Database Process Automation Using DNA IQ™, Plexor® HY and PowerPlex® 16 HS Systems on the Biomek® 3000 Workstation
Idaho State Police Forensic Services
Publication Date: 2011
Introduction
The Idaho State Police Forensic Services (ISPFS) Laboratory previously validated the automated extraction of convicted offender buccal FTA® samples using the DNA IQ™ System on the Biomek® 3000 workstation and found it to be more effective than manual extraction. Due to the wide range of DNA concentrations obtained, quantification and normalization was a necessary part of the routine processing of database samples for ISPFS. The lack of automation procedures beyond extraction required multiple steps in which the analyst performed hands-on dilutions, master mix preparation and sample transfers. Manual quantification setup of 96-well plates was relatively efficient when using multichannel pipettes, but some pipetting variability was observed between analysts. Manual normalization of samples and amplification setup was extremely laborious and time-intensive because they were not amenable to the use of multichannel pipettes. Although the Biomek® 3000 can be programmed to perform these functions, the ISPFS laboratory did not have the time or expertise to write effective or efficient methods. ISPFS sought readily available and reliable methods to automate DNA quantitation setup, normalization and amplification setup processes to reduce the potential for pipetting errors, increase efficiency and allow analyst efforts to be focused on downstream data analysis and review. The laboratory desired that the automation methods reduce the hands-on time of the analysts, perform the same (or better) than manual methods and produce consistent and reproducible results. The ISPFS laboratory evaluated the Promega Identity Automation™ methods for DNA IQ™ extraction, Plexor® HY setup, sample normalization and PowerPlex® 16 HS amplification setup on the Beckman Coulter Biomek® 3000 with these criteria in mind.
Method Comparison
The Biomek® 3000 DNA IQ™ extraction procedure was previously validated and has been used routinely by the ISPFS laboratory for processing convicted offender buccal FTA® samples; however, the method released with the Identity Automation™ procedures was modified by Promega and, therefore, was reevaluated as part of this study. The new extraction method includes addition of a second processing plate. The resin is transferred to this second plate during the first alcohol wash, increasing sample purity by preventing lysate residue from contaminating samples during heated elution. Additional features in the new Automated DNA IQ™ method for the Biomek® 3000 include the use of partial tip boxes, tip disposal, a sample start column prompt and a sample start volume prompt. The sample start volume prompt allows the preprocessing sample digest to be done with lysis buffer or Stain Extraction Buffer/proteinase K (SEB/proK). If SEB/proK is used instead of lysis buffer, the instrument will add lysis buffer to the sample at the beginning of the method. Three 3.2mm punches from eight different buccal FTA® samples were extracted using the original ISPFS-validated DNA IQ™ method, the updated DNA IQ™ method with a lysis buffer digest and the updated method with a SEB/proK digest. All three digests were performed in a 70°C water bath for 20 minutes. Extraction was evaluated by manual quantification on a single qPCR plate with the Quantifiler® Human DNA Quantification Kit per the manufacturer’s instructions. No significant differences in DNA yield were observed between the three extraction procedures (data not shown).
The Automated Plexor® HY Setup method for the Biomek® 3000 allows manual or automated standard preparation as well as manual or automated master mix preparation. Each option was tested throughout the evaluation period with acceptable results. The instrument dispenses the master mix to the amplification plate, then adds samples and standards to the master mix. If the user desires the samples to be quantified multiple times on the qPCR plate, the Automated Plexor® HY method provides a prompt for the analyst to indicate the number of replicates to be sequentially transferred. As a verification of the method, all 24 extracts previously obtained from the three DNA IQ™ methods were quantified with the Plexor® HY System both manually and using the Automated Plexor® HY method. Each plate was amplified on an Applied Biosystems 7500 Real-Time PCR System and data analyzed with the Promega Plexor® Analysis Software. Table 1 lists the DNA concentrations obtained for each preparation method. Although some differences were obtained, especially with higher-concentration samples, overall a good correlation of results between the two methods was observed. The analyst performing this study does not routinely mix each sample after adding it to the well containing master mix, whereas the Biomek® 3000 performs thorough mixing when adding each sample to the already dispensed master mix. The different sample addition methods used by the analyst and the robot could possibly account for some of the variation observed.
