calculations in immigration cases, which often originate from Africa. The distribution of haplotypes in Blacks and Caucasians also was significantly different as determined by statistical analyses.
The FESFPS and Penta E loci have been suggested to exist about 6cM apart on chromosome 15 (7) (8) ; 1cM represents 1% average recombination between two loci (9) and typically encompasses about a million base pairs of chromosomal distance (10) .
Our collection of family trios contained 18 multichild families from the Black group with a total of 51 children. Among the 51 children, eight exhibited recombinant haplotypes confirmed through comparison to other children in the family. Thus, in our study, there was about 16% recombination between FESFPS and Penta E. It should be noted that Maha and collaborators (George Maha, Laboratory Corporation of America, personal communication) reported a similar recombination rate between FESFPS and Penta E in a study with a larger number of multichildren families. In a recent report from Phillips et al. (8) the recombination rate on chromosome 15 in the region of the FESFPS and Penta E loci was estimated at about 18%. Based on these results, recombination between the loci occurs with a higher frequency than expected based on their estimated distance apart on chromosome 15. It is possible that a recombination hot spot exists between these loci. It is also possible that, since both loci are located on chromosome 15 towards the end of the “q” arm (10) , recombination in this region is naturally higher than that towards the more central areas. Such an explanation has been proposed by Phillips et al. (8)
While the increase in LR provided to casework by using haplotypes was modest, the concept appears potentially useful. Hill et al. (1) reported the development of a 26-plex of STR loci distinct from those loci commonly included in commercially available multiplex kits. It is possible that some of the loci in the 26-plex described by Hill et al. (1) are linked to loci in the commercial kits. Thus, other haplotype systems are possible besides FESFPS-Penta E for use in complicated cases. It is also possible that other markers, linked to FESFPS-Penta E, would further increase the discriminatory power of this haplotype system.
Acknowledgements
The authors gratefully acknowledge Dr. Mark Payton, Oklahoma State University, for the assistance with statistical analysis of data.
Article References
- Hill, C.R., Butler, J.M and Vallone, P.M. (2009) A 26plex autosomal STR assay to aid human identity testing. J. Forensic Sci. 54, 1008–15.
- Lewis, K. et al. (2010) Identification of distant relatives using haplotypes constructed from multiple linked autosomal STRs. Paper presented at the American Academy of Forensic Sciences Annual Meeting, Seattle, Washington.
- Lamm, L.U. Gurtler, H. and Hansen, H. (1983) The HLA system. In: Inclusion probabilities in parentage testing, Walker, R.H., ed., AABB Press, Arlington, VA. 381–96.
- Smouse, P.E. and Adams, J. (1983) The use of closely linked genetic markers for paternity analysis. In: Inclusion probabilities in paternity testing, Walker, R.H., ed., AABB Press, Arlington, VA. 397–410.
- Chakravarti, A. and Li, C.C. (1983) The effect of linkage on paternity calculations. In: Inclusion probabilities in paternity testing, Walker, R.H., ed., AABB Press, Arlington, VA. 411–22.
- Butler, J.M. (2006) Genetics and genomics of short tandem repeat loci used in human identity testing. J. Forensic Sci. 51, 253–65.
- Hartl, D.L. and Jones, E.W. (2009) Genetics: Analysis of genes and genomes, 7th edition, Jones and Bartlett Publishers, Boston, MA. 156.
- Phillips, C. et al. (2012) The recombination landscape around forensic STRs: Accurate measurement of genetic distances between syntenic STR pairs using HapMap high density SNP data. Forensic Sci. Int. Genet. 6, 354–65.
- AABB (2010) Linked loci, FESFPS and Penta E. AABB Accreditation Relationship Testing News. 5(1),
- Lodish, H. et al. (2004) Molecular Cell Biology, fifth edition, W.H. Freeman, San Francisco, CA. 396.
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Worthen, C. et al. Evaluation of Haplotypes to Enhance the Power of STR Test Kits in Cases of Questioned Family Relationships. [Internet] 2012. [cited: year, month, date]. Available from: https://www.promega.com/resources/profiles-in-dna/2012/haplotypes-in-family-relationship-testing/
American Medical Association, Manual of Style, 10th edition, 2007
Worthen, C. et al. Evaluation of Haplotypes to Enhance the Power of STR Test Kits in Cases of Questioned Family Relationships. Promega Corporation Web site. https://www.promega.com/resources/profiles-in-dna/2012/haplotypes-in-family-relationship-testing/ Updated 2012. Accessed Month Day, Year.
PowerPlex is a registered trademark of Promega Corporation.
Contribution of an article to Profiles in DNA does not constitute an endorsement of Promega products.