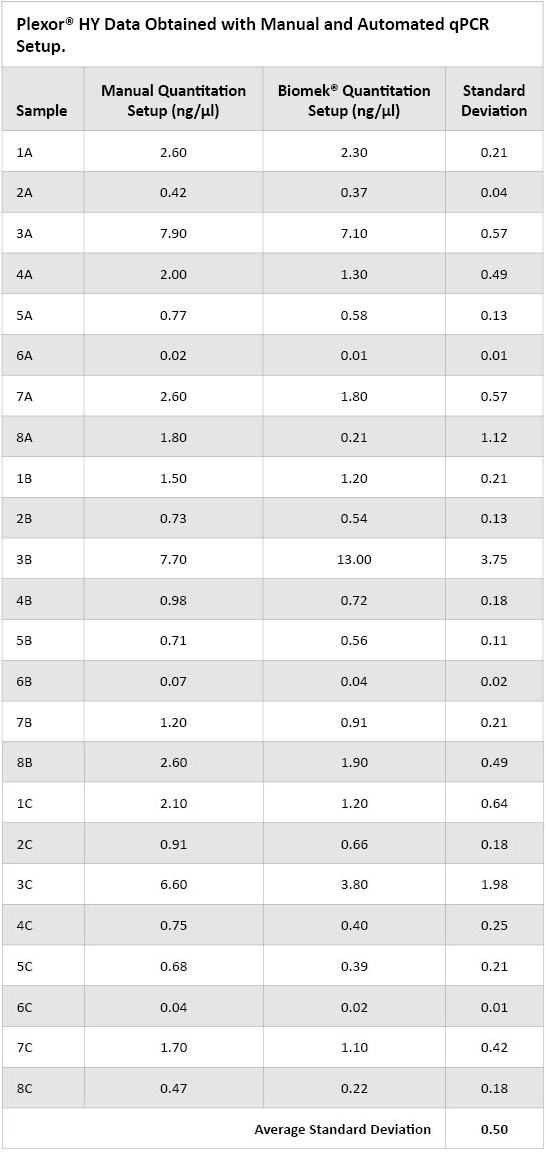 Table 1. Plexor® HY Data Obtained Using Manual qPCR Setup and Automated qPCR Setup.
Table 1. Plexor® HY Data Obtained Using Manual qPCR Setup and Automated qPCR Setup. The Automated Normalization and STR Reaction Setup method for the Biomek® 3000 works in conjunction with the STR Normalization Manager™ software. The STR Normalization Manager™ software is used to create templates that define the amount of DNA to be amplified (concentration and volume), number and types of controls as well as analysis methods and protocols to create a genetic analyzer plate document. The software also has several options for amplification plate setup based on sample type and concentration. For example, the user can choose whether or not to amplify samples if quantification values are low (cutoff determined by the laboratory) and, if a sample is not amplified, whether to skip that well or consolidate samples on the amplification plate. When the Biomek® Automated Normalization method is run, the STR Normalization Manager™ software automatically opens and prompts the user to choose the previously created template to be used as well as to input sample concentration information. The program allows manual entry of quantitation results or direct import of a .tab file. ISPFS currently uses a reduced-volume PowerPlex® 16 HS amplification. The Automated Normalization and STR Reaction Setup method can accommodate the small-volume transfers necessary for a reduced-volume reaction. Sixteen previously extracted and quantified samples were normalized and PowerPlex® 16 HS reduced-reaction amplification plates set up both manually and using the Biomek® Automated Normalization method with a target DNA concentration of 0.25ng. The plates were amplified on Applied Biosystems 9700 thermal cyclers using a 10/20 cycling protocol. The samples were subsequently run on an Applied Biosystems 3130xl Genetic Analyzer with 1µl amplified product and a 3-second injection time. Figures 1 and 2 show the average peak heights and peak height ratios observed at each locus for both methods. The overall peak heights and peak height ratios obtained were comparable between the two methods.
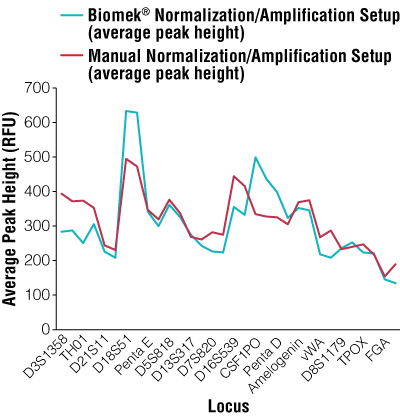 Figure 1. Average peak heights per locus for manual normalization and amplification setup versus automated normalization and amplification setup.
Figure 1. Average peak heights per locus for manual normalization and amplification setup versus automated normalization and amplification setup.
Overall manual peak height average = 306RFU; overall automated peak height average = 317RFU.
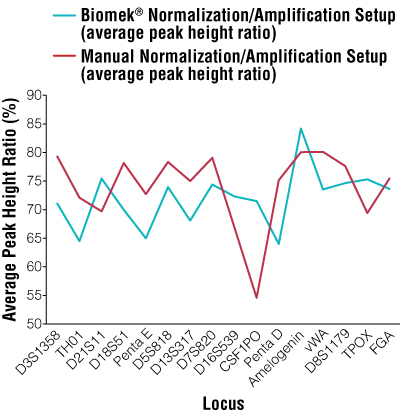 Figure 2. Average peak height ratios per locus for manual normalization and amplification setup versus automated normalization and amplification setup.
Figure 2. Average peak height ratios per locus for manual normalization and amplification setup versus automated normalization and amplification setup.
Overall manual peak height ratio average = 74%; overall automated peak height ratio average = 72%.
The 0.25ng target used in the normalization and amplification setup comparison was based on ISPFS’s original PowerPlex® 16 HS validation, in which quantification was performed with the Quantifiler® Human kit. Concentration values obtained with the Plexor® HY System differ from those of Quantifiler® Human kit. Based on the Plexor® HY data gathered during this study, re-optimization was performed by ISPFS, and subsequent amplifications were done using 0.5ng of template DNA.
Precision
Twenty-four different buccal FTA® samples were punched with the BSD600-Duet semi-automated puncher (three 3.2mm punches). Each sample was punched twice into two separate plates and extracted using the updated Automated DNA IQ™ method for the Biomek® 3000. Preprocessing for both plates was done using a lysis buffer digest at 70°C for 25 minutes. The first plate was digested in a water bath, and the second was done using the Biomek® heater with the Slicprep™ adaptor (Slicprep™ 96 Device, Cat. V1391). Quantification setup for each extracted plate was performed on the Biomek® 3000 using the Automated Plexor® HY method and samples replicated three times on each qPCR amplification plate. Both plates were amplified on an Applied Biosystems 7500 Real-Time PCR System and data analyzed using the Plexor® Analysis Software. Good correlation of replicates was achieved with no significant differences in quantitation results between the water bath and heater/adaptor digests (data not shown). All subsequent plates throughout the evaluation were preprocessed with the heater/adaptor digest.
An additional 16 buccal FTA® samples were punched on a single plate and extracted using the lysis buffer/heater digest at 70°C for 25 minutes. Quantification setup was performed using the Automated Plexor® HY Setup method for the Biomek® 3000. Each of the 16 samples was replicated five times on a single qPCR plate. The qPCR plate was amplified on an Applied Biosystems 7500 Real-Time PCR System and data analyzed using the Plexor® Analysis Software. Table 2 shows the concentration results for the five replicates of these 16 samples. Samples dispensed by the instrument provided reproducible results with very good correlation between replicates.
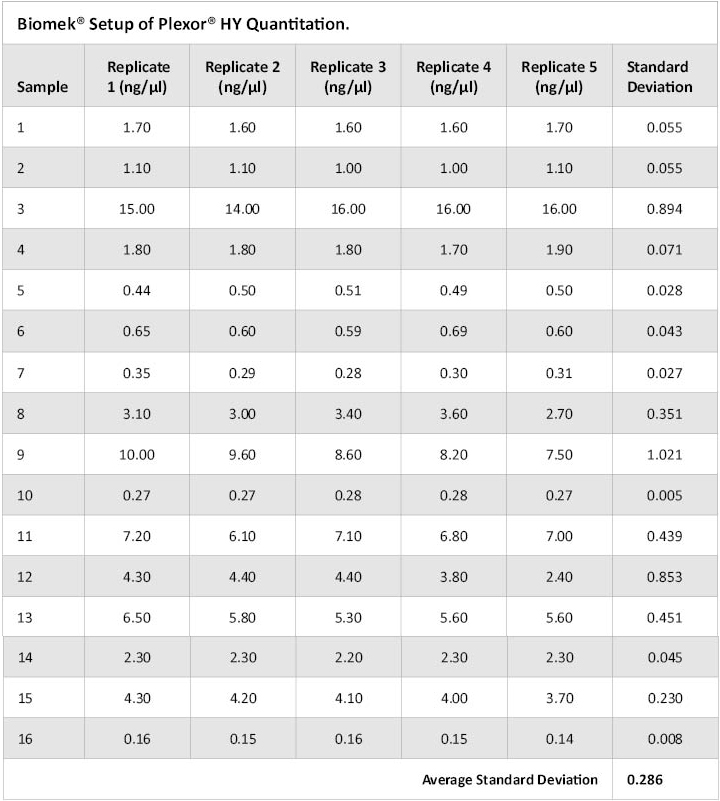 Table 2. Plexor® HY Concentrations Obtained for Five Replicates of 16 Samples Dispensed by the Biomek® 3000.
Table 2. Plexor® HY Concentrations Obtained for Five Replicates of 16 Samples Dispensed by the Biomek® 3000. A second Biomek® 3000 was purchased by ISPFS during the evaluation period. Quantification of previously extracted DNA from the 24 buccal FTA® samples (stored at –20°C) was set up in duplicate using the Automated Plexor® HY method on the new instrument. Samples were replicated three times on each of two qPCR plates, amplified on an Applied Biosystems 7500 Real-Time PCR System and analyzed with the Plexor® Analysis Software. Although concentration values were different than those previously obtained, likely due to sample age and evaporation, replicate precision was consistent with that of the first instrument (data not shown).
Accuracy
The 16 previously extracted and quantified samples used as part of the normalization precision study were used to check the normalization accuracy of the instrument. These samples were normalized both manually and with the Biomek® Automated Normalization method using a target concentration of 0.06ng/µl. In each case, 15µl of each normalized sample was dispensed to a 96-well plate for quantitation setup on the Biomek® 3000. An automated Plexor® HY qPCR assay was performed on each sample, and quantitation results were compared for the manual and Biomek® normalizations (Figure 3). Additionally, the DNA concentrations obtained from the Biomek®-normalized samples were compared to the 0.06ng/µl target value (Table 3). Although some variation is expected with any quantitation system, the concentrations obtained for the Biomek®-normalized samples were accurate with a very low standard deviation. The Biomek® 3000 performed as well as, and in a few cases slightly better than, manual normalization.
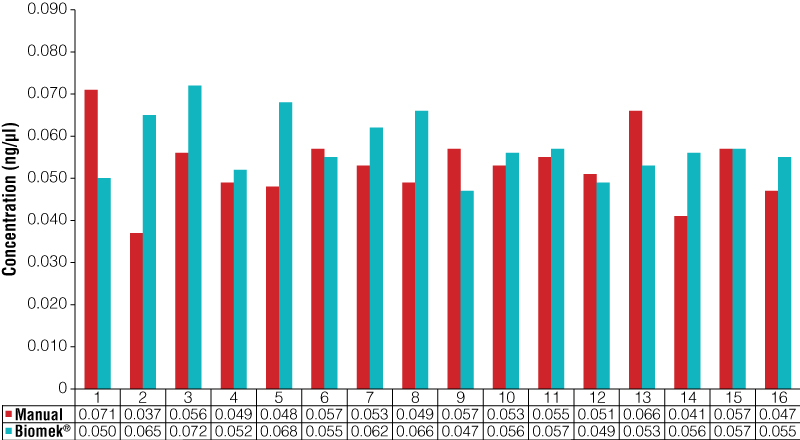 Figure 3. DNA concentrations obtained using the Plexor® HY System for samples normalized to 0.06ng/µl manually and using automation.
Figure 3. DNA concentrations obtained using the Plexor® HY System for samples normalized to 0.06ng/µl manually and using automation. 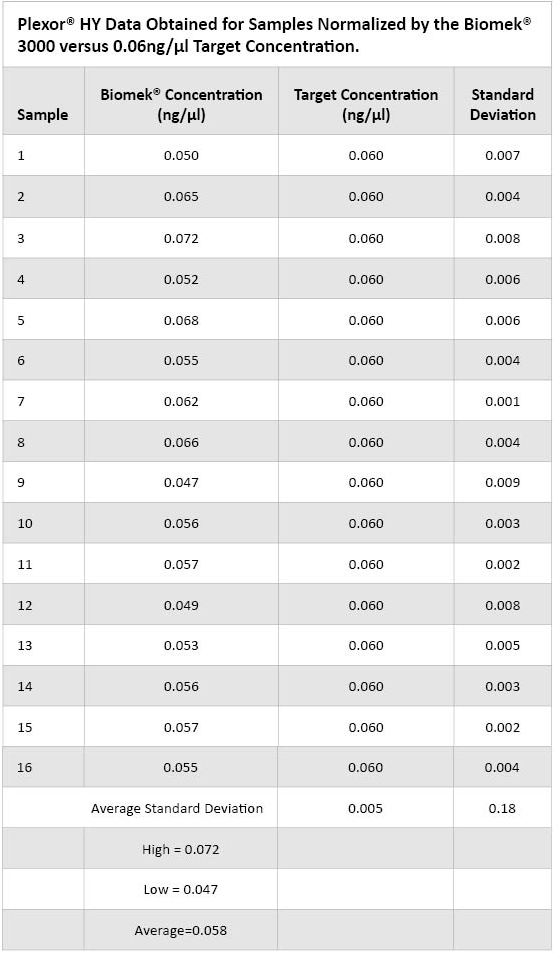 Table 3. Plexor® HY Data Obtained for Samples Normalized By the Biomek® 3000 versus the 0.06ng/µl Target Concentration.
Table 3. Plexor® HY Data Obtained for Samples Normalized By the Biomek® 3000 versus the 0.06ng/µl Target Concentration. Conclusions
The availability of prewritten and validated methods for automation is a benefit to laboratories interested in implementing robotic technology. Analyst time can be spent on internal validation and implementation rather than the time-consuming and often frustrating process of writing programs for the robotic workstation. The Identity Automation™ methods are a viable option that allows flexibility in chemistry and lab-specific modifications. Automation from extraction to amplification setup using the DNA IQ™, Plexor® HY and normalization methods on the Biomek® 3000 provides reproducible and accurate results that are consistent with those obtained manually. Efficiency is increased and the analyst’s hands-on time is greatly reduced when using this system for extraction, quantitation, normalization and amplification setup.
How to Cite This Article
Scientific Style and Format, 7th edition, 2006
Cunnington, C. Database Process Automation Using DNA IQ™, Plexor® HY and PowerPlex® 16 HS Systems on the Biomek® 3000 Workstation. [Internet] 2011. [cited: year, month, date]. Available from: https://www.promega.com/resources/profiles-in-dna/2011/database-process-automation/
American Medical Association, Manual of Style, 10th edition, 2007
Cunnington, C. Database Process Automation Using DNA IQ™, Plexor® HY and PowerPlex® 16 HS Systems on the Biomek® 3000 Workstation. Promega Corporation Web site. https://www.promega.com/resources/profiles-in-dna/2011/database-process-automation/ Updated 2011. Accessed Month Day, Year.
Contribution of an article to Profiles in DNA does not constitute an endorsement of Promega products.
Plexor and PowerPlex are registered trademarks of Promega Corporation. DNA IQ, Identity Automation, STR Normalization Manager and Slicprep are trademarks of Promega Corporation.
Biomek is a registered trademark of Beckman Coulter, Inc. FTA is a registered trademark of Flinders Technologies, Pty, Ltd., and is licensed to Whatman. Quantifiler is a registered trademark of Life Technologies